
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 19,420,168 – 19,420,326 |
| Length | 158 |
| Max. P | 0.952981 |

| Location | 19,420,168 – 19,420,286 |
|---|---|
| Length | 118 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.21 |
| Mean single sequence MFE | -39.89 |
| Consensus MFE | -28.82 |
| Energy contribution | -28.02 |
| Covariance contribution | -0.80 |
| Combinations/Pair | 1.44 |
| Mean z-score | -2.72 |
| Structure conservation index | 0.72 |
| SVM decision value | 1.43 |
| SVM RNA-class probability | 0.952981 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19420168 118 - 27905053 UACAACUGGGGCAAGAAGGCAGCUUCUGGAAUCGCAGUACUUGUGGUUUUGCUAUCUGGCUGCCUUUUUGCUACCCAGAUGGUCAACCACUACACGUUGCUUAUUAAGUAGGGUUAC--U ..((.(((((((((((((((((((..(..((((((((...))))))))..)......))))))))))))))..))))).))((.((((.((((..............))))))))))--. ( -46.34) >DroSim_CAF1 28220 118 - 1 UACAAGUGGGGCAAGAAGGCAGCUGCUGGAAUCGCAGUACUUGUGGUUUUGCUAUCUGGCUGCCUUUUUGCCACCCAGAUGGUCAACCACUACUCGUUGCUUAUUAAGUAGGUUUAC--U ....((((((((((((((((((((((.((((((((((...)))))))))))).....)))))))))))))))(((.....)))...)))))....((.((((((...))))))..))--. ( -45.90) >DroEre_CAF1 20323 117 - 1 UACAAAUGGGGCAAGAAGGCGGCUGCUGGAAUUGCAGUUCUCGUGGUUUUGCUAUCCGGCUGUCUUUUUGCUACCCAGAUGGUCAACCACUACUCGAUGCUUAUUAAGUAAGA-UAC--U ......((((((((((((((((((...(((...((((..(.....)..))))..)))))))))))))))))..))))..(((....))).((((..((....))..))))...-...--. ( -37.00) >DroYak_CAF1 28219 118 - 1 UACAAAUGGGGCAAGAAGGCGGCUGCUGGAAUCGCAGUACUUGUGGUUUUGCUAUCCGGCUGCCUUUUUGCCACCCAGAUGGUCAACCACUACUCGCUGCUUAUUAAGUAAGGUUAC--U .......((((((((((((((((((.(..((((((((...))))))))..).....)))))))))))))))).)).((.(((....))))).((..((........))..)).....--. ( -42.40) >DroAna_CAF1 31718 117 - 1 UACAAAUGGGGCAAAAAGGCUGCUGCUGGAAUAGUGGUAUUAAUAGUGUUGCUCUCUGGCUGCCUUUUUGCCACACAAAUGGUCAACAAAUAUUCAAUACUGUUUAAGUAUGU-UUC--U ..((..((.(((((((((((....((..((..(((..(((.....)))..))).))..)).)))))))))))...))..))...............(((((.....)))))..-...--. ( -34.50) >DroPer_CAF1 29777 120 - 1 UAUAAAUGGGGCAAAAAGGCAGCUGCUGGCAUUGUAGUGCUGUUGCUUCUAGUCACGACCUGCCUCUUUGUCACGAUGAUGGUCAACAAUGAAUCAGUGCUCACGAAGUAGGUCCACAAU .......(((((((...((((.((((.......)))))))).))))))).......(((((((.((..((.((((((..((.....))....))).)))..)).)).)))))))...... ( -33.20) >consensus UACAAAUGGGGCAAGAAGGCAGCUGCUGGAAUCGCAGUACUUGUGGUUUUGCUAUCUGGCUGCCUUUUUGCCACCCAGAUGGUCAACCACUACUCGAUGCUUAUUAAGUAGGUUUAC__U ..((..((((((((((((((((((((((......))))....((((.....))))..)))))))))))))))..)))..))................((((.....)))).......... (-28.82 = -28.02 + -0.80)

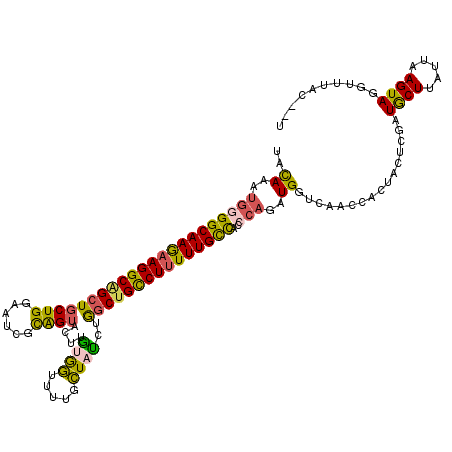

| Location | 19,420,206 – 19,420,326 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.11 |
| Mean single sequence MFE | -37.95 |
| Consensus MFE | -27.47 |
| Energy contribution | -26.28 |
| Covariance contribution | -1.19 |
| Combinations/Pair | 1.52 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.89 |
| SVM RNA-class probability | 0.874698 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19420206 120 - 27905053 GCAGCUAUUUAUCAUCUCCCCGAUCCUCUUGAUGGCUCUGUACAACUGGGGCAAGAAGGCAGCUUCUGGAAUCGCAGUACUUGUGGUUUUGCUAUCUGGCUGCCUUUUUGCUACCCAGAU ..(((((((....(((.....)))......)))))))........(((((((((((((((((((..(..((((((((...))))))))..)......))))))))))))))..))))).. ( -44.10) >DroSim_CAF1 28258 120 - 1 GCAGCUAUUUAUCAUCUCCCCGAUCCUCCUGAUCGCUCUGUACAAGUGGGGCAAGAAGGCAGCUGCUGGAAUCGCAGUACUUGUGGUUUUGCUAUCUGGCUGCCUUUUUGCCACCCAGAU ....................(((((.....))))).((((.......(((((((((((((((((((.((((((((((...)))))))))))).....))))))))))))))).)))))). ( -42.71) >DroEre_CAF1 20360 120 - 1 GCAGCUCUUUAUCAUCUCUCCAAUCCUCCUGAUCGCUCUUUACAAAUGGGGCAAGAAGGCGGCUGCUGGAAUUGCAGUUCUCGUGGUUUUGCUAUCCGGCUGUCUUUUUGCUACCCAGAU ..............................................((((((((((((((((((...(((...((((..(.....)..))))..)))))))))))))))))..))))... ( -32.80) >DroYak_CAF1 28257 120 - 1 GCAGCUAUUCAUCAUCUCUCCAAUCCUCCUGAUGGCUCUUUACAAAUGGGGCAAGAAGGCGGCUGCUGGAAUCGCAGUACUUGUGGUUUUGCUAUCCGGCUGCCUUUUUGCCACCCAGAU ..(((((((.....................)))))))..........((((((((((((((((((.(..((((((((...))))))))..).....)))))))))))))))).))..... ( -40.00) >DroAna_CAF1 31755 120 - 1 GCAGUUGUACUUCAUCUCACCUUUUCUCCUGAUAGCCAUUUACAAAUGGGGCAAAAAGGCUGCUGCUGGAAUAGUGGUAUUAAUAGUGUUGCUCUCUGGCUGCCUUUUUGCCACACAAAU ....((((...........................(((((....)))))(((((((((((....((..((..(((..(((.....)))..))).))..)).)))))))))))..)))).. ( -35.70) >DroPer_CAF1 29817 120 - 1 GCAACUAUACAUAAUCUCUCCGCUUCUGCUGAUUGCUCUGUAUAAAUGGGGCAAAAAGGCAGCUGCUGGCAUUGUAGUGCUGUUGCUUCUAGUCACGACCUGCCUCUUUGUCACGAUGAU (((((..((((((((....((((..(((((..(((((((((....)))))))))...)))))..)).)).))))).)))..))))).((..((.((((.........)))).))...)). ( -32.40) >consensus GCAGCUAUUCAUCAUCUCUCCGAUCCUCCUGAUCGCUCUGUACAAAUGGGGCAAGAAGGCAGCUGCUGGAAUCGCAGUACUUGUGGUUUUGCUAUCUGGCUGCCUUUUUGCCACCCAGAU .................................................(((((((((((((((((((......))))....((((.....))))..)))))))))))))))........ (-27.47 = -26.28 + -1.19)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:43:33 2006