| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 19,805,208 – 19,805,307 |
| Length | 99 |
| Max. P | 0.742529 |

| Location | 19,805,208 – 19,805,307 |
|---|---|
| Length | 99 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 81.87 |
| Mean single sequence MFE | -28.92 |
| Consensus MFE | -21.60 |
| Energy contribution | -22.05 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.672319 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19805208 99 + 27905053 ACCACAUCAUUAAUUUAUUAUCA-------------UCCAGGGUGGUCACAUCAAGAUACACUUGAAUACCGAUAUGCCCGUCCAGGUGGACGGUGAGCCCUGGAUUCAGAG ......................(-------------(((((((((((....(((((.....)))))..))).......((((((....))))))...)))))))))...... ( -32.40) >DroVir_CAF1 38778 100 + 1 AU-AUAUAUUUAAUA-----ACCG-CUCG-----CUUACAGGGUGGCCACAUCAAAAUACAUUUAAACACCGAUAUGCCGGUGCAGGUGGACGGCGAGCCAUGGAUUCAGAG ..-............-----.(((-((((-----((.....((((....))))......(((((...(((((......))))).)))))...)))))))...))........ ( -25.40) >DroSec_CAF1 35632 104 + 1 ACCAUAUCAUUAAUUUAUGAACUUUAA--------CUCCAGGGUGGUCACAUCAAGAUACAUUUGAACACCGAUAUGCCCGUCCAGGUGGACGGUGAGCCCUGGAUUCAGAG ......((((......)))).((((.(--------.(((((((((((....(((((.....)))))..))).......((((((....))))))...)))))))).).)))) ( -31.30) >DroSim_CAF1 35688 112 + 1 ACCAUAUCAUUAAUUUAUGAACUUUAACUACUUAACUACAGGGUGGUCACAUCAAGAUACAUUUGAACACCGAUAUGCCCGUCCAGGUGGACGGUGAGCCCUGGAUCCAGAG ......((((......))))..................(((((((((....(((((.....)))))..))).......((((((....))))))...))))))......... ( -26.30) >DroEre_CAF1 36244 104 + 1 ACCUUAUUCUCAAUGUAUGUACUUUAA-------CUU-CAGGGUGGUCAUAUCAAGAUACACUUGAAUACCGAUAUGCCCGUCCAGGUGGACGGUGAGCCCUGGAUCCAGAG .....................((((..-------.((-(((((((((.((.(((((.....)))))))))).......((((((....))))))...))))))))...)))) ( -29.70) >DroYak_CAF1 36270 105 + 1 ACCAUGUAAUUAAUGUAUGAAUUUUAA-------AUUGCAGGGUGGUCACAUCAAGAUACACUUGAAUACCGAUAUGCCCGUCCAGGUGGACGGUGAGCCCUGGAUCCAGAG .(((((((((((((......)))...)-------))))))(((((((....(((((.....)))))..))).......((((((....))))))...)))))))........ ( -28.40) >consensus ACCAUAUCAUUAAUUUAUGAACUUUAA________UUACAGGGUGGUCACAUCAAGAUACACUUGAACACCGAUAUGCCCGUCCAGGUGGACGGUGAGCCCUGGAUCCAGAG ......................................(((((((((....(((((.....)))))..))).......((((((....))))))...))))))......... (-21.60 = -22.05 + 0.45)


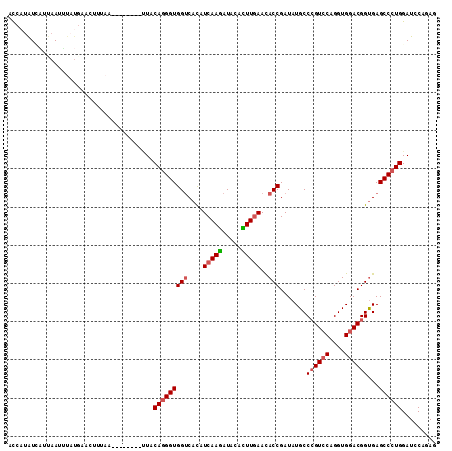
| Location | 19,805,208 – 19,805,307 |
|---|---|
| Length | 99 |
| Sequences | 6 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 81.87 |
| Mean single sequence MFE | -29.83 |
| Consensus MFE | -22.79 |
| Energy contribution | -22.90 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.742529 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19805208 99 - 27905053 CUCUGAAUCCAGGGCUCACCGUCCACCUGGACGGGCAUAUCGGUAUUCAAGUGUAUCUUGAUGUGACCACCCUGGA-------------UGAUAAUAAAUUAAUGAUGUGGU ......((((((((....((((((....)))))).......(((..(((((.....)))))....))).)))))))-------------)...................... ( -34.30) >DroVir_CAF1 38778 100 - 1 CUCUGAAUCCAUGGCUCGCCGUCCACCUGCACCGGCAUAUCGGUGUUUAAAUGUAUUUUGAUGUGGCCACCCUGUAAG-----CGAG-CGGU-----UAUUAAAUAUAU-AU ..........((.((((((.(.((((..(((((((....)))))))(((((.....))))).)))).)((...))..)-----))))-).))-----............-.. ( -23.00) >DroSec_CAF1 35632 104 - 1 CUCUGAAUCCAGGGCUCACCGUCCACCUGGACGGGCAUAUCGGUGUUCAAAUGUAUCUUGAUGUGACCACCCUGGAG--------UUAAAGUUCAUAAAUUAAUGAUAUGGU ((.(((.(((((((....((((((....)))))).((((((((.((........)).))))))))....))))))).--------))).)).((((......))))...... ( -33.10) >DroSim_CAF1 35688 112 - 1 CUCUGGAUCCAGGGCUCACCGUCCACCUGGACGGGCAUAUCGGUGUUCAAAUGUAUCUUGAUGUGACCACCCUGUAGUUAAGUAGUUAAAGUUCAUAAAUUAAUGAUAUGGU (((((....)))))....((((((....)))))).((((((((((.(((.((........)).))).)))).....(((((....(((.......))).))))))))))).. ( -29.40) >DroEre_CAF1 36244 104 - 1 CUCUGGAUCCAGGGCUCACCGUCCACCUGGACGGGCAUAUCGGUAUUCAAGUGUAUCUUGAUAUGACCACCCUG-AAG-------UUAAAGUACAUACAUUGAGAAUAAGGU .(((.(((.(((((....((((((....)))))).......(((..(((((.....)))))....))).)))))-..(-------(......))....))).)))....... ( -30.90) >DroYak_CAF1 36270 105 - 1 CUCUGGAUCCAGGGCUCACCGUCCACCUGGACGGGCAUAUCGGUAUUCAAGUGUAUCUUGAUGUGACCACCCUGCAAU-------UUAAAAUUCAUACAUUAAUUACAUGGU ...(((((.(((((....((((((....)))))).......(((..(((((.....)))))....))).)))))..))-------)))........................ ( -28.30) >consensus CUCUGAAUCCAGGGCUCACCGUCCACCUGGACGGGCAUAUCGGUAUUCAAAUGUAUCUUGAUGUGACCACCCUGGAA________UUAAAGUUCAUAAAUUAAUGAUAUGGU .........(((((.....(((((....)))))((((((((((.((........)).)))))))).)).)))))...................................... (-22.79 = -22.90 + 0.11)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:46:02 2006