
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 18,711,616 – 18,711,784 |
| Length | 168 |
| Max. P | 0.942624 |

| Location | 18,711,616 – 18,711,726 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 68.90 |
| Mean single sequence MFE | -28.72 |
| Consensus MFE | -17.88 |
| Energy contribution | -17.53 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.40 |
| Mean z-score | -1.17 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.680112 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 18711616 110 + 27905053 CUCAUUAAAAGAGAUAUUAUUUACUAUUCCCAAAAACAGCUAGAAAAAAGCUA----AGGCGCU----ACCUUCAAACCAGGAAGAGCAGCAACUGAAGGGGGCGUGGCAGGCGCCAG (((.......)))........................((((.......)))).----.((((((----..((((.......)))).((.((..(.....)..))...)).)))))).. ( -24.60) >DroSec_CAF1 145436 110 + 1 UUAAUUAAAAGAGGUAUCAUUCACUGUUUCCAAAAAUAGCUAGAAAAUAGCUA----GGGCGCU----ACCUUCGAAUCAGGAAGAGCAGCAACUGAGGGGGGCGUGGCAGGCGCCAG ............(((..(..((((.((..((.....((((((.....))))))----((..(((----.(((((.......)))).).)))..))...))..))))))..)..))).. ( -30.60) >DroSim_CAF1 137088 110 + 1 UUAAUUAAAAGAGGUAUCAUUCACUGUUUCCAAAAAUAGCUAGAAAAUAGCUA----GGGCGCU----ACCUUCGAAUCAGGAAGAGCAGCAACUGAAGGGGGCGUGGCAGGCGCCAG ............(((..(..((((.((..((.....((((((.....))))))----((..(((----.(((((.......)))).).)))..))...))..))))))..)..))).. ( -30.80) >DroEre_CAF1 142930 89 + 1 --------------------GUAAUAUUCC-AAAGAUAGCUAGAAAAGAGCUA----AGGCGCU----ACCUUCGAACCAGGAACAUAAGCAGCUGAAGGGGGCGUGGCAGGCGCCAG --------------------..........-.....(((((.......)))))----.((((((----.(((.......))).......((.(((......)))...)).)))))).. ( -23.00) >DroYak_CAF1 141036 114 + 1 CUAAAUGAAUGUGAUGCCAUCGAAUAUUCCUAAAAAUAGCUAGAAAAUAGCUA----AGGCGCUUUGAGCCUUUGAACCAGGAACAGAAGCAGCUAAAGGGGGCGUGGCAGGCGCCAG ..........(((.(((((.((....(((((.....((((((.....))))))----.((((((((...(((.......)))....))))).)))..))))).)))))))..)))... ( -33.70) >DroAna_CAF1 129791 109 + 1 CUAAACGAAUUGUGUGGCACAGAA--CUCGGGUACUUGGCUUGCAACAAGCCAAAUGAGAAGCUCU--AAUUUUGGGUAACGAACGG----GGAU-GAGGAGGCGUGGCAGGCGCCAG ...........((.((.(((....--((((.....((((((((...)))))))).))))...((((--.(((((.(........).)----))))-..))))..))).)).))..... ( -29.60) >consensus CUAAUUAAAAGAGGUAUCAUUCAAUAUUCCCAAAAAUAGCUAGAAAAUAGCUA____AGGCGCU____ACCUUCGAACCAGGAACAGCAGCAACUGAAGGGGGCGUGGCAGGCGCCAG ....................................((((((.....))))))......(((((.....((((((...................)))))).)))))(((....))).. (-17.88 = -17.53 + -0.36)



| Location | 18,711,656 – 18,711,762 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 84.43 |
| Mean single sequence MFE | -37.80 |
| Consensus MFE | -29.86 |
| Energy contribution | -29.78 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.690401 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 18711656 106 + 27905053 UAGAAAAAAGCUA----AGGCGCU----ACCUUCAAACCAGGAAGAGCAGCAACUGAAGGGGGCGUGGCAGGCGCCAGUUGCCCCAUUUGGGAAAUGGAGGCAGCAGCCUGGGA ..........((.----(((((((----.((((((..(((((.....(((...)))...((((((((((....))))..)))))).)))))....)))))).))).)))).)). ( -40.80) >DroSec_CAF1 145476 106 + 1 UAGAAAAUAGCUA----GGGCGCU----ACCUUCGAAUCAGGAAGAGCAGCAACUGAGGGGGGCGUGGCAGGCGCCAGUUGCCCCAUUUGGGAAAUGGAGGCAGCAGCCUGGGA ..........(((----((((((.----.((((((......(.....)......))))))..))))(((....))).(((((((((((.....))))).))))))..))))).. ( -39.90) >DroSim_CAF1 137128 106 + 1 UAGAAAAUAGCUA----GGGCGCU----ACCUUCGAAUCAGGAAGAGCAGCAACUGAAGGGGGCGUGGCAGGCGCCAGUUGCCCCAUUUGGGAAAUGGAGGCAGCAGCCUGGGA ..........(((----((((((.----.((((((......(.....)......))))))..))))(((....))).(((((((((((.....))))).))))))..))))).. ( -40.20) >DroEre_CAF1 142949 106 + 1 UAGAAAAGAGCUA----AGGCGCU----ACCUUCGAACCAGGAACAUAAGCAGCUGAAGGGGGCGUGGCAGGCGCCAGUUGCCCCAUUUGGGAAAUGGAGGCAGCAGCCAGGGA ..........((.----.((((((----.((((((..(((((..((........))...((((((((((....))))..)))))).)))))....)))))).))).)))..)). ( -37.30) >DroYak_CAF1 141076 110 + 1 UAGAAAAUAGCUA----AGGCGCUUUGAGCCUUUGAACCAGGAACAGAAGCAGCUAAAGGGGGCGUGGCAGGCGCCAGUUGCCCCAUUUGGGAAAUGGAGGCAGCAGCCAGGGA .........(((.----.(((((((((..(((.......)))..)))).((.(((......)))...))..))))).(((((((((((.....))))).)))))))))...... ( -38.20) >DroAna_CAF1 129829 107 + 1 UUGCAACAAGCCAAAUGAGAAGCUCU--AAUUUUGGGUAACGAACGG----GGAU-GAGGAGGCGUGGCAGGCGCCAGUUUCCCCAUUUGGGAAAUGGAGGCAGCAGCCAGGGA ..((.....))...........((((--.(((((.(........).)----))))-..)))).(.((((..(((((.(((((((.....)))))))...))).)).)))).).. ( -30.40) >consensus UAGAAAAUAGCUA____AGGCGCU____ACCUUCGAACCAGGAACAGCAGCAACUGAAGGGGGCGUGGCAGGCGCCAGUUGCCCCAUUUGGGAAAUGGAGGCAGCAGCCAGGGA .........(((......)))........((((((...................)))))).(((.((((....))))(((((((((((.....))))).)))))).)))..... (-29.86 = -29.78 + -0.08)

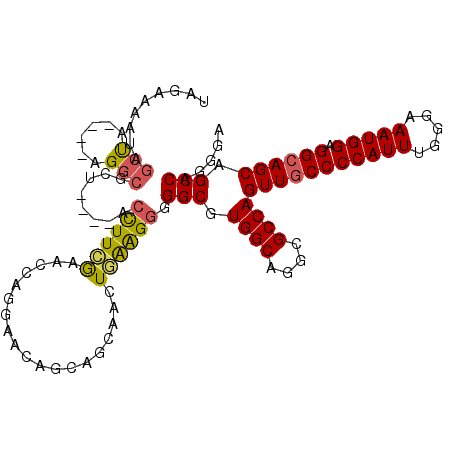

| Location | 18,711,656 – 18,711,762 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 84.43 |
| Mean single sequence MFE | -32.87 |
| Consensus MFE | -22.55 |
| Energy contribution | -22.75 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.69 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.529000 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 18711656 106 - 27905053 UCCCAGGCUGCUGCCUCCAUUUCCCAAAUGGGGCAACUGGCGCCUGCCACGCCCCCUUCAGUUGCUGCUCUUCCUGGUUUGAAGGU----AGCGCCU----UAGCUUUUUUCUA ....((((.(((((((.((....(((...(((((...((((....)))).)))))....(((....))).....)))..)).))))----)))))))----............. ( -35.60) >DroSec_CAF1 145476 106 - 1 UCCCAGGCUGCUGCCUCCAUUUCCCAAAUGGGGCAACUGGCGCCUGCCACGCCCCCCUCAGUUGCUGCUCUUCCUGAUUCGAAGGU----AGCGCCC----UAGCUAUUUUCUA ...((((..(..((..(((((.....)))))(((((((((((.......))))......))))))))).)..))))....((((((----(((....----..))))))))).. ( -33.90) >DroSim_CAF1 137128 106 - 1 UCCCAGGCUGCUGCCUCCAUUUCCCAAAUGGGGCAACUGGCGCCUGCCACGCCCCCUUCAGUUGCUGCUCUUCCUGAUUCGAAGGU----AGCGCCC----UAGCUAUUUUCUA ...((((..(..((..(((((.....)))))(((((((((((.......))))......))))))))).)..))))....((((((----(((....----..))))))))).. ( -33.90) >DroEre_CAF1 142949 106 - 1 UCCCUGGCUGCUGCCUCCAUUUCCCAAAUGGGGCAACUGGCGCCUGCCACGCCCCCUUCAGCUGCUUAUGUUCCUGGUUCGAAGGU----AGCGCCU----UAGCUCUUUUCUA .....(((.(((((((.(...........(((((...((((....)))).)))))..((((..((....))..))))...).))))----)))))).----............. ( -36.10) >DroYak_CAF1 141076 110 - 1 UCCCUGGCUGCUGCCUCCAUUUCCCAAAUGGGGCAACUGGCGCCUGCCACGCCCCCUUUAGCUGCUUCUGUUCCUGGUUCAAAGGCUCAAAGCGCCU----UAGCUAUUUUCUA ....((((((.((((.(((((.....)))))))))...(((((..(((......((...(((.......)))...))......))).....))))).----))))))....... ( -31.40) >DroAna_CAF1 129829 107 - 1 UCCCUGGCUGCUGCCUCCAUUUCCCAAAUGGGGAAACUGGCGCCUGCCACGCCUCCUC-AUCC----CCGUUCGUUACCCAAAAUU--AGAGCUUCUCAUUUGGCUUGUUGCAA .....(((....)))........(((((((((((....((((.......)))).....-.)))----))((((.............--.)))).....)))))).......... ( -26.34) >consensus UCCCAGGCUGCUGCCUCCAUUUCCCAAAUGGGGCAACUGGCGCCUGCCACGCCCCCUUCAGCUGCUGCUCUUCCUGAUUCGAAGGU____AGCGCCU____UAGCUAUUUUCUA .....(((((.((((.(((((.....)))))))))...(((((.......(((....((((..(......)..))))......))).....))))).....)))))........ (-22.55 = -22.75 + 0.20)



| Location | 18,711,688 – 18,711,784 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 90.49 |
| Mean single sequence MFE | -35.22 |
| Consensus MFE | -32.51 |
| Energy contribution | -32.67 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.92 |
| SVM decision value | 1.33 |
| SVM RNA-class probability | 0.942624 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 18711688 96 + 27905053 GGAAGAGCAGCAACUGAAGGGGGCGUGGCAGGCGCCAGUUGCCCCAUUUGGGAAAUGGAGGCAGCAGCCUGGGAGUUUUGCCCAAAAAGUCUCGUA ....((((............((((.((((....))))(((((((((((.....))))).)))))).))))(((.......))).....).)))... ( -34.30) >DroSec_CAF1 145508 96 + 1 GGAAGAGCAGCAACUGAGGGGGGCGUGGCAGGCGCCAGUUGCCCCAUUUGGGAAAUGGAGGCAGCAGCCUGGGAGUUUUGGCCAAAAAGUCUCGCA .........((.........((((.((((....))))(((((((((((.....))))).)))))).))))((((.((((.....)))).)))))). ( -36.60) >DroSim_CAF1 137160 96 + 1 GGAAGAGCAGCAACUGAAGGGGGCGUGGCAGGCGCCAGUUGCCCCAUUUGGGAAAUGGAGGCAGCAGCCUGGGAGUUUUGGCCAAAAAGUCUCGCA .........((.........((((.((((....))))(((((((((((.....))))).)))))).))))((((.((((.....)))).)))))). ( -36.60) >DroEre_CAF1 142981 96 + 1 GGAACAUAAGCAGCUGAAGGGGGCGUGGCAGGCGCCAGUUGCCCCAUUUGGGAAAUGGAGGCAGCAGCCAGGGAGUUUUGGCCAAAAAGUCUCGCA .........((..........(((.((((....))))(((((((((((.....))))).)))))).))).((((.((((.....)))).)))))). ( -35.80) >DroYak_CAF1 141112 96 + 1 GGAACAGAAGCAGCUAAAGGGGGCGUGGCAGGCGCCAGUUGCCCCAUUUGGGAAAUGGAGGCAGCAGCCAGGGAGUUUUGGCCAAAAAGUCUCGCA .........((..........(((.((((....))))(((((((((((.....))))).)))))).))).((((.((((.....)))).)))))). ( -35.80) >DroAna_CAF1 129867 89 + 1 CGAACGG----GGAU-GAGGAGGCGUGGCAGGCGCCAGUUUCCCCAUUUGGGAAAUGGAGGCAGCAGCCAGGGAGUUC-GGCCA-GAAGUCUCGCA .......----....-......((.((((..(((((.(((((((.....)))))))...))).)).))))((((.(((-.....-))).)))))). ( -32.20) >consensus GGAACAGCAGCAACUGAAGGGGGCGUGGCAGGCGCCAGUUGCCCCAUUUGGGAAAUGGAGGCAGCAGCCAGGGAGUUUUGGCCAAAAAGUCUCGCA .........((..........(((.((((....))))(((((((((((.....))))).)))))).))).((((.((((.....)))).)))))). (-32.51 = -32.67 + 0.16)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:37:12 2006