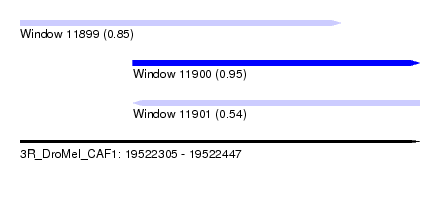

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 19,522,305 – 19,522,447 |
| Length | 142 |
| Max. P | 0.950007 |

| Location | 19,522,305 – 19,522,419 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 79.53 |
| Mean single sequence MFE | -31.52 |
| Consensus MFE | -20.80 |
| Energy contribution | -21.83 |
| Covariance contribution | 1.03 |
| Combinations/Pair | 1.41 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.79 |
| SVM RNA-class probability | 0.851548 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19522305 114 + 27905053 CAGUACAGCUUAUUCCACAUCCAUGGAGUUGUAAAGUUGUUUAUACCUAGAUCCCCGCCCGCUGCUUUGGAGCAGCAUAUCAUUAUGCUGUUCCCUAAGGGCCUGUUAAAAAAA (((.(((((((((((((......))))))....)))))))................((((........(((((((((((....)))))))))))....)))))))......... ( -35.10) >DroGri_CAF1 15949 110 + 1 CAGU---GUUUAUUCCAAGUCCAUGGAGUUGUAGAGCUGCUUAUAGCGGGAGCCACGUCCACUGGAUUGCAUAAGCAAAUUAUUUUGAUGCUCCCUCAGCCUCUGUUAAUGAA- ....---....((((((......))))))..(((((((((.....))((((((....(((...)))((((....))))...........))))))..)).)))))........- ( -26.60) >DroSim_CAF1 17275 114 + 1 CAGUACAGCUUAUUCCACAUCCAUGGAGUUGUAAAGUUGUUUAUUUUUUUAUCCCCGCCCGCUGCUUUGGAGCACCAUAUCAUUAUGCUGUUCCCUCAGGGCCUGUUAAAUAAA (((.(((((((((((((......))))))....)))))))................((((........((((((.((((....)))).))))))....)))))))......... ( -28.50) >DroEre_CAF1 17225 114 + 1 CAGUACAGCUUAUUCCACAUCCAUGGAGUUGUAAAGUUGCUUAUACCUAGAUCCCCGCCCGCUGCUUUGGAGCAGCAUAUCGUUAUGCCGCUCCCUCAGCGCCUGUGGAACAAA .....((((((((((((......))))))....))))))...............((((.(((((....(((((.(((((....))))).)))))..)))))...))))...... ( -36.10) >DroYak_CAF1 17308 114 + 1 CAGUACAGCUUAUUCCACAUCCAUGGAGUUGUACAGUUGCUUAUACCUAGAUCCUCGCCCGCUGCUCUGGAGCAGCAUAUCGUUAUGCCGUUCCCUCAGGGCCUGUAAAACCAA ..(((((((....((((......)))))))))))......................((((........(((((.(((((....))))).)))))....))))............ ( -30.50) >DroAna_CAF1 15545 111 + 1 --GUAUA-UUUACUCCACGUCCAUGGAAUUGUAUAGCUGCUUGUAUCGGGAUCCCUGUCCGCUGGUUUGGAGCAACAUGUCAUUGUGUUGCUCCCUCAGGGCCUGUAAAGAAAU --.((((-(....((((......))))...)))))....(((....((((..(((((.((...))...((((((((((......))))))))))..)))))))))..))).... ( -32.30) >consensus CAGUACAGCUUAUUCCACAUCCAUGGAGUUGUAAAGUUGCUUAUACCUAGAUCCCCGCCCGCUGCUUUGGAGCAGCAUAUCAUUAUGCUGCUCCCUCAGGGCCUGUUAAAAAAA .....((((((((((((......))))))....)))))).................((((........(((((((((((....)))))))))))....))))............ (-20.80 = -21.83 + 1.03)


| Location | 19,522,345 – 19,522,447 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 78.28 |
| Mean single sequence MFE | -24.75 |
| Consensus MFE | -18.45 |
| Energy contribution | -18.62 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.75 |
| SVM decision value | 1.40 |
| SVM RNA-class probability | 0.950007 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19522345 102 + 27905053 UUAUACCUAGAUCCCCGCCCGCUGCUUUGGAGCAGCAUAUCAUUAUGCUGUUCCCUAAGGGCCUGUUAAAAAAAAAAACACAUA--UAAAUAGUG----ACGGCUUAA ................(((.(((.(((.(((((((((((....)))))))))))..))))))................(((...--......)))----..))).... ( -27.30) >DroSec_CAF1 17126 98 + 1 UUAUACCUAGAUCCCCGCCCGCUGCUUUGGAGCAGCAUAUCAUUAUGCUGUUCCCUCAGGGCCUGUUAAAUAAAA----ACAUA--UAAAUAGAG----GAAGUUUAA .......(((((..((((((........(((((((((((....)))))))))))....))))(((((........----.....--..))))).)----)..))))). ( -28.46) >DroSim_CAF1 17315 98 + 1 UUAUUUUUUUAUCCCCGCCCGCUGCUUUGGAGCACCAUAUCAUUAUGCUGUUCCCUCAGGGCCUGUUAAAUAAAA----ACAUA--UAAAUAGAG----GAAGUUUAA ...(((((((((....((((........((((((.((((....)))).))))))....)))).((((.......)----)))..--...))))))----)))...... ( -20.80) >DroEre_CAF1 17265 99 + 1 UUAUACCUAGAUCCCCGCCCGCUGCUUUGGAGCAGCAUAUCGUUAUGCCGCUCCCUCAGCGCCUGUGGAACAAAA----GCAUA--UAAAUAGUU---AAUAGUUUCA ..............((((.(((((....(((((.(((((....))))).)))))..)))))...)))).......----.....--.........---.......... ( -27.10) >DroYak_CAF1 17348 99 + 1 UUAUACCUAGAUCCUCGCCCGCUGCUCUGGAGCAGCAUAUCGUUAUGCCGUUCCCUCAGGGCCUGUAAAACCAAA----ACAUA--UAAAUAGUU---CACAGUUUCA .........((..((.((((........(((((.(((((....))))).)))))....))))((((.........----.....--...))))..---...))..)). ( -20.03) >DroPer_CAF1 14987 101 + 1 UUAUAACGGGACCCACGACCACUGCUUUGGAGCAGCAUGUCAUUAUGUUGUUCUCUUAGGAUCUGCGGAUACGAA----AGAUAGUUAAGCAAAGGUU---UGUUUUA .......(((((....((((..(((((.(((((((((((....)))))))))))......((((.((....))..----))))....)))))..))))---.))))). ( -24.80) >consensus UUAUACCUAGAUCCCCGCCCGCUGCUUUGGAGCAGCAUAUCAUUAUGCUGUUCCCUCAGGGCCUGUUAAAAAAAA____ACAUA__UAAAUAGAG____AAAGUUUAA ................((((........(((((((((((....)))))))))))....)))).............................................. (-18.45 = -18.62 + 0.17)


| Location | 19,522,345 – 19,522,447 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 78.28 |
| Mean single sequence MFE | -27.38 |
| Consensus MFE | -14.28 |
| Energy contribution | -15.08 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.543440 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19522345 102 - 27905053 UUAAGCCGU----CACUAUUUA--UAUGUGUUUUUUUUUUUAACAGGCCCUUAGGGAACAGCAUAAUGAUAUGCUGCUCCAAAGCAGCGGGCGGGGAUCUAGGUAUAA ....(((..----(((......--...)))..............(((((((...(((.(((((((....))))))).)))...((.....)))))).))).))).... ( -25.20) >DroSec_CAF1 17126 98 - 1 UUAAACUUC----CUCUAUUUA--UAUGU----UUUUAUUUAACAGGCCCUGAGGGAACAGCAUAAUGAUAUGCUGCUCCAAAGCAGCGGGCGGGGAUCUAGGUAUAA .......((----(((......--..(((----(.......))))(.(((((..(((.(((((((....))))))).)))....))).)).))))))........... ( -28.80) >DroSim_CAF1 17315 98 - 1 UUAAACUUC----CUCUAUUUA--UAUGU----UUUUAUUUAACAGGCCCUGAGGGAACAGCAUAAUGAUAUGGUGCUCCAAAGCAGCGGGCGGGGAUAAAAAAAUAA .......((----(((......--..(((----(.......))))(.(((((..(((.((.((((....)))).)).)))....))).)).))))))........... ( -24.30) >DroEre_CAF1 17265 99 - 1 UGAAACUAUU---AACUAUUUA--UAUGC----UUUUGUUCCACAGGCGCUGAGGGAGCGGCAUAACGAUAUGCUGCUCCAAAGCAGCGGGCGGGGAUCUAGGUAUAA ..........---.........--(((((----((..(((((.....(((((..(((((((((((....)))))))))))....)))))....)))))..))))))). ( -39.80) >DroYak_CAF1 17348 99 - 1 UGAAACUGUG---AACUAUUUA--UAUGU----UUUGGUUUUACAGGCCCUGAGGGAACGGCAUAACGAUAUGCUGCUCCAGAGCAGCGGGCGAGGAUCUAGGUAUAA .....(((((---(((((....--.....----..))).)))))))((((((..(((.(((((((....))))))).)))....))..))))................ ( -27.80) >DroPer_CAF1 14987 101 - 1 UAAAACA---AACCUUUGCUUAACUAUCU----UUCGUAUCCGCAGAUCCUAAGAGAACAACAUAAUGACAUGCUGCUCCAAAGCAGUGGUCGUGGGUCCCGUUAUAA ...(((.---.((((...((((...((((----..((....)).))))..))))...........(((((.(((((((....))))))))))))))))...))).... ( -18.40) >consensus UUAAACUGU____CACUAUUUA__UAUGU____UUUUAUUUAACAGGCCCUGAGGGAACAGCAUAAUGAUAUGCUGCUCCAAAGCAGCGGGCGGGGAUCUAGGUAUAA .............................................(.(((((..(((.(((((((....))))))).)))....))).)).)................ (-14.28 = -15.08 + 0.81)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:44:01 2006