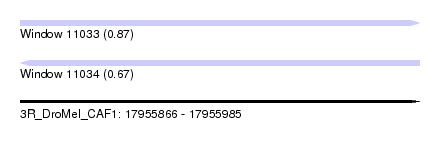

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 17,955,866 – 17,955,985 |
| Length | 119 |
| Max. P | 0.873165 |

| Location | 17,955,866 – 17,955,985 |
|---|---|
| Length | 119 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 82.24 |
| Mean single sequence MFE | -42.87 |
| Consensus MFE | -31.02 |
| Energy contribution | -31.30 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.41 |
| Mean z-score | -2.56 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.873165 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
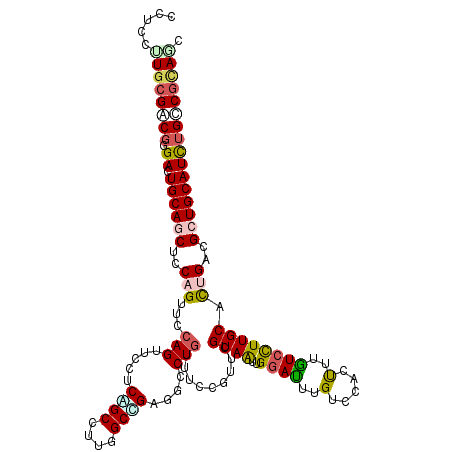
>3R_DroMel_CAF1 17955866 119 + 27905053 ACUGCGGCAGAUGCAGCGGCAGUGCAAGGACAAGACGGACAAGUCCAACUACGAGCGAAAGCAGGCGACCGCCAAGGCGGAGGAGCUGGAACUGGAGUUGCAGUCCCGCCGUAGGGAGU (((((.((.......)).)))))....((((...........))))..((((..((....)).(((..((((....)))).((..(((.(((....))).)))..)))))))))..... ( -50.40) >DroVir_CAF1 789 119 + 1 GAUGCGGCAAAUGCAGCGUCAGUGCAAGGAUAAACUGGACAAAUCCAAUUACGAACGCAAACAGGCAACGGCCAAGGCCGAGGAACUGGAACUGGAGCUGCAAUCACGUCGCAAAGAGG ..((((((...((((((..((((((.((......))(((....)))..........))...(((.(..((((....))))..)..)))..))))..)))))).....))))))...... ( -39.90) >DroGri_CAF1 407 119 + 1 GAUGCGUCAAAUGCAGCGCCAGUGCAAGGACAAACUUGACAAAUCCAAUUACGAACGCAAACAGGCAUCGGCCAAGGCUGAGGAACUGGAUCUGGAGCUGCAGUCGCGACGCAAGGAGG ..((((((...((((((.((((((((((......)))).)).(((((.........((......)).(((((....))))).....))))))))).)))))).....))))))...... ( -44.50) >DroWil_CAF1 353 119 + 1 GUUAAGGCAGAUGCAGCGUCAAUGCAAAGAUAAAGUCGACAAAUGCAAUUACGAAAGGAAACAGGCUUCAGCCAAGGCUGAUGAACUUGAACAGGAGUUGCAAUCGCGUCGCAAGGAGA ......((.(((((.(((....)))..................(((((((.(....)....((((..(((((....)))))....))))......)))))))...)))))))....... ( -30.30) >DroAna_CAF1 354 119 + 1 GCUGCGCCAGAUGCAGCGCCAGUGCAAGGACAAAGUGGACAAGUCCAACUACGAGCGGAAGCAGGCCGCUGCCAAGGCGGAGGAGCUGGAGCAGGAGCUGCAGUCCAGGCGCAAGGAGU ..((((((....((((((((.(((...((((...........))))...)))..((....)).)).)))))).........(((.(((.(((....))).)))))).))))))...... ( -49.40) >DroPer_CAF1 357 119 + 1 GCUGCGGCAAAUGCAGCGUCAGUGCAAGGACAAGAUGGACAAAUCCAAUUACGAACGGAAACAGGCCUCAGCCAAGGCCGAGGAACUGGAACUGGAGCUGCAGUCCCGUCGCAAAGAGG ((.((((....((((((..((((....(..(.....)..)...((((.........(....)...((((.((....)).))))...))))))))..))))))...)))).))....... ( -42.70) >consensus GCUGCGGCAAAUGCAGCGUCAGUGCAAGGACAAAAUGGACAAAUCCAAUUACGAACGGAAACAGGCAUCAGCCAAGGCCGAGGAACUGGAACUGGAGCUGCAGUCCCGUCGCAAGGAGG ..((((((...((((((.(((((.(..((((...........))))...............(((....((((....)))).....)))).))))).)))))).....))))))...... (-31.02 = -31.30 + 0.28)



| Location | 17,955,866 – 17,955,985 |
|---|---|
| Length | 119 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 82.24 |
| Mean single sequence MFE | -39.47 |
| Consensus MFE | -25.05 |
| Energy contribution | -25.58 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.43 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.671042 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17955866 119 - 27905053 ACUCCCUACGGCGGGACUGCAACUCCAGUUCCAGCUCCUCCGCCUUGGCGGUCGCCUGCUUUCGCUCGUAGUUGGACUUGUCCGUCUUGUCCUUGCACUGCCGCUGCAUCUGCCGCAGU .....((((((((((((((......)))))))(((.(..((((....))))..)...)))...)).)))))..((((..(.....)..))))....(((((.((.......)).))))) ( -43.00) >DroVir_CAF1 789 119 - 1 CCUCUUUGCGACGUGAUUGCAGCUCCAGUUCCAGUUCCUCGGCCUUGGCCGUUGCCUGUUUGCGUUCGUAAUUGGAUUUGUCCAGUUUAUCCUUGCACUGACGCUGCAUUUGCCGCAUC ......((((..(..(.((((((..((((..(((.....((((....))))....)))...(((.....(((((((....)))))))......)))))))..)))))).)..))))).. ( -40.20) >DroGri_CAF1 407 119 - 1 CCUCCUUGCGUCGCGACUGCAGCUCCAGAUCCAGUUCCUCAGCCUUGGCCGAUGCCUGUUUGCGUUCGUAAUUGGAUUUGUCAAGUUUGUCCUUGCACUGGCGCUGCAUUUGACGCAUC ......(((((((....((((((.((((((((((((.....((....))((((((......))).))).)))))))).((.((((......)))))))))).))))))..))))))).. ( -42.20) >DroWil_CAF1 353 119 - 1 UCUCCUUGCGACGCGAUUGCAACUCCUGUUCAAGUUCAUCAGCCUUGGCUGAAGCCUGUUUCCUUUCGUAAUUGCAUUUGUCGACUUUAUCUUUGCAUUGACGCUGCAUCUGCCUUAAC ......((((.((((((((((((....)))((.(((..(((((....)))))))).)).........))))))))....(((((.............)))))).))))........... ( -24.52) >DroAna_CAF1 354 119 - 1 ACUCCUUGCGCCUGGACUGCAGCUCCUGCUCCAGCUCCUCCGCCUUGGCAGCGGCCUGCUUCCGCUCGUAGUUGGACUUGUCCACUUUGUCCUUGCACUGGCGCUGCAUCUGGCGCAGC .....(((((((.(((.((((((.((((((((((((.(...((....))(((((.......))))).).)))))))...(..(.....)..)..)))..)).)))))))))))))))). ( -45.60) >DroPer_CAF1 357 119 - 1 CCUCUUUGCGACGGGACUGCAGCUCCAGUUCCAGUUCCUCGGCCUUGGCUGAGGCCUGUUUCCGUUCGUAAUUGGAUUUGUCCAUCUUGUCCUUGCACUGACGCUGCAUUUGCCGCAGC .....(((((.((((..((((((..((((..(((..(((((((....))))))).))).........((((..((((..(.....)..))))))))))))..)))))))))).))))). ( -41.30) >consensus CCUCCUUGCGACGGGACUGCAGCUCCAGUUCCAGUUCCUCAGCCUUGGCCGAGGCCUGUUUCCGUUCGUAAUUGGAUUUGUCCACUUUGUCCUUGCACUGACGCUGCAUCUGCCGCAGC .....((((((((.((.((((((..(((...(((.....((((....))))....))).........((((..((((..(.....)..)))))))).)))..)))))))))))))))). (-25.05 = -25.58 + 0.53)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:30:11 2006