
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 19,291,538 – 19,291,663 |
| Length | 125 |
| Max. P | 0.717965 |

| Location | 19,291,538 – 19,291,636 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 72.56 |
| Mean single sequence MFE | -26.23 |
| Consensus MFE | -7.22 |
| Energy contribution | -8.20 |
| Covariance contribution | 0.98 |
| Combinations/Pair | 1.29 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.28 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.559275 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19291538 98 + 27905053 AAGUGCGUAACAAGAAUUGAGGCAG--------UUGCGGUCUAUAAUGGAUUUAUGGGGUUAGCGCGAAGGCUGCCCGGUAGAUAGACGGUUAUAGACAGCUAGAU .(((.((((((.............)--------)))))(((((((((.(((((((.((((.(((......))))))).))))))...).))))))))).))).... ( -32.62) >DroGri_CAF1 100818 94 + 1 AAGUGCGUAGCAAGAAUUGAGCAAAGCGAAACUUUGCUUAUAAAAAUAAGUUUAUGAGGUUACACUGAAGGUUGCA-GCUAAAGUG-----------CAUUUAAAU (((((((((((.......((((((((.....))))))))(((((......)))))...)))))(((...(((....-)))..))))-----------))))).... ( -21.40) >DroSec_CAF1 87355 98 + 1 AAGUGCGUAACAAGAAUUGAGGCAG--------UUGCGGGCUAUAAUGGAUUUAUGGGGUUAGCGCGAAGGCUGCCCGGUAGAUAGACGGUUAUAGACAGCUAGAU .(((.((((((.............)--------)))))..(((((((.(((((((.((((.(((......))))))).))))))...).)))))))...))).... ( -27.62) >DroSim_CAF1 89273 98 + 1 AAGUGCGUAACAAGAAUUGAGGCAG--------UUGCGGGCUAUAAUGGAUUUAUGGGGUUAGCGCGAAGGCUGCCCGGUAGAUAGACGGUUAUAGACAGCUAGAU .(((.((((((.............)--------)))))..(((((((.(((((((.((((.(((......))))))).))))))...).)))))))...))).... ( -27.62) >DroEre_CAF1 88527 98 + 1 AAGUGCGUAACAAGAAUUGAGGCAG--------UUGCGGGCUAUAAUGGAUUUAUGGGGUUACCGCGAAGGCUGCCCGGUAGAUAGACGCUUAUAGACAGAUAGAU ....((((.......((((.(((((--------((((((.((((((.....)))))).....)))))...))))))))))......))))................ ( -27.12) >DroMoj_CAF1 104792 76 + 1 AAGUGCGUAGCAAGAGCUGAGCU------------------AAAAGUUAGCUUAUGAGGUUAGCCCGAAGGCCGCA-GCUAAAUUG-----------CGUUUAAAU .........((((.(((((.(((------------------....(((((((.....))))))).....)))..))-)))...)))-----------)........ ( -21.00) >consensus AAGUGCGUAACAAGAAUUGAGGCAG________UUGCGGGCUAUAAUGGAUUUAUGGGGUUAGCGCGAAGGCUGCCCGGUAGAUAGACGGUUAUAGACAGCUAGAU ..((.....))......................................((((((.((((....((....)).)))).))))))...................... ( -7.22 = -8.20 + 0.98)



| Location | 19,291,563 – 19,291,663 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 75.23 |
| Mean single sequence MFE | -25.73 |
| Consensus MFE | -12.64 |
| Energy contribution | -11.08 |
| Covariance contribution | -1.55 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.49 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.717965 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19291563 100 + 27905053 UUGCGGUCUAUAAUGGAUUUAUGGGGUUAGCGCGAAGGCUGCCCGGUAGAUAGACGGUUAUAGAC--AGCUAGAUGGAUAUGUAUAUAGAUAAAUGUCUAUG ..((.(((((((((.(((((((.((((.(((......))))))).))))))...).)))))))))--.))...((((((((.(((....))).)))))))). ( -35.30) >DroPse_CAF1 103956 90 + 1 AU------AAUAAUAAAUUUAUGGGGUUAGCACGAAGGUUGUUCGAUGAAUGGUCU---AUGCCUGGAGCU--CUGGGCACACAUAUAG-CAUAUGCUCAUA ..------...........((((((..((((....(((..((((...))))..)))---.((((..(....--)..))))........)-).))..)))))) ( -20.50) >DroSec_CAF1 87380 100 + 1 UUGCGGGCUAUAAUGGAUUUAUGGGGUUAGCGCGAAGGCUGCCCGGUAGAUAGACGGUUAUAGAC--AGCUAGAUGGAUAUGUAUAUAGAUAAAUGGCUAUG ....(..(((((((.(((((((.((((.(((......))))))).))))))...).))))))).)--(((((..((..(((....)))..))..)))))... ( -27.00) >DroSim_CAF1 89298 100 + 1 UUGCGGGCUAUAAUGGAUUUAUGGGGUUAGCGCGAAGGCUGCCCGGUAGAUAGACGGUUAUAGAC--AGCUAGAUGGAUAUGUAUAUAGAUAAAUGGCUAUG ....(..(((((((.(((((((.((((.(((......))))))).))))))...).))))))).)--(((((..((..(((....)))..))..)))))... ( -27.00) >DroEre_CAF1 88552 100 + 1 UUGCGGGCUAUAAUGGAUUUAUGGGGUUACCGCGAAGGCUGCCCGGUAGAUAGACGCUUAUAGAC--AGAUAGAUGGAUAUGUAUAUAGAUAAAUGGCUAUG .....((((((.....((((((.((((..((.....))..)))).)))))).......((((..(--(......))..))))...........))))))... ( -24.10) >DroPer_CAF1 104073 90 + 1 AU------AAUAAUAAAUUUAUGGGGUUAGCACGAAGGUUGUUCUAUGAAUGGUCU---AUGCCUGGAGCU--CUGGGCACAUAUAUAG-CAUAUGCUCAUA ..------...........((((((..((((....(((..((((...))))..)))---.((((..(....--)..))))........)-).))..)))))) ( -20.50) >consensus UUGCGGGCUAUAAUGGAUUUAUGGGGUUAGCGCGAAGGCUGCCCGGUAGAUAGACGGUUAUAGAC__AGCUAGAUGGAUAUGUAUAUAGAUAAAUGGCUAUG ................((((((.((((....((....)).)))).))))))................................................... (-12.64 = -11.08 + -1.55)

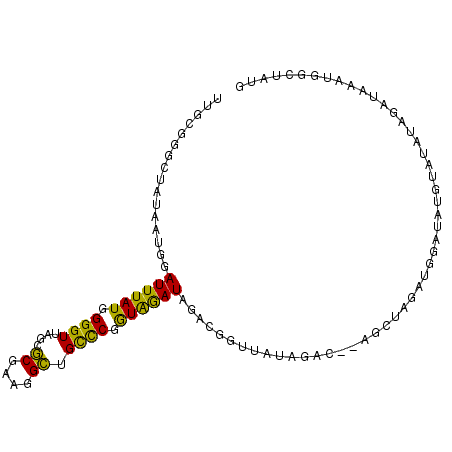

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:42:41 2006