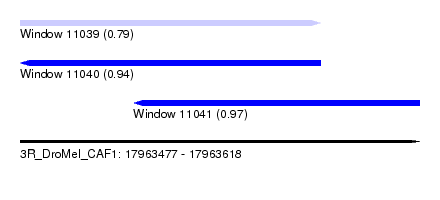

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 17,963,477 – 17,963,618 |
| Length | 141 |
| Max. P | 0.974276 |

| Location | 17,963,477 – 17,963,583 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 72.76 |
| Mean single sequence MFE | -29.50 |
| Consensus MFE | -19.00 |
| Energy contribution | -20.72 |
| Covariance contribution | 1.72 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.786773 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17963477 106 + 27905053 GCGCGGUUUUCGUACCCGAGAUCCUCGUGGAUUAUGCGGAUAUACUGCGUUUGACCAUCAAAAUGUAAUGGUCAAAAGGAACAACUUUUAUUUGC-----UCAAGCUUUAA (((((((.((((((((((((...)))).))....))))))...)))))))..((((((.........))))))(((((......)))))....((-----....))..... ( -29.10) >DroSim_CAF1 7855 111 + 1 GCGCGGUUUUCGUACCCGAGAUCCUCGUGGAUUAUGCGGAUAUACCGCGUUUGACCAUCAAAAUGUAAUGGUCAAAAGGUACAACUUUUAUUUGUUUGUUUCCAGCUUAAA (((((((.((((((((((((...)))).))....))))))...)))))))((((((((.........))))))))..((.(((((........).))))..))........ ( -33.30) >DroEre_CAF1 7104 111 + 1 GCGCGGUUUUCGUACCCGAGAUCCUCGUGGAUUAUGGGGGUAUGCUGGGUUUGACCAUCAAAAUGUAAUGGUCAAAAGGAACAACUUUCAUUUGCUUGCUUCAAGCCUUAA ....(((((.....((((.(((((....))))).))))((((.((....(((((((((.........))))))))).((((....))))....)).))))..))))).... ( -30.70) >DroYak_CAF1 7400 111 + 1 GCGCGGUUUUCGUACCCGAGAUCCUCGUGGAUUAUGGGGGUAUACGGCGUUUGACCAUCAAAAUGUAAUGGUCAAAAGGAACAACUUUUGUUUGCUUGCUUAAAGCUUUAA ((((.((.......((((.(((((....))))).)))).....)).))))((((((((.........))))))))...(((((.....)))))((((.....))))..... ( -33.30) >DroAna_CAF1 6682 80 + 1 GCGCGCUUUUAUUACCCG--GUCCUUUUGGAUCGCAGGGGUAUAUUGUAUUAAAGCCGCAAAAUGACACGAUUGCAGAGUGC----------------------------- (((.((((((((((((((--((((....)))))....)))))....)))..)))))))).......(((.........))).----------------------------- ( -21.10) >consensus GCGCGGUUUUCGUACCCGAGAUCCUCGUGGAUUAUGGGGGUAUACUGCGUUUGACCAUCAAAAUGUAAUGGUCAAAAGGAACAACUUUUAUUUGCUUG_UUCAAGCUUUAA (((((((....((((((..(((((....)))))....)))))))))))))((((((((.........)))))))).................................... (-19.00 = -20.72 + 1.72)

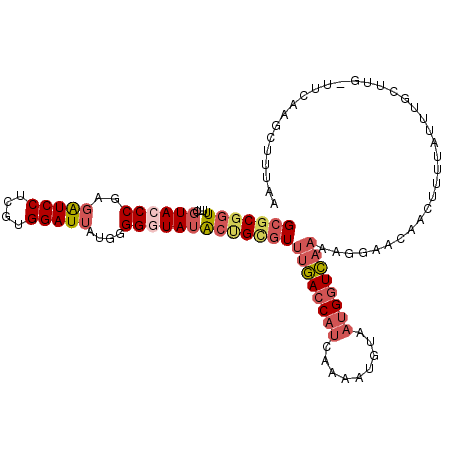

| Location | 17,963,477 – 17,963,583 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 72.76 |
| Mean single sequence MFE | -26.74 |
| Consensus MFE | -15.98 |
| Energy contribution | -16.86 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.60 |
| SVM decision value | 1.33 |
| SVM RNA-class probability | 0.942879 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17963477 106 - 27905053 UUAAAGCUUGA-----GCAAAUAAAAGUUGUUCCUUUUGACCAUUACAUUUUGAUGGUCAAACGCAGUAUAUCCGCAUAAUCCACGAGGAUCUCGGGUACGAAAACCGCGC .........((-----((((.......))))))..(((((((((((.....)))))))))))(((.((.((((((....((((....))))..)))))).....)).))). ( -27.80) >DroSim_CAF1 7855 111 - 1 UUUAAGCUGGAAACAAACAAAUAAAAGUUGUACCUUUUGACCAUUACAUUUUGAUGGUCAAACGCGGUAUAUCCGCAUAAUCCACGAGGAUCUCGGGUACGAAAACCGCGC .....((((....))............(((((((((((((((((((.....))))))))))).((((.....))))...((((....))))...)))))))).....)).. ( -35.40) >DroEre_CAF1 7104 111 - 1 UUAAGGCUUGAAGCAAGCAAAUGAAAGUUGUUCCUUUUGACCAUUACAUUUUGAUGGUCAAACCCAGCAUACCCCCAUAAUCCACGAGGAUCUCGGGUACGAAAACCGCGC .....(((((...)))))........((((.....(((((((((((.....)))))))))))..)))).(((((.....((((....))))...)))))............ ( -28.20) >DroYak_CAF1 7400 111 - 1 UUAAAGCUUUAAGCAAGCAAACAAAAGUUGUUCCUUUUGACCAUUACAUUUUGAUGGUCAAACGCCGUAUACCCCCAUAAUCCACGAGGAUCUCGGGUACGAAAACCGCGC .....((((.....)))).................(((((((((((.....)))))))))))(((.((.(((((.....((((....))))...))))).....)).))). ( -26.80) >DroAna_CAF1 6682 80 - 1 -----------------------------GCACUCUGCAAUCGUGUCAUUUUGCGGCUUUAAUACAAUAUACCCCUGCGAUCCAAAAGGAC--CGGGUAAUAAAAGCGCGC -----------------------------((.((((((((..........)))))).............(((((....(.(((....))).--)))))).....)).)).. ( -15.50) >consensus UUAAAGCUUGAA_CAAGCAAAUAAAAGUUGUUCCUUUUGACCAUUACAUUUUGAUGGUCAAACGCAGUAUACCCCCAUAAUCCACGAGGAUCUCGGGUACGAAAACCGCGC .............................((....(((((((((((.....))))))))))).......(((((.....((((....))))...)))))........)).. (-15.98 = -16.86 + 0.88)

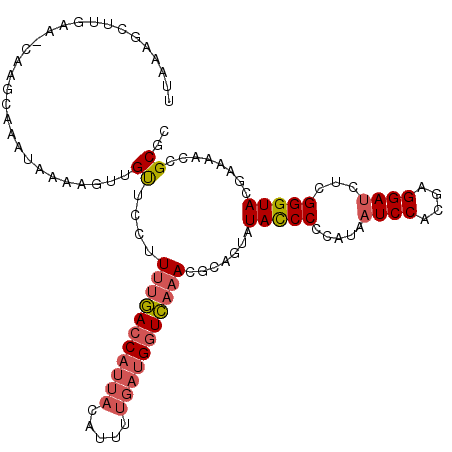
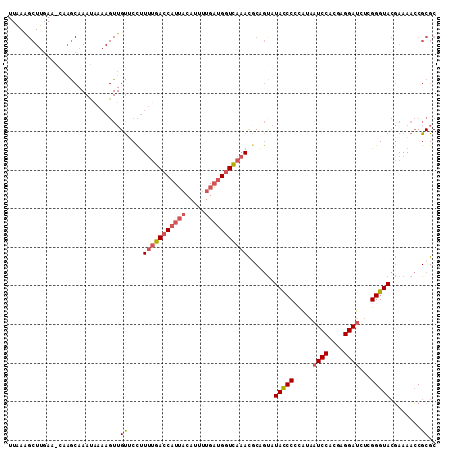
| Location | 17,963,517 – 17,963,618 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 76.70 |
| Mean single sequence MFE | -23.75 |
| Consensus MFE | -12.60 |
| Energy contribution | -12.35 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.53 |
| SVM decision value | 1.73 |
| SVM RNA-class probability | 0.974276 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17963517 101 - 27905053 UAAA----UUGUA-AAAAUCUAAAAACUCCUUUAUUUCGCUUAAAGCUUGA-----GCAAAUAAAAGUUGUUCCUUUUGACCAUUACAUUUUGAUGGUCAAACGCAGUAUA ...(----((((.-..........((((..(((((((.(((((.....)))-----)))))))))))))......(((((((((((.....))))))))))).)))))... ( -23.80) >DroSim_CAF1 7895 106 - 1 UAUA----AUGUU-GAAAUCUAAAAACUCCUUUAUUUCGCUUUAAGCUGGAAACAAACAAAUAAAAGUUGUACCUUUUGACCAUUACAUUUUGAUGGUCAAACGCGGUAUA ....----.....-..........((((..(((((((.((.....))((....))...)))))))))))(((((.(((((((((((.....)))))))))))...))))). ( -23.60) >DroEre_CAF1 7144 111 - 1 CUCAGUUUCCGUUUGAAGAUGAAAAACUCUUUUUGUAAUCUUAAGGCUUGAAGCAAGCAAAUGAAAGUUGUUCCUUUUGACCAUUACAUUUUGAUGGUCAAACCCAGCAUA ....(((((.((((.(((((..((((....))))...))))).))))..)))))..((.(((....)))......(((((((((((.....)))))))))))....))... ( -25.00) >DroYak_CAF1 7440 110 - 1 CUUAGUUUCUGUUUAAAGAUAUACUACUCAUAUU-UCCGCUUAAAGCUUUAAGCAAGCAAACAAAAGUUGUUCCUUUUGACCAUUACAUUUUGAUGGUCAAACGCCGUAUA ..((((.(((......)))...))))........-...((((((....))))))..((.((((.....))))...(((((((((((.....))))))))))).))...... ( -22.60) >consensus CAUA____CUGUU_AAAAACUAAAAACUCCUUUAUUUCGCUUAAAGCUUGAAGCAAGCAAAUAAAAGUUGUUCCUUUUGACCAUUACAUUUUGAUGGUCAAACGCAGUAUA .............................................(((........((((.......))))....(((((((((((.....)))))))))))...)))... (-12.60 = -12.35 + -0.25)

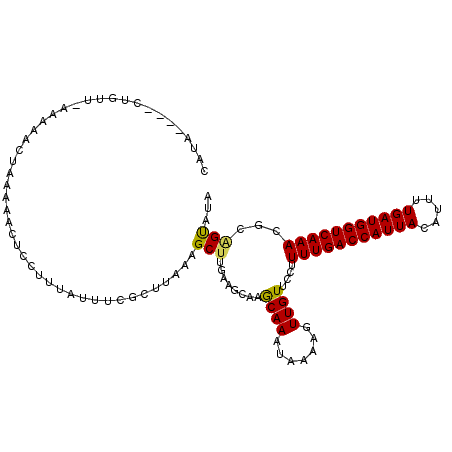

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:30:17 2006