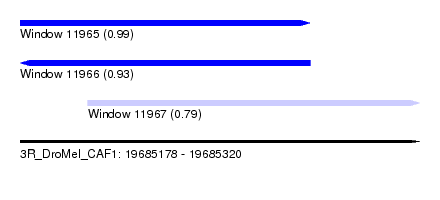
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 19,685,178 – 19,685,320 |
| Length | 142 |
| Max. P | 0.985697 |

| Location | 19,685,178 – 19,685,281 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.70 |
| Mean single sequence MFE | -29.15 |
| Consensus MFE | -28.30 |
| Energy contribution | -28.30 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.97 |
| SVM decision value | 2.02 |
| SVM RNA-class probability | 0.985697 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19685178 103 + 27905053 ---------------AUUA-UAAAUGUCGGGACGGCGCGAACGCCAUUCCCGGCUACCAAAAAUUACACGCACGAGUUAUUCACAUUAGGAAUAAUCGCAGAGGUCAACUCAGCCGAG-G ---------------....-.....(((((((.((((....))))..)))))))...............((.(((.((((((.......)))))))))..(((.....))).))....-. ( -28.30) >DroSec_CAF1 52403 103 + 1 ---------------AUUA-UAAAUGUCGGGACGGCGCGAACGCCAUUCCCGGCUACCAAAAAUUACACGCACGAGUUAUUCACAUUAGGAAUAAUCGCAGAGGUCAACUCAGCCGAG-G ---------------....-.....(((((((.((((....))))..)))))))...............((.(((.((((((.......)))))))))..(((.....))).))....-. ( -28.30) >DroWil_CAF1 98784 105 + 1 ---------------AUUAAUUAAGGUCGGGACGGCGCGAACGCCAUUCCCGGCUACCAAAAAUUACACGCACGAGUUAUUCACAUUAGGAAUAAUCGCAGAGGUCAACUCAGCCGAAUG ---------------.........((((((((.((((....))))..))))))))..............((.(((.((((((.......)))))))))..(((.....))).))...... ( -29.70) >DroYak_CAF1 53209 103 + 1 ---------------AUUA-UUAAUGUCGGGACGGCGCGAACGCCAUUCCCGGCUACCAAAAAUUACACGCACGAGUUAUUCACAUUAGGAAUAAUCGCAGAGGUCAACUCAGCCGAG-G ---------------....-.....(((((((.((((....))))..)))))))...............((.(((.((((((.......)))))))))..(((.....))).))....-. ( -28.30) >DroAna_CAF1 57272 103 + 1 ---------------AUUA-UUUAUGUCGGGACGGCGCGAACGCCAUUCCCGGCUACCAAAAAUUACACGCACGAGUUAUUCACAUUAGGAAUAAUCGCAGAGGUCAACUCAGCCGAG-G ---------------....-.....(((((((.((((....))))..)))))))...............((.(((.((((((.......)))))))))..(((.....))).))....-. ( -28.30) >DroPer_CAF1 75091 118 + 1 AAUGGUUCGAUAUUUAUUA-UUUAUGUCGGGACGGCGCGAACGCCAUUCCCGGCUACCAAAAAUUACACGCACGAGUUAUUCACAUUAGGAAUAAUCGCAGAGGUCAACUCAGCCGCA-G ..((((..((((.....))-))...(((((((.((((....))))..))))))).))))..........((..((.((((((.......))))))))((.(((.....))).)).)).-. ( -32.00) >consensus _______________AUUA_UUAAUGUCGGGACGGCGCGAACGCCAUUCCCGGCUACCAAAAAUUACACGCACGAGUUAUUCACAUUAGGAAUAAUCGCAGAGGUCAACUCAGCCGAG_G .........................(((((((.((((....))))..)))))))...............((.(((.((((((.......)))))))))..(((.....))).))...... (-28.30 = -28.30 + 0.00)



| Location | 19,685,178 – 19,685,281 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.70 |
| Mean single sequence MFE | -31.03 |
| Consensus MFE | -30.90 |
| Energy contribution | -30.90 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.03 |
| Structure conservation index | 1.00 |
| SVM decision value | 1.17 |
| SVM RNA-class probability | 0.925304 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19685178 103 - 27905053 C-CUCGGCUGAGUUGACCUCUGCGAUUAUUCCUAAUGUGAAUAACUCGUGCGUGUAAUUUUUGGUAGCCGGGAAUGGCGUUCGCGCCGUCCCGACAUUUA-UAAU--------------- .-...(((((((.....))).(((((((((((....).)))))).))))................))))((((.(((((....)))))))))........-....--------------- ( -30.90) >DroSec_CAF1 52403 103 - 1 C-CUCGGCUGAGUUGACCUCUGCGAUUAUUCCUAAUGUGAAUAACUCGUGCGUGUAAUUUUUGGUAGCCGGGAAUGGCGUUCGCGCCGUCCCGACAUUUA-UAAU--------------- .-...(((((((.....))).(((((((((((....).)))))).))))................))))((((.(((((....)))))))))........-....--------------- ( -30.90) >DroWil_CAF1 98784 105 - 1 CAUUCGGCUGAGUUGACCUCUGCGAUUAUUCCUAAUGUGAAUAACUCGUGCGUGUAAUUUUUGGUAGCCGGGAAUGGCGUUCGCGCCGUCCCGACCUUAAUUAAU--------------- ......((.(((.....))).(((((((((((....).)))))).))))))...(((((...(((...(((((.(((((....)))))))))))))..)))))..--------------- ( -31.30) >DroYak_CAF1 53209 103 - 1 C-CUCGGCUGAGUUGACCUCUGCGAUUAUUCCUAAUGUGAAUAACUCGUGCGUGUAAUUUUUGGUAGCCGGGAAUGGCGUUCGCGCCGUCCCGACAUUAA-UAAU--------------- .-...(((((((.....))).(((((((((((....).)))))).))))................))))((((.(((((....)))))))))........-....--------------- ( -30.90) >DroAna_CAF1 57272 103 - 1 C-CUCGGCUGAGUUGACCUCUGCGAUUAUUCCUAAUGUGAAUAACUCGUGCGUGUAAUUUUUGGUAGCCGGGAAUGGCGUUCGCGCCGUCCCGACAUAAA-UAAU--------------- .-...(((((((.....))).(((((((((((....).)))))).))))................))))((((.(((((....)))))))))........-....--------------- ( -30.90) >DroPer_CAF1 75091 118 - 1 C-UGCGGCUGAGUUGACCUCUGCGAUUAUUCCUAAUGUGAAUAACUCGUGCGUGUAAUUUUUGGUAGCCGGGAAUGGCGUUCGCGCCGUCCCGACAUAAA-UAAUAAAUAUCGAACCAUU .-.((.((..((...(((.(.(((((((((((....).)))))).))))).).))....))..)).))(((((.(((((....)))))))))).......-................... ( -31.30) >consensus C_CUCGGCUGAGUUGACCUCUGCGAUUAUUCCUAAUGUGAAUAACUCGUGCGUGUAAUUUUUGGUAGCCGGGAAUGGCGUUCGCGCCGUCCCGACAUUAA_UAAU_______________ .....(((((((.....))).(((((((((((....).)))))).))))................))))((((.(((((....)))))))))............................ (-30.90 = -30.90 + 0.00)



| Location | 19,685,202 – 19,685,320 |
|---|---|
| Length | 118 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.04 |
| Mean single sequence MFE | -33.38 |
| Consensus MFE | -29.50 |
| Energy contribution | -29.50 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.794634 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19685202 118 + 27905053 ACGCCAUUCCCGGCUACCAAAAAUUACACGCACGAGUUAUUCACAUUAGGAAUAAUCGCAGAGGUCAACUCAGCCGAG-GUGCAAUGGCCAAGCCUCACUCCGGAGGAAC-CGCCUUCGU ..(((((((((((((.................(((.((((((.......)))))))))..(((.....)))))))).)-)...))))))............((((((...-..)))))). ( -33.60) >DroPse_CAF1 73406 119 + 1 ACGCCAUUCCCGGCUACCAAAAAUUACACGCACGAGUUAUUCACAUUAGGAAUAAUCGCAGAGGUCAACUCAGCCGAA-GUGCAAUGGCCAAGCCUCACUCCGGAGGAACACGCCUUGGU ..(((......))).(((((.........((((((.((((((.......))))))))((.(((.....))).))....-))))...(((....((((......)))).....)))))))) ( -33.70) >DroWil_CAF1 98809 119 + 1 ACGCCAUUCCCGGCUACCAAAAAUUACACGCACGAGUUAUUCACAUUAGGAAUAAUCGCAGAGGUCAACUCAGCCGAAUGUGCAAUGGCCAAGCCUCACUCCGGAGGAAC-CGCCUUCGU ..(((((((..((((.................(((.((((((.......)))))))))..(((.....)))))))))))).))...(((....((((......))))...-.)))..... ( -32.10) >DroYak_CAF1 53233 118 + 1 ACGCCAUUCCCGGCUACCAAAAAUUACACGCACGAGUUAUUCACAUUAGGAAUAAUCGCAGAGGUCAACUCAGCCGAG-GUGCAAUGGCCAAGCCUCACUCCGGAGGAAC-CGCCUUCGU ..(((((((((((((.................(((.((((((.......)))))))))..(((.....)))))))).)-)...))))))............((((((...-..)))))). ( -33.60) >DroAna_CAF1 57296 118 + 1 ACGCCAUUCCCGGCUACCAAAAAUUACACGCACGAGUUAUUCACAUUAGGAAUAAUCGCAGAGGUCAACUCAGCCGAG-GUGCAAUGGCCAAGCCUCACUCCGGAGGAAC-CGCCUUCGU ..(((((((((((((.................(((.((((((.......)))))))))..(((.....)))))))).)-)...))))))............((((((...-..)))))). ( -33.60) >DroPer_CAF1 75130 119 + 1 ACGCCAUUCCCGGCUACCAAAAAUUACACGCACGAGUUAUUCACAUUAGGAAUAAUCGCAGAGGUCAACUCAGCCGCA-GUGCAAUGGCCAAGCCUCACUCCGGAGGAACACGCCUUGGU ..(((......))).(((((.........((((((.((((((.......))))))))((.(((.....))).))....-))))...(((....((((......)))).....)))))))) ( -33.70) >consensus ACGCCAUUCCCGGCUACCAAAAAUUACACGCACGAGUUAUUCACAUUAGGAAUAAUCGCAGAGGUCAACUCAGCCGAA_GUGCAAUGGCCAAGCCUCACUCCGGAGGAAC_CGCCUUCGU ..(((......)))...............((((((.((((((.......))))))))((.(((.....))).)).....))))...(((....((((......)))).....)))..... (-29.50 = -29.50 + 0.00)

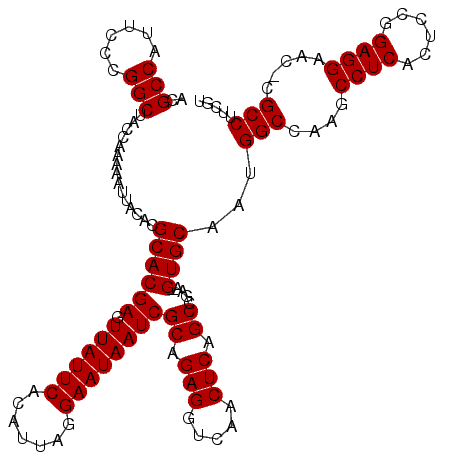
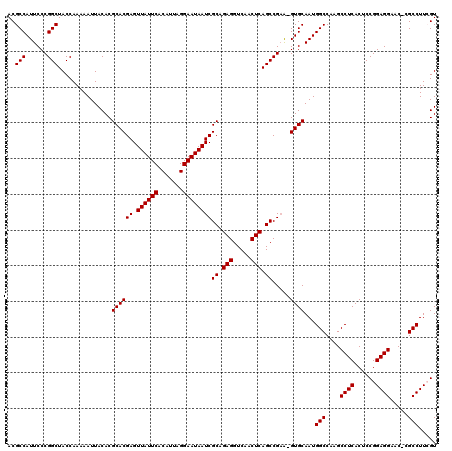
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:45:05 2006