
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 19,048,002 – 19,048,141 |
| Length | 139 |
| Max. P | 0.890260 |

| Location | 19,048,002 – 19,048,103 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 74.37 |
| Mean single sequence MFE | -22.04 |
| Consensus MFE | -11.22 |
| Energy contribution | -12.66 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.51 |
| SVM decision value | 0.00 |
| SVM RNA-class probability | 0.533890 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19048002 101 + 27905053 ---------GACAUGGU----GGCUAUUGCAAUUUUAUGUAGAUAUGGUGUAAGCUUAUAAGCUUUUGACACGUGUCCAUGCAAAUUUUUCAGUUAAUUUGCAUUAGUCAUUUU ---------.....(((----(((((.((((......))))((((((((..(((((....)))))...)).)))))).(((((((((........))))))))))))))))).. ( -25.60) >DroSec_CAF1 60762 99 + 1 ---------GACAUAGU----GGCGAUAGCUAUUUUAUGUAGAUAUGGUGUAAGCUUAUAAGCUUUUGACACGUGUCCAUGCAAAUUUUUCAAUUAAUUUGCAU-AGUAAUU-U ---------.(((((((----(((....)))))..))))).((((((((..(((((....)))))...)).)))))).(((((((((........)))))))))-.......-. ( -26.70) >DroSim_CAF1 62492 99 + 1 ---------GACAUAGU----GGCGAUUGCAAUUUUAUGUAGAUAUGAUGUAAGCUUAUAAGCUUUUGACACGUGUCCAUGCAAAUUUUGCAAUUAAUUUGCAU-AGUAAUU-U ---------.((((((.----.((....))....)))))).((((((.((((((((....)))))...))))))))).(((((((((........)))))))))-.......-. ( -23.40) >DroEre_CAF1 63778 95 + 1 ----UAUUAGAAAUAUUAAGCCGCCUAUA-AAUUUUAUGUAGAUACGGUGUAAGUUUACACGCUUUGUACACGUGUUCA------------AAUUAAUUAGCAU-AGUAAUU-G ----(((((.....(((((((((.(((((-.......)))))...))))....(...(((((.........))))).).------------..))))).....)-))))...-. ( -12.70) >DroYak_CAF1 64293 112 + 1 UGCAUAUUAUAAAUAUUAAGCCGCGCAGGCUACUUUAUGUAGAUAUAGUGUAAGUUUACAUGCUUUGCACACGUGUCCAUGCAAAUUUUUCAAUUCAUUUGCAU-ACUAACU-U ((((((............((((.....))))....))))))(((((.((((((((......)).))))))..))))).((((((((..........))))))))-.......-. ( -21.79) >consensus _________GACAUAGU____GGCGAUUGCAAUUUUAUGUAGAUAUGGUGUAAGCUUAUAAGCUUUUGACACGUGUCCAUGCAAAUUUUUCAAUUAAUUUGCAU_AGUAAUU_U .........................................((((((.(((((((......))))...))))))))).(((((((((........))))))))).......... (-11.22 = -12.66 + 1.44)

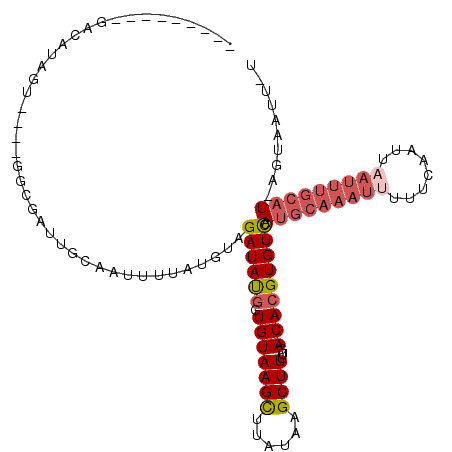

| Location | 19,048,002 – 19,048,103 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 74.37 |
| Mean single sequence MFE | -18.20 |
| Consensus MFE | -6.48 |
| Energy contribution | -8.52 |
| Covariance contribution | 2.04 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.36 |
| SVM decision value | 0.96 |
| SVM RNA-class probability | 0.890260 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19048002 101 - 27905053 AAAAUGACUAAUGCAAAUUAACUGAAAAAUUUGCAUGGACACGUGUCAAAAGCUUAUAAGCUUACACCAUAUCUACAUAAAAUUGCAAUAGCC----ACCAUGUC--------- .....(((....(((((((........)))))))((((....((((...(((((....))))))))).................((....)).----.)))))))--------- ( -17.80) >DroSec_CAF1 60762 99 - 1 A-AAUUACU-AUGCAAAUUAAUUGAAAAAUUUGCAUGGACACGUGUCAAAAGCUUAUAAGCUUACACCAUAUCUACAUAAAAUAGCUAUCGCC----ACUAUGUC--------- .-.....((-(((((((((........)))))))))))....((((...(((((....))))))))).......(((((.....((....)).----..))))).--------- ( -20.40) >DroSim_CAF1 62492 99 - 1 A-AAUUACU-AUGCAAAUUAAUUGCAAAAUUUGCAUGGACACGUGUCAAAAGCUUAUAAGCUUACAUCAUAUCUACAUAAAAUUGCAAUCGCC----ACUAUGUC--------- .-.....((-(((((((((........)))))))))))....(((....(((((....)))))....)))....(((((.....((....)).----..))))).--------- ( -18.80) >DroEre_CAF1 63778 95 - 1 C-AAUUACU-AUGCUAAUUAAUU------------UGAACACGUGUACAAAGCGUGUAAACUUACACCGUAUCUACAUAAAAUU-UAUAGGCGGCUUAAUAUUUCUAAUA---- .-.((((..-.((((.((.((((------------(..((((((.......))))))...........((....))...)))))-.)).))))...))))..........---- ( -11.60) >DroYak_CAF1 64293 112 - 1 A-AGUUAGU-AUGCAAAUGAAUUGAAAAAUUUGCAUGGACACGUGUGCAAAGCAUGUAAACUUACACUAUAUCUACAUAAAGUAGCCUGCGCGGCUUAAUAUUUAUAAUAUGCA (-((((..(-((((....(((((....)))))(((((....))))).....)))))..))))).............(((((..((((.....)))).....)))))........ ( -22.40) >consensus A_AAUUACU_AUGCAAAUUAAUUGAAAAAUUUGCAUGGACACGUGUCAAAAGCUUAUAAGCUUACACCAUAUCUACAUAAAAUUGCAAUCGCC____ACUAUGUC_________ ..........(((((((((........)))))))))......((((...((((......))))))))............................................... ( -6.48 = -8.52 + 2.04)



| Location | 19,048,023 – 19,048,141 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.77 |
| Mean single sequence MFE | -24.47 |
| Consensus MFE | -16.43 |
| Energy contribution | -16.81 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.811765 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19048023 118 + 27905053 UUAUGUAGAUAUGGUGUAAGCUUAUAAGCUUUUGACACGUGUCCAUGCAAAUUUUUCAGUUAAUUUGCAUUAGUCAUUUUUUUAACUAGUUCUACAUAACAAAG--UUGAAAUUUGUAUU .((((((((....(((((((((....)))))...))))(((.(.(((((((((........)))))))))..).))).............))))))))(((((.--......)))))... ( -26.20) >DroSec_CAF1 60783 118 + 1 UUAUGUAGAUAUGGUGUAAGCUUAUAAGCUUUUGACACGUGUCCAUGCAAAUUUUUCAAUUAAUUUGCAU-AGUAAUU-UGUUAACUAGUUGUACAUAACAAACUUAUGAAAUUGGUAUU .......((((((((..(((((....)))))...)).)))))).(((((((((........)))))))))-.......-.....(((((((...(((((.....))))).)))))))... ( -25.10) >DroSim_CAF1 62513 113 + 1 UUAUGUAGAUAUGAUGUAAGCUUAUAAGCUUUUGACACGUGUCCAUGCAAAUUUUGCAAUUAAUUUGCAU-AGUAAUU-UGUUAACUAGUUGUAGUU-----ACAUUUGAAAUUAGUAUU .......((((((.((((((((....)))))...)))))))))...(((((((.(((((.....))))).-...))))-)))..(((((((.(((..-----....))).)))))))... ( -23.80) >DroYak_CAF1 64327 117 + 1 UUAUGUAGAUAUAGUGUAAGUUUACAUGCUUUGCACACGUGUCCAUGCAAAUUUUUCAAUUCAUUUGCAU-ACUAACU-UUGCAACUAGUUGUAAAUUACAAACACUUUUAGCU-GUUUA .....(((....(((((.((((((((.((((((((...((....((((((((..........))))))))-....)).-.)))))..)))))))))))....))))).....))-).... ( -22.80) >consensus UUAUGUAGAUAUGGUGUAAGCUUAUAAGCUUUUGACACGUGUCCAUGCAAAUUUUUCAAUUAAUUUGCAU_AGUAAUU_UGUUAACUAGUUGUACAUAACAAACAUUUGAAAUU_GUAUU .......((((((.(((((((......))))...))))))))).(((((((((........)))))))))..............(((((((...................)))))))... (-16.43 = -16.81 + 0.38)


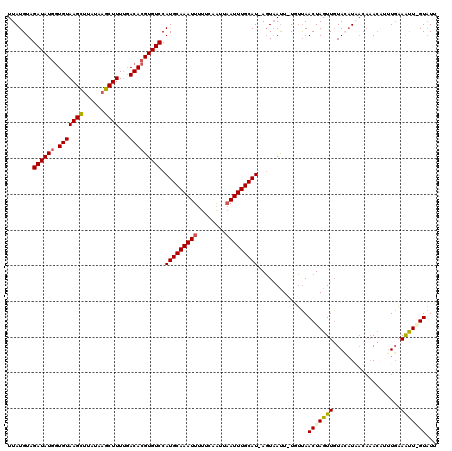
| Location | 19,048,023 – 19,048,141 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.77 |
| Mean single sequence MFE | -24.02 |
| Consensus MFE | -11.94 |
| Energy contribution | -12.50 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.05 |
| Mean z-score | -3.52 |
| Structure conservation index | 0.50 |
| SVM decision value | 0.86 |
| SVM RNA-class probability | 0.867499 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 19048023 118 - 27905053 AAUACAAAUUUCAA--CUUUGUUAUGUAGAACUAGUUAAAAAAAUGACUAAUGCAAAUUAACUGAAAAAUUUGCAUGGACACGUGUCAAAAGCUUAUAAGCUUACACCAUAUCUACAUAA ...(((((......--.)))))((((((((..((((((......))))))(((((((((........)))))))))......((((...(((((....)))))))))....)))))))). ( -26.60) >DroSec_CAF1 60783 118 - 1 AAUACCAAUUUCAUAAGUUUGUUAUGUACAACUAGUUAACA-AAUUACU-AUGCAAAUUAAUUGAAAAAUUUGCAUGGACACGUGUCAAAAGCUUAUAAGCUUACACCAUAUCUACAUAA ...............(((((((((............)))))-)))).((-(((((((((........)))))))))))....((((...(((((....)))))))))............. ( -23.90) >DroSim_CAF1 62513 113 - 1 AAUACUAAUUUCAAAUGU-----AACUACAACUAGUUAACA-AAUUACU-AUGCAAAUUAAUUGCAAAAUUUGCAUGGACACGUGUCAAAAGCUUAUAAGCUUACAUCAUAUCUACAUAA .((((..........(((-----(((((....))))).)))-.....((-(((((((((........)))))))))))....))))...(((((....)))))................. ( -20.70) >DroYak_CAF1 64327 117 - 1 UAAAC-AGCUAAAAGUGUUUGUAAUUUACAACUAGUUGCAA-AGUUAGU-AUGCAAAUGAAUUGAAAAAUUUGCAUGGACACGUGUGCAAAGCAUGUAAACUUACACUAUAUCUACAUAA .....-.......(((((.(((((((.......)))))))(-((((..(-((((....(((((....)))))(((((....))))).....)))))..))))))))))............ ( -24.90) >consensus AAUAC_AAUUUCAAAUGUUUGUUAUGUACAACUAGUUAAAA_AAUUACU_AUGCAAAUUAAUUGAAAAAUUUGCAUGGACACGUGUCAAAAGCUUAUAAGCUUACACCAUAUCUACAUAA ..................................................(((((((((........)))))))))(((...((((...(((((....)))))))))....)))...... (-11.94 = -12.50 + 0.56)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:40:52 2006