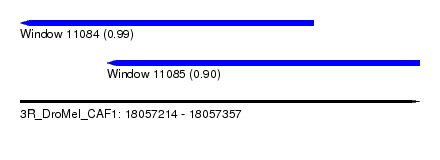

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 18,057,214 – 18,057,357 |
| Length | 143 |
| Max. P | 0.986874 |

| Location | 18,057,214 – 18,057,319 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 71.31 |
| Mean single sequence MFE | -26.13 |
| Consensus MFE | -8.55 |
| Energy contribution | -9.83 |
| Covariance contribution | 1.28 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.72 |
| Structure conservation index | 0.33 |
| SVM decision value | 2.06 |
| SVM RNA-class probability | 0.986874 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 18057214 105 - 27905053 CAAUCAGUCCUGGUCAAGUGGGUGUGGAAACACCGGAAAAUAAUUUGGCAACAAAGUUGAGGU-UAU-CAAUCAAA----U---AU-AUUAUUUUAACC----UACUAAUCAGUAAAUA .........(((((..((((((((((....)))...(((((((((((....))))(((((...-..)-))))....----.---..-.))))))).)))----)))).)))))...... ( -26.50) >DroSec_CAF1 99381 108 - 1 CAAUCAGUCCUGGUCAAGUGGGUGUGGAAACACCGGAAAAUAAUGUGGCAACAAAGUUGAGGA-AAU-CAAUCAAA----U---AU-AUUAUUUUAACCU-UCUACUAAUCAGUAAAAA .........(((((..(((((((((....)))))(((((((((((((....)...(((((...-..)-))))....----.---))-))))))))..)).-..)))).)))))...... ( -24.30) >DroSim_CAF1 102234 105 - 1 CAAUCAGUCCUGGUCAAGUGGGUGUGGAAACACCGGAAAAUAAUAUGGCAACAAAGUUGAGAA-AAU-CAAUCAAA----U---AU-AUUAUUUUAACC----UACUAAUCAGUAAAAA .........(((((..((((((((((....)))...(((((((((((....)...(((((...-..)-))))....----.---))-)))))))).)))----)))).)))))...... ( -27.20) >DroEre_CAF1 102887 109 - 1 CAAUCAGUCCUGGUCAAGUGGGUGUGGAAACACCGGAAAAUAAUUUGGCAACAAAGCUGGUUG-AAUGCGAAAUAA----U---AU-GUUCUUUUCGCCUACCUACUAAUCAGGAAAA- .......(((((((..((((((((.(....)(((((.......((((....)))).)))))..-...((((((...----.---..-.....)))))).)))))))).)))))))...- ( -33.90) >DroYak_CAF1 103814 114 - 1 CAAUCAGUCCUGGUCAAGUGGGCGUGGAAACACCGGAAAAUAAUUUGGUAACAAAGUGGUUUG-CAUAGGAACUAAACUAU---AUAGUUCCUUUAACCUAUUUGCUAAUCAGGAAAA- .......(((((((((((((((.(((....))).(.(((....((((....))))....))).-)..((((((((......---.))))))))....))))))))...)))))))...- ( -34.60) >DroAna_CAF1 96621 92 - 1 CAAUCAAACCAGGCUAAGAGGGUAUUCAUGCACUGAGAAAAG---------UUAAGCUGAGGAUUAU-GA---AAA----UGCCAC-AUUAUUAU---------ACAGAUCAAUCUCCA ...........((.(((...((((((((((..((.((.....---------.....)).))...)))-))---..)----))))..-.)))....---------..(((....))))). ( -10.30) >consensus CAAUCAGUCCUGGUCAAGUGGGUGUGGAAACACCGGAAAAUAAUUUGGCAACAAAGUUGAGGA_AAU_CAAUCAAA____U___AU_AUUAUUUUAACC____UACUAAUCAGUAAAAA .........(((((..(((((((((....)))))...........((....))..................................................)))).)))))...... ( -8.55 = -9.83 + 1.28)


| Location | 18,057,245 – 18,057,357 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 77.72 |
| Mean single sequence MFE | -27.23 |
| Consensus MFE | -13.71 |
| Energy contribution | -15.27 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.34 |
| Structure conservation index | 0.50 |
| SVM decision value | 1.03 |
| SVM RNA-class probability | 0.904200 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 18057245 112 - 27905053 AGAUGAUCAAAUCUAAUGGAAAG--GGCUAGCAAAGCGAGCAAUCAGUCCUGGUCAAGUGGGUGUGGAAACACCGGAAAAUAAUUUGGCAACAAAGUUGAGGU-UAU-CAAUCAAA--- .(((((((...(((..(((..((--((((.((...))((....))))))))..)))....(((((....))))))))......((((....)))).....)))-)))-).......--- ( -33.60) >DroSec_CAF1 99415 112 - 1 AGAUGAUCAAAUCUAAUGGAAAG--GGCUAGCAAAGCGAGCAAUCAGUCCUGGUCAAGUGGGUGUGGAAACACCGGAAAAUAAUGUGGCAACAAAGUUGAGGA-AAU-CAAUCAAA--- .(((..((((.(((..(((..((--((((.((...))((....))))))))..)))....(((((....))))))))........((....))...))))...-.))-).......--- ( -28.30) >DroSim_CAF1 102265 112 - 1 AGAUGAUCAAAUCUAAUGGAAAG--GGCUAGCAAAGCGAGCAAUCAGUCCUGGUCAAGUGGGUGUGGAAACACCGGAAAAUAAUAUGGCAACAAAGUUGAGAA-AAU-CAAUCAAA--- .(((..((((.(((..(((..((--((((.((...))((....))))))))..)))....(((((....))))))))........((....))...))))...-.))-).......--- ( -28.40) >DroEre_CAF1 102921 113 - 1 AGAUGAUCAAAUCUAAUGGAAAG--GGCUGGCUAAGCGAGCAAUCAGUCCUGGUCAAGUGGGUGUGGAAACACCGGAAAAUAAUUUGGCAACAAAGCUGGUUG-AAUGCGAAAUAA--- ....(((((..(((..(((..((--((((((((.....)))...)))))))..)))....(((((....))))))))......((((....))))..))))).-............--- ( -30.70) >DroYak_CAF1 103850 116 - 1 AGAUGAUCAAAUCUAAUGGAAGG--GGCCAGCUAAGCGAGCAAUCAGUCCUGGUCAAGUGGGCGUGGAAACACCGGAAAAUAAUUUGGUAACAAAGUGGUUUG-CAUAGGAACUAAACU ((((......))))..(((....--..))).(((.((((((..((.((((.........))))(((....)))..))......((((....))))...)))))-).))).......... ( -25.20) >DroAna_CAF1 96650 95 - 1 --------AAAUCUAAUGGGAAGAGUGCUGGCAAAACGAGCAAUCAAACCAGGCUAAGAGGGUAUUCAUGCACUGAGAAAAG---------UUAAGCUGAGGAUUAU-GA---AAA--- --------.(((((..(((...((.((((.(.....).)))).))...)))((((..(((....)))....(((......))---------)..))))..)))))..-..---...--- ( -17.20) >consensus AGAUGAUCAAAUCUAAUGGAAAG__GGCUAGCAAAGCGAGCAAUCAGUCCUGGUCAAGUGGGUGUGGAAACACCGGAAAAUAAUUUGGCAACAAAGUUGAGGA_AAU_CAAUCAAA___ ...((((((........(((......(((.((...)).)))......)))))))))....(((((....)))))............................................. (-13.71 = -15.27 + 1.56)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:30:58 2006