
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 19,364,802 – 19,364,898 |
| Length | 96 |
| Max. P | 0.856816 |

| Location | 19,364,802 – 19,364,898 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 77.05 |
| Mean single sequence MFE | -19.34 |
| Consensus MFE | -4.16 |
| Energy contribution | -5.64 |
| Covariance contribution | 1.48 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.22 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.588137 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19364802 96 + 27905053 UUUUGUUUUCUAAAUUUUAACCAGAUGGCGAUACACCAAAAUUUUUUGUUUGUACCUUG-ACUUCAUAUUUUUUAUGGAUCUUAGACAUAAUUAUAG ..((((..(((...........)))..))))........((((...((((((......(-(.((((((.....)))))))).)))))).)))).... ( -9.40) >DroSec_CAF1 14673 96 + 1 UUUUGUUUUCUAAAUUUUAACCAGAGGGCGGUACACCCACAUU-UUGGUUUGUACCGUGUAUUUCCUAUUUUUUAUAGAUCUUUGACAUAAUUAUAG .....................((((((((((((((.(((....-.)))..)))))))).......((((.....)))).))))))............ ( -19.70) >DroSim_CAF1 19993 96 + 1 UUUUGUUUUCUAAAUUUUAACCAGAGGGCGGUACAUCCAAAUU-UUGGUUUGUACCGUGUAUUUCAUUUUUUUUAUGGAUCUUAGACAUAAUUAUAG ....(((((((...........)))))))((((((.(((....-.)))..))))))((.((.(((((.......)))))...)).)).......... ( -19.30) >DroEre_CAF1 18136 84 + 1 UUUUGUUUUUUAAAUUUUAGCCAGAGGGCGGCACAUCUACACU-UU--UGUGUUCGGUGUUUUCCAUACUU-U---------UGGCCACAAUUAUAG ...................((((((((..((.(((((.((((.-..--.))))..)))))...))...)))-)---------))))........... ( -20.70) >DroYak_CAF1 18183 95 + 1 UUAUGAUUUCCAAAUUGCAAACAGAGGGCGACACAUCUUAAUU-UUGGUGUGUACCGUGCUUUCCAUACUU-UUAUGGCUCUUAGCCGCAAUUAUGG .........(((((((((.....((((((((((((((......-..)))))))....))))))).......-....(((.....))))))))).))) ( -27.60) >consensus UUUUGUUUUCUAAAUUUUAACCAGAGGGCGGUACAUCCAAAUU_UUGGUUUGUACCGUGUAUUUCAUAUUUUUUAUGGAUCUUAGACAUAAUUAUAG ...........................((((((((.(((......)))..))))))))....................................... ( -4.16 = -5.64 + 1.48)

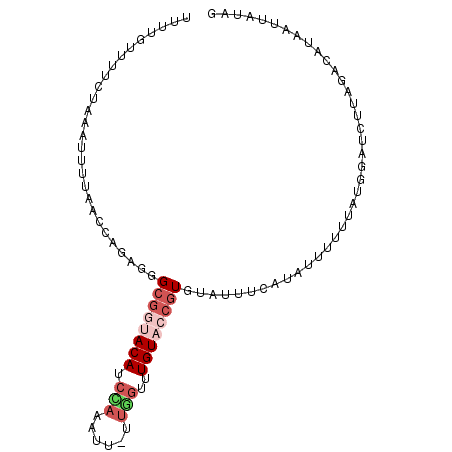

| Location | 19,364,802 – 19,364,898 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 77.05 |
| Mean single sequence MFE | -17.18 |
| Consensus MFE | -3.76 |
| Energy contribution | -4.36 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.22 |
| SVM decision value | 0.81 |
| SVM RNA-class probability | 0.856816 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19364802 96 - 27905053 CUAUAAUUAUGUCUAAGAUCCAUAAAAAAUAUGAAGU-CAAGGUACAAACAAAAAAUUUUGGUGUAUCGCCAUCUGGUUAAAAUUUAGAAAACAAAA ...........((((((((.((((.....))))..))-)..((((((..((((....)))).))))))(((....)))......)))))........ ( -12.90) >DroSec_CAF1 14673 96 - 1 CUAUAAUUAUGUCAAAGAUCUAUAAAAAAUAGGAAAUACACGGUACAAACCAA-AAUGUGGGUGUACCGCCCUCUGGUUAAAAUUUAGAAAACAAAA (((.((((.((....((((((((.....))))).......(((((((..(((.-....))).)))))))...)))...)).)))))))......... ( -15.10) >DroSim_CAF1 19993 96 - 1 CUAUAAUUAUGUCUAAGAUCCAUAAAAAAAAUGAAAUACACGGUACAAACCAA-AAUUUGGAUGUACCGCCCUCUGGUUAAAAUUUAGAAAACAAAA ...........((((((........................((((((..(((.-....))).))))))(((....))).....))))))........ ( -14.70) >DroEre_CAF1 18136 84 - 1 CUAUAAUUGUGGCCA---------A-AAGUAUGGAAAACACCGAACACA--AA-AGUGUAGAUGUGCCGCCCUCUGGCUAAAAUUUAAAAAACAAAA ....((((.((((((---------.-.((..(((...(((.(..((((.--..-.)))).).))).)))..)).)))))).))))............ ( -17.90) >DroYak_CAF1 18183 95 - 1 CCAUAAUUGCGGCUAAGAGCCAUAA-AAGUAUGGAAAGCACGGUACACACCAA-AAUUAAGAUGUGUCGCCCUCUGUUUGCAAUUUGGAAAUCAUAA (((.((((((((...(((((((((.-...))))).......((((((((.(..-......).))))).)))))))..)))))))))))......... ( -25.30) >consensus CUAUAAUUAUGUCUAAGAUCCAUAAAAAAUAUGAAAUACACGGUACAAACCAA_AAUUUAGAUGUACCGCCCUCUGGUUAAAAUUUAGAAAACAAAA (((.((((.................................((((((..(((......))).))))))(((....)))...)))))))......... ( -3.76 = -4.36 + 0.60)


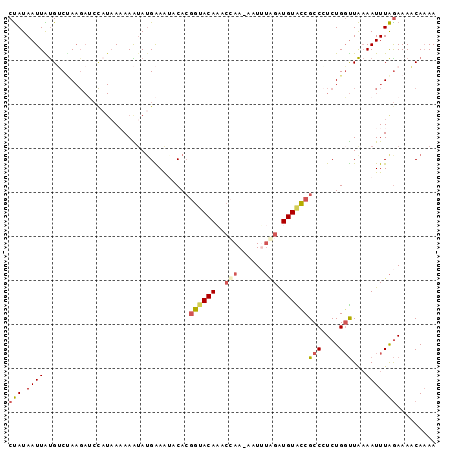
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:43:08 2006