| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
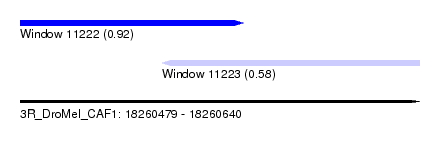
| Location | 18,260,479 – 18,260,640 |
| Length | 161 |
| Max. P | 0.921133 |

| Location | 18,260,479 – 18,260,569 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 72.30 |
| Mean single sequence MFE | -27.83 |
| Consensus MFE | -13.08 |
| Energy contribution | -13.05 |
| Covariance contribution | -0.03 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.47 |
| SVM decision value | 1.14 |
| SVM RNA-class probability | 0.921133 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 18260479 90 + 27905053 UUGGCUAAUUUUGGGCUGGAGAAAUCAGCGGC-------CGGCAAGGAAGUCAACGUUU---------------UUUUUU---UUUUUUGCCAACGCAAACA--GAAAUGUGAAA--AA ...((........(((((((....))..))))-------)((((((((((.........---------------.....)---)))))))))...))..(((--....)))....--.. ( -23.14) >DroPse_CAF1 20687 105 + 1 AUGGCUAAUUUUGGGCUGGUGAAAUCUGCAGGCAGAAAUUGGCAAGGAAGCUGCCGAGCUG--------CAGAG-CUUCU---UUUUUCGCCAAGGCAAACUAGUAAAAGCGAAA--AA ...(((((((((.(.(((..(....)..))).).)))))))))(((((((((((......)--------)..))-)))))---))((((((....((......))....))))))--.. ( -30.90) >DroSec_CAF1 16504 84 + 1 UUGGCUAAUUUUGGGCUGGAGAAAUCAGCGGC-------CGGCAAGGAAGCCAUCGUUU---------------U---------UUUUUGCCAACGCAAACA--GAAAUGUGUAA--AA .(((((..((((.(((((((....))..))))-------).).)))..))))).(((((---------------(---------(.(((((....))))).)--)))))).....--.. ( -25.10) >DroEre_CAF1 17075 87 + 1 UUGGCUAAUUUUGGGCUGGAGAAAUCAGCGGC-------CGGCAAGGAAGCCUCCGUAU---------------UUUUU--------UUGCCAACGCAAACA--GAAAUGUGAAAAAAA ..((((..((((.(((((((....))..))))-------).).)))..))))..(((((---------------(((((--------((((....))))).)--))))))))....... ( -22.80) >DroYak_CAF1 16814 93 + 1 UUGGCUAAUUUUGGGCUGGAGAAAUCAGCGGC-------CGGCAAGGAAGCCUCUGUUU---------------UUUUUUUGCUUUUUUGCCAACGCAAACA--GAAAUGUGAAA--AA ...((........(((((((....))..))))-------)(((((((((((........---------------.......)))))))))))...))..(((--....)))....--.. ( -27.66) >DroPer_CAF1 20894 113 + 1 AUGGCUAAUUUUGGGCUGGUGAAAUCUGCAGGCAGAAAUUGGCAAGGAAGCUGCCGAGCUGCAGAGCUGCAGAG-CUUCU---UUUUUCGCCAAGGCAAACUAGUAAAAGCGAAA--AA ..((((.......))))...(((.(((((((((((...((((((.(....))))))).))))....))))))).-.))).---((((((((....((......))....))))))--)) ( -37.40) >consensus UUGGCUAAUUUUGGGCUGGAGAAAUCAGCGGC_______CGGCAAGGAAGCCACCGUUU_______________UUUUUU___UUUUUUGCCAACGCAAACA__GAAAUGUGAAA__AA .((((.........(((((.....)))))...........(((......))).....................................)))).......................... (-13.08 = -13.05 + -0.03)



| Location | 18,260,536 – 18,260,640 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.25 |
| Mean single sequence MFE | -18.95 |
| Consensus MFE | -16.34 |
| Energy contribution | -15.98 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.576543 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 18260536 104 - 27905053 AAAUUUCACUUUUCACCAUUGCAUAAUUGCAGUUUGACAAUGCUUAAAAUUGAUUAUGCUGCGUU--------UUC-CUUUU--UUUCACAUUUC--UGUUUGCGUUGGCAAAAAA---A .........((((..(((.((((.....(((((.(((((((.......)))).))).)))))...--------...-.....--....(((....--))).)))).)))..)))).---. ( -17.30) >DroPse_CAF1 20757 115 - 1 AAAUUUCACUUUUCACCAUUGCAUAAUUGCAGUUUGACAAUGCUUAAAAUUGAUUAUGCUGCGUUACUUUUCUCUCUCCUUU--UUUCGCUUUUACUAGUUUGCCUUGGCGAAAAA---A .........(((((.(((..(((.....(((((.(((((((.......)))).))).)))))....................--....(((......))).)))..))).))))).---. ( -21.80) >DroEre_CAF1 17132 101 - 1 AAAUUUCACUUUUCACCAUUGCAUAAUUGCAGUUUGACAAUGCUUAAAAUUGAUUAUGCUGCGUU--------UUC-CUUUUUUUUUCACAUUUC--UGUUUGCGUUGGCAA-------- ...............(((.((((.....(((((.(((((((.......)))).))).)))))...--------...-...........(((....--))).)))).)))...-------- ( -16.50) >DroYak_CAF1 16871 107 - 1 AAAUUUCACUUUUCACCAUUGCAUAAUUGCAGUUUGACAAUGCUUAAAAUUGAUUAUGCUGCGUU--------UUC-CUUUU--UUUCACAUUUC--UGUUUGCGUUGGCAAAAAAGCAA ........(((((..(((.((((.....(((((.(((((((.......)))).))).)))))...--------...-.....--....(((....--))).)))).)))...)))))... ( -18.30) >DroAna_CAF1 17128 105 - 1 AAAUUUCACUUUUCACCGUUGCAUAAUUGCAGUUUGACAAUGCUUAAAAUUGAUUAUGCUGCACU--------CGCACUUUU--UUUCGCUUUUC--UGUUUGCGUUGGCAAAAAU---A .........((((..(((.((((....((((((.(((((((.......)))).))).))))))..--------.((......--....)).....--....)))).)))..)))).---. ( -18.00) >DroPer_CAF1 20972 115 - 1 AAAUUUCACUUUUCACCAUUGCAUAAUUGCAGUUUGACAAUGCUUAAAAUUGAUUAUGCUGCGUUACUUUUCUCUCUCCUUU--UUUCGCUUUUACUAGUUUGCCUUGGCGAAAAA---A .........(((((.(((..(((.....(((((.(((((((.......)))).))).)))))....................--....(((......))).)))..))).))))).---. ( -21.80) >consensus AAAUUUCACUUUUCACCAUUGCAUAAUUGCAGUUUGACAAUGCUUAAAAUUGAUUAUGCUGCGUU________CUC_CUUUU__UUUCACAUUUC__UGUUUGCGUUGGCAAAAAA___A ...............(((.((((....((((((.(((((((.......)))).))).)))))).........................(((......))).)))).)))........... (-16.34 = -15.98 + -0.36)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:33:11 2006