
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 18,282,082 – 18,282,219 |
| Length | 137 |
| Max. P | 0.999941 |

| Location | 18,282,082 – 18,282,193 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 83.41 |
| Mean single sequence MFE | -24.07 |
| Consensus MFE | -15.02 |
| Energy contribution | -15.74 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.62 |
| SVM decision value | 1.11 |
| SVM RNA-class probability | 0.916674 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 18282082 111 + 27905053 AGUCUGGAAUCAUUUAAAAAAUCCCCUUUGGUUUUAAACGCAAUUCGAAUAUUCUUUGCCUUGUUGAUAAACACGUUUUUGGAUUUUUCCAAAAAUGUUGUUUGCUACUUA .((.((((((((................)))))))).))((((............))))...((.(.((((((((((((((((....)))))))))).))))))).))... ( -23.29) >DroSec_CAF1 7639 110 + 1 AGUCCUGAAACAUUUAAAA-AUUCCAUUUAGGUUUAAACGCAUUUCAAAUAUUCUUUGCGUUGUUGACAAACAUGUUUUUGGAUUUUUCCAAAAAUGUUGUUUGUUACUUA ...((((((..........-......))))))....((((((..............))))))..(((((((((((((((((((....)))))))))).))))))))).... ( -26.83) >DroSim_CAF1 8423 110 + 1 AGUCUGGAAACAUUUAAAA-AUUCCCUUUGGGUUUAAACGCAUUUCAAAUAUUCUUUGCGUUGUUGACAAACAUGUUUUUGGAUUUUUCCAAAAAUUUUGUUUGUUACUUA ....((....)).......-...(((...)))....((((((..............))))))..(((((((((.(((((((((....)))))))))..))))))))).... ( -27.94) >DroEre_CAF1 13713 109 + 1 AGUCUGGAAACAUUUUCAA-AUCCCCU-UAGUUUUAAACGCUUUUAAAAUGCUCACUGACCUGUUGACAAACACUUUUUCGGAUUUUUCCAAAAAUGUUCGUUUUUACUCA .(((((....)).......-.......-..((((((((....)))))))).......)))..((.(((.((((.(((((.(((....)))))))))))).)))...))... ( -15.30) >DroYak_CAF1 7731 109 + 1 AGUCUGGAAACAUUUCAAA-AUCCCCU-UAAGUUUAAACGUUUUUAAAACAAUCACUGGGCUGUUGACAAGCACGUUUUUGGAUUUUUCCAGGAAUGUUGUUUGUUACUUA ((((..(((((........-.......-...))))....(((.....))).....)..))))..(((((((((((((((((((....)))))))))).))))))))).... ( -26.97) >consensus AGUCUGGAAACAUUUAAAA_AUCCCCUUUAGGUUUAAACGCAUUUCAAAUAUUCUUUGCGUUGUUGACAAACACGUUUUUGGAUUUUUCCAAAAAUGUUGUUUGUUACUUA ................................................................(((((((((((((((((((....)))))))))).))))))))).... (-15.02 = -15.74 + 0.72)
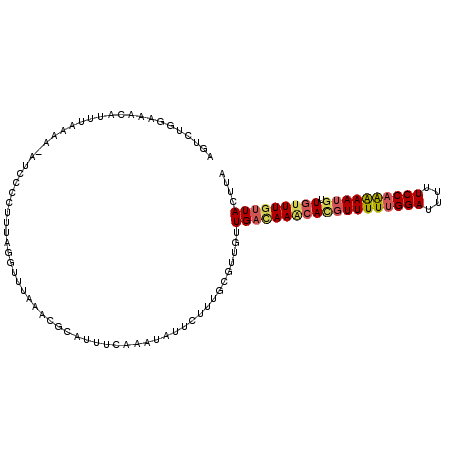



| Location | 18,282,082 – 18,282,193 |
|---|---|
| Length | 111 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 83.41 |
| Mean single sequence MFE | -21.43 |
| Consensus MFE | -13.24 |
| Energy contribution | -14.16 |
| Covariance contribution | 0.92 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.646564 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 18282082 111 - 27905053 UAAGUAGCAAACAACAUUUUUGGAAAAAUCCAAAAACGUGUUUAUCAACAAGGCAAAGAAUAUUCGAAUUGCGUUUAAAACCAAAGGGGAUUUUUUAAAUGAUUCCAGACU ..(((((((((((...((((((((....))))))))..))))).........((((.((....))...)))))))...........((((((........))))))..))) ( -18.80) >DroSec_CAF1 7639 110 - 1 UAAGUAACAAACAACAUUUUUGGAAAAAUCCAAAAACAUGUUUGUCAACAACGCAAAGAAUAUUUGAAAUGCGUUUAAACCUAAAUGGAAU-UUUUAAAUGUUUCAGGACU ...((.(((((((...((((((((....))))))))..)))))))..))........(((((((((((((.((((((....)))))).)..-))))))))))))....... ( -22.80) >DroSim_CAF1 8423 110 - 1 UAAGUAACAAACAAAAUUUUUGGAAAAAUCCAAAAACAUGUUUGUCAACAACGCAAAGAAUAUUUGAAAUGCGUUUAAACCCAAAGGGAAU-UUUUAAAUGUUUCCAGACU ...((.(((((((...((((((((....))))))))..)))))))..))((((((..............))))))..........((((((-........))))))..... ( -23.14) >DroEre_CAF1 13713 109 - 1 UGAGUAAAAACGAACAUUUUUGGAAAAAUCCGAAAAAGUGUUUGUCAACAGGUCAGUGAGCAUUUUAAAAGCGUUUAAAACUA-AGGGGAU-UUGAAAAUGUUUCCAGACU .........(((((((((((((((....)))))))..)))))))).....((((.(..((((((((.(((.(.((((....))-)).)..)-)).))))))))..).)))) ( -25.00) >DroYak_CAF1 7731 109 - 1 UAAGUAACAAACAACAUUCCUGGAAAAAUCCAAAAACGUGCUUGUCAACAGCCCAGUGAUUGUUUUAAAAACGUUUAAACUUA-AGGGGAU-UUUGAAAUGUUUCCAGACU (((((...............((((....)))).((((((..(((..(((((........))))).)))..))))))..)))))-.((..((-........))..))..... ( -17.40) >consensus UAAGUAACAAACAACAUUUUUGGAAAAAUCCAAAAACGUGUUUGUCAACAACGCAAAGAAUAUUUGAAAUGCGUUUAAACCUAAAGGGGAU_UUUUAAAUGUUUCCAGACU ...((.(((((((...((((((((....))))))))..)))))))..))........((((((((((((...((((...........)))).))))))))))))....... (-13.24 = -14.16 + 0.92)



| Location | 18,282,113 – 18,282,219 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 69.80 |
| Mean single sequence MFE | -20.75 |
| Consensus MFE | -13.11 |
| Energy contribution | -15.86 |
| Covariance contribution | 2.75 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.90 |
| Structure conservation index | 0.63 |
| SVM decision value | 4.71 |
| SVM RNA-class probability | 0.999941 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 18282113 106 + 27905053 UUUUAAACGCAAUUCGAAUAUUCUUUGCCUUGUUGAUAAACACGUUUUUGGAUUUUUCCAAAAAUGUUGUUUGCUACUUA-------------UCUUAAAAAAAAUACCCUGUCUAAUA ((((((..((((............))))...((.(.((((((((((((((((....)))))))))).))))))).))...-------------..)))))).................. ( -19.10) >DroSec_CAF1 7669 92 + 1 GUUUAAACGCAUUUCAAAUAUUCUUUGCGUUGUUGACAAACAUGUUUUUGGAUUUUUCCAAAAAUGUUGUUUGUUACUUA-------------UCUU--------------AAAAAGAA .....((((((..............))))))..(((((((((((((((((((....)))))))))).)))))))))....-------------....--------------........ ( -24.24) >DroSim_CAF1 8453 92 + 1 GUUUAAACGCAUUUCAAAUAUUCUUUGCGUUGUUGACAAACAUGUUUUUGGAUUUUUCCAAAAAUUUUGUUUGUUACUUA-------------CCCU--------------GCCUAAUA .....((((((..............))))))..(((((((((.(((((((((....)))))))))..)))))))))....-------------....--------------........ ( -24.24) >DroEre_CAF1 13742 118 + 1 UUUUAAACGCUUUUAAAAUGCUCACUGACCUGUUGACAAACACUUUUUCGGAUUUUUCCAAAAAUGUUCGUUUUUACUCAAAAGGUGUACUUAUCUCAUA-AAAAUUCCCGGCAUAAUC ........(((.......((.((((......).))))).(((((((((.(((....)))((((((....))))))....)))))))))............-.........)))...... ( -15.40) >consensus GUUUAAACGCAUUUCAAAUAUUCUUUGCCUUGUUGACAAACACGUUUUUGGAUUUUUCCAAAAAUGUUGUUUGUUACUUA_____________UCUU______________GCAUAAUA .....((((((..............))))))..(((((((((((((((((((....)))))))))).)))))))))........................................... (-13.11 = -15.86 + 2.75)


| Location | 18,282,113 – 18,282,219 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 69.80 |
| Mean single sequence MFE | -19.04 |
| Consensus MFE | -9.70 |
| Energy contribution | -10.32 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.51 |
| SVM decision value | 0.92 |
| SVM RNA-class probability | 0.881865 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 18282113 106 - 27905053 UAUUAGACAGGGUAUUUUUUUUAAGA-------------UAAGUAGCAAACAACAUUUUUGGAAAAAUCCAAAAACGUGUUUAUCAACAAGGCAAAGAAUAUUCGAAUUGCGUUUAAAA ..((((((..(.(((((.((....))-------------.))))).)(((((...((((((((....))))))))..))))).........((((.((....))...)))))))))).. ( -18.60) >DroSec_CAF1 7669 92 - 1 UUCUUUUU--------------AAGA-------------UAAGUAACAAACAACAUUUUUGGAAAAAUCCAAAAACAUGUUUGUCAACAACGCAAAGAAUAUUUGAAAUGCGUUUAAAC ........--------------....-------------...((.(((((((...((((((((....))))))))..)))))))..))((((((..............))))))..... ( -17.64) >DroSim_CAF1 8453 92 - 1 UAUUAGGC--------------AGGG-------------UAAGUAACAAACAAAAUUUUUGGAAAAAUCCAAAAACAUGUUUGUCAACAACGCAAAGAAUAUUUGAAAUGCGUUUAAAC ......((--------------(...-------------(((((((((((((...((((((((....))))))))..)))))))......(.....)..))))))...)))........ ( -18.10) >DroEre_CAF1 13742 118 - 1 GAUUAUGCCGGGAAUUUU-UAUGAGAUAAGUACACCUUUUGAGUAAAAACGAACAUUUUUGGAAAAAUCCGAAAAAGUGUUUGUCAACAGGUCAGUGAGCAUUUUAAAAGCGUUUAAAA ...........((((.((-(.((((((...(((((((.((((.......((((((((((((((....)))))))..))))))))))).))))..)))...)))))).))).)))).... ( -21.81) >consensus UAUUAGGC______________AAGA_____________UAAGUAACAAACAACAUUUUUGGAAAAAUCCAAAAACAUGUUUGUCAACAACGCAAAGAAUAUUUGAAAUGCGUUUAAAA ..........................................((.(((((((...((((((((....))))))))..)))))))..))............................... ( -9.70 = -10.32 + 0.63)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:33:36 2006