
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 18,743,874 – 18,743,984 |
| Length | 110 |
| Max. P | 0.939977 |

| Location | 18,743,874 – 18,743,984 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 76.68 |
| Mean single sequence MFE | -29.72 |
| Consensus MFE | -12.90 |
| Energy contribution | -13.04 |
| Covariance contribution | 0.14 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.43 |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.895307 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 18743874 110 + 27905053 C------GCU--GC-AUUUUUGCAUAUCUUGCUGUUUUGUUUGGCAUUUUUAGUGAUUUCACUUAAAUUUUCAUAACAAAAUAUGCACAGUGCGUGCUUG-CAGCUCACUUUUGGCUCAA .------(((--((-(....(((((....(((((((((((((((...(((.((((....)))).)))...))).))))))))).)))..)))))....))-))))............... ( -31.20) >DroVir_CAF1 17453 109 + 1 C------GCC--GU-AUAUCUGCAUAUCUGGCAGUUUUGUUUGGCAUUUUUAGUGAUUUCACUUAAAUUUUCAUAACAAAACAUGAAGAGUGCGUGCAUG-CAGCUCGGCUAA-GCUGAG .------((.--((-((.(.(((((.(((..(((((((((((((...(((.((((....)))).)))...))).)))))))).)).)))))))).).)))-).))(((((...-))))). ( -29.50) >DroGri_CAF1 3163 116 + 1 GCAUGUUGUU--GUGUUAUUUGCAUAUGAAGCAGUUUUGUUUGGCAUUUUUAGUGAUUUCACUUAAAUUUUCAUAACAAAACAAGAAGAGUACGAGCUUG-UAGCUCGGUUAA-GCUGGG ((.((((..(--((((.....)))))...))))(((((((((((...(((.((((....)))).)))...))).))))))))..........(((((...-..))))).....-)).... ( -28.70) >DroSec_CAF1 2703 110 + 1 C------GCU--GC-AUUUUUGCAUAUCUGGCUGUUUUGUUUGGCAUUUUUAGUGAUUUCACUUAAAUUUUCAUAACAAAAUAUGCACAGUGCGUGCUUG-CAGCUCACUUUUGGCUCAA .------(((--((-(.....((((..(((((((((((((((((...(((.((((....)))).)))...))).))))))))).)).)))...)))).))-))))............... ( -32.50) >DroMoj_CAF1 23672 112 + 1 C------GCCUGGU-GGGCCCGCCCCUUUGAUAGUUUUGUUUGGCAUUUUAAGUGAUUUCACUUAAAUUUUCAUAACAAAACAUGAAGAGUGCGUGCUCGGGCGCUCGUGUAU-GCUGAG (------((((...-((((.((((.((((.((.(((((((((((...((((((((....))))))))...))).)))))))))))))).).))).)))))))))((((.(...-.))))) ( -36.30) >DroPer_CAF1 10369 99 + 1 G------ACU--GG-AUUUUUGCAUAUCUUGCCGUUUUGUUUGCCAUU-UUAGUGAUUUCACUUAAAUUUUCAUAACAAAAUAUGCACAGUGCGUGCUUG-CAACAG----------CAG (------.((--((-((........)))((((.((((((((((..(((-(.((((....)))).))))...)).))))))))..((((.....))))..)-))))))----------).. ( -20.10) >consensus C______GCU__GU_AUUUUUGCAUAUCUGGCAGUUUUGUUUGGCAUUUUUAGUGAUUUCACUUAAAUUUUCAUAACAAAACAUGAACAGUGCGUGCUUG_CAGCUCGCUUAU_GCUCAG ....................(((((........(((((((((((...(((.((((....)))).)))...))).)))))))).......))))).......................... (-12.90 = -13.04 + 0.14)



| Location | 18,743,874 – 18,743,984 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 76.68 |
| Mean single sequence MFE | -25.85 |
| Consensus MFE | -10.40 |
| Energy contribution | -10.46 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.69 |
| Structure conservation index | 0.40 |
| SVM decision value | 1.31 |
| SVM RNA-class probability | 0.939977 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 18743874 110 - 27905053 UUGAGCCAAAAGUGAGCUG-CAAGCACGCACUGUGCAUAUUUUGUUAUGAAAAUUUAAGUGAAAUCACUAAAAAUGCCAAACAAAACAGCAAGAUAUGCAAAAAU-GC--AGC------G ...............((((-((.(((.....)))((((((((((((.......(((.((((....)))).))).((.....))....)))))))))))).....)-))--)))------. ( -27.10) >DroVir_CAF1 17453 109 - 1 CUCAGC-UUAGCCGAGCUG-CAUGCACGCACUCUUCAUGUUUUGUUAUGAAAAUUUAAGUGAAAUCACUAAAAAUGCCAAACAAAACUGCCAGAUAUGCAGAUAU-AC--GGC------G ......-...(((((.(((-(((((..(((........((((((((.((..(.(((.((((....)))).))).)..)))))))))))))..).)))))))...)-.)--)))------. ( -26.90) >DroGri_CAF1 3163 116 - 1 CCCAGC-UUAACCGAGCUA-CAAGCUCGUACUCUUCUUGUUUUGUUAUGAAAAUUUAAGUGAAAUCACUAAAAAUGCCAAACAAAACUGCUUCAUAUGCAAAUAACAC--AACAACAUGC ....((-.....(((((..-...)))))........(((((.((((((.....(((.((((....)))).)))..............(((.......))).)))))).--)))))...)) ( -21.10) >DroSec_CAF1 2703 110 - 1 UUGAGCCAAAAGUGAGCUG-CAAGCACGCACUGUGCAUAUUUUGUUAUGAAAAUUUAAGUGAAAUCACUAAAAAUGCCAAACAAAACAGCCAGAUAUGCAAAAAU-GC--AGC------G ...............((((-((.....((((((.((...(((((((.((..(.(((.((((....)))).))).)..)))))))))..)))))...))).....)-))--)))------. ( -25.30) >DroMoj_CAF1 23672 112 - 1 CUCAGC-AUACACGAGCGCCCGAGCACGCACUCUUCAUGUUUUGUUAUGAAAAUUUAAGUGAAAUCACUUAAAAUGCCAAACAAAACUAUCAAAGGGGCGGGCCC-ACCAGGC------G ....((-........))(((.(.((.(((.(((((...((((((((.((..(.((((((((....)))))))).)..)))))))))).....)))))))).)).)-....)))------. ( -31.60) >DroPer_CAF1 10369 99 - 1 CUG----------CUGUUG-CAAGCACGCACUGUGCAUAUUUUGUUAUGAAAAUUUAAGUGAAAUCACUAA-AAUGGCAAACAAAACGGCAAGAUAUGCAAAAAU-CC--AGU------C .((----------((....-..))))...(((((((((((((((((......((((.((((....)))).)-)))(.....).....))))))))))))).....-.)--)))------. ( -23.10) >consensus CUCAGC_AUAACCGAGCUG_CAAGCACGCACUCUGCAUAUUUUGUUAUGAAAAUUUAAGUGAAAUCACUAAAAAUGCCAAACAAAACUGCCAGAUAUGCAAAAAU_AC__AGC______G ...............((......))..(((.........(((((((.((..(.(((.((((....)))).))).)..)))))))))..........)))..................... (-10.40 = -10.46 + 0.06)

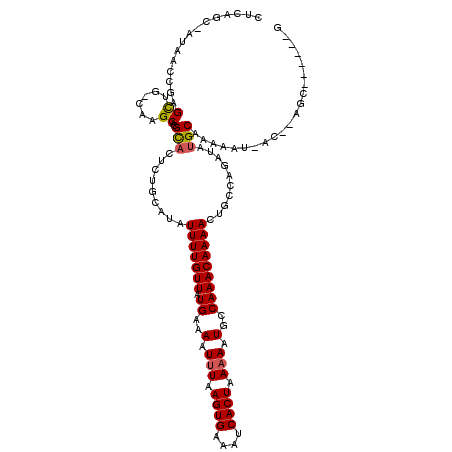

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:37:31 2006