
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 2,455,471 – 2,455,591 |
| Length | 120 |
| Max. P | 0.999991 |

| Location | 2,455,471 – 2,455,591 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.67 |
| Mean single sequence MFE | -31.10 |
| Consensus MFE | -28.25 |
| Energy contribution | -28.83 |
| Covariance contribution | 0.59 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.93 |
| Structure conservation index | 0.91 |
| SVM decision value | 3.75 |
| SVM RNA-class probability | 0.999581 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 2455471 120 + 27905053 AGAUUCCGAACAUUUGUUACCCCAAACAAAAUGGCUACACAGCCAUUUCCGACGAAUUUCCACAUGGAAGGCUUUACACCCAUCUCAGCGCUUUUGUUUGGAUUAUUAUAUUCGCGUUUC ............((((((......))))))((((((....))))))...((.(((((.((((.(..(((((((.............))).))))..).)))).......))))))).... ( -24.92) >DroSec_CAF1 14468 120 + 1 AGAUUCCGAACAUUUGUUACCCCAAACAAAAUGGCUACACAGCCAUUUCCGCCGAAUUUCCACAUGGAAGGCUUUACACCCAUCUCGGCGCUUUUGUUUGGAUUAUUAUAUUCGCGUUUC .((..(((((.((....((..(((((((((((((((....)))))....((((((..((((....))))((........))...)))))).))))))))))..))..)).)))).)..)) ( -33.80) >DroSim_CAF1 11187 120 + 1 AGAUUCCGAACAUUUGUUACCCCAAACAAAAUGGCUACACAGCCAUUUCCGCCGAAUUUCCACAUGGAAGGCUUUACACCCAUCUCGGCGCUUUUGUUUGGAUUAUUAUAUUCGCGUUUC .((..(((((.((....((..(((((((((((((((....)))))....((((((..((((....))))((........))...)))))).))))))))))..))..)).)))).)..)) ( -33.80) >DroEre_CAF1 15428 120 + 1 AGAUUCCGAACAUUUGUUACCCCGAACAAAAUGGCUACACAGCCAUUUCCGCCGAAUUUCCACAUGGAAGGCUUUACACCCAUCUCGGCGCUUUUGUUUGGAUUAUUAUAUUCGCGUUUC .((..(((((.((....((..(((((((((((((((....)))))....((((((..((((....))))((........))...)))))).))))))))))..))..)).)))).)..)) ( -33.50) >DroYak_CAF1 15286 120 + 1 AGAUUCCGAACAUUUGUUACCCCAAACAAAAUGGCUACACAGCCAUUUCCGCUGAAUUUCCACAUGGAAGGCUUUACACCCAUCGCGGCGCUUUUGUUUGGAUUAUUAUAUUCGCGUUUC .((..(((((.((....((..(((((((((((((((....)))))....(((((...((((....))))((........))....))))).))))))))))..))..)).)))).)..)) ( -30.10) >DroAna_CAF1 14796 120 + 1 GCAGACCAAACAUUUGUUACCCCAAACAAAAUGGCUACAUAGCCAUUUCCGCCAAAUUUCCACAUGGAAGGCUUUACACCCAUCUCGGCGCUUUUGUUUGGAUUAUUAUAUUCGCGUUUC ((......(((....)))...(((((((((((((((....)))))....((((....((((....))))((........)).....)))).))))))))))............))..... ( -30.50) >consensus AGAUUCCGAACAUUUGUUACCCCAAACAAAAUGGCUACACAGCCAUUUCCGCCGAAUUUCCACAUGGAAGGCUUUACACCCAUCUCGGCGCUUUUGUUUGGAUUAUUAUAUUCGCGUUUC ....((((((((((((((......))))))((((((....))))))...((((((..((((....))))((........))...))))))....)))))))).................. (-28.25 = -28.83 + 0.59)

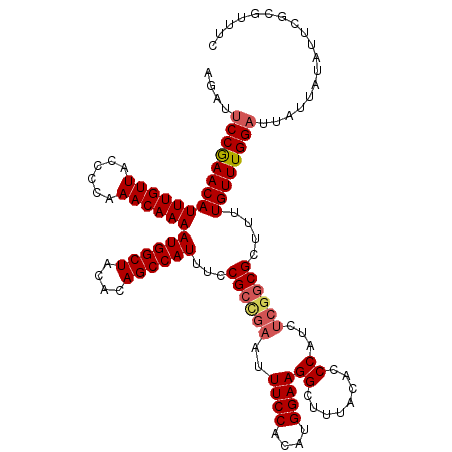
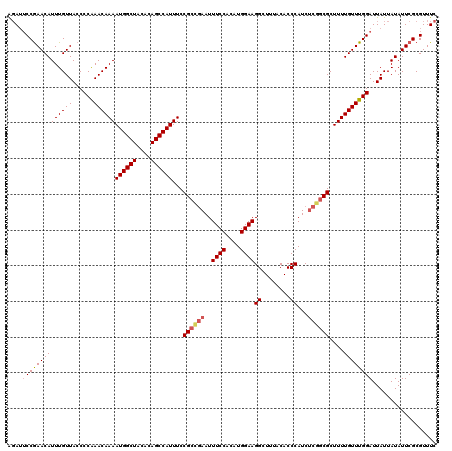
| Location | 2,455,471 – 2,455,591 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.67 |
| Mean single sequence MFE | -38.90 |
| Consensus MFE | -38.24 |
| Energy contribution | -38.52 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.08 |
| Mean z-score | -3.23 |
| Structure conservation index | 0.98 |
| SVM decision value | 5.65 |
| SVM RNA-class probability | 0.999991 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 2455471 120 - 27905053 GAAACGCGAAUAUAAUAAUCCAAACAAAAGCGCUGAGAUGGGUGUAAAGCCUUCCAUGUGGAAAUUCGUCGGAAAUGGCUGUGUAGCCAUUUUGUUUGGGGUAACAAAUGUUCGGAAUCU ......(((((((.....(((((((...........(((((((.....)))((((....))))...)))).(((((((((....)))))))))))))))).......)))))))...... ( -34.30) >DroSec_CAF1 14468 120 - 1 GAAACGCGAAUAUAAUAAUCCAAACAAAAGCGCCGAGAUGGGUGUAAAGCCUUCCAUGUGGAAAUUCGGCGGAAAUGGCUGUGUAGCCAUUUUGUUUGGGGUAACAAAUGUUCGGAAUCU ......(((((((.....(((((((.....(((((((...(((.....)))((((....)))).)))))))(((((((((....)))))))))))))))).......)))))))...... ( -42.40) >DroSim_CAF1 11187 120 - 1 GAAACGCGAAUAUAAUAAUCCAAACAAAAGCGCCGAGAUGGGUGUAAAGCCUUCCAUGUGGAAAUUCGGCGGAAAUGGCUGUGUAGCCAUUUUGUUUGGGGUAACAAAUGUUCGGAAUCU ......(((((((.....(((((((.....(((((((...(((.....)))((((....)))).)))))))(((((((((....)))))))))))))))).......)))))))...... ( -42.40) >DroEre_CAF1 15428 120 - 1 GAAACGCGAAUAUAAUAAUCCAAACAAAAGCGCCGAGAUGGGUGUAAAGCCUUCCAUGUGGAAAUUCGGCGGAAAUGGCUGUGUAGCCAUUUUGUUCGGGGUAACAAAUGUUCGGAAUCU ......(((((((.....(((.(((.....(((((((...(((.....)))((((....)))).)))))))(((((((((....)))))))))))).))).......)))))))...... ( -38.30) >DroYak_CAF1 15286 120 - 1 GAAACGCGAAUAUAAUAAUCCAAACAAAAGCGCCGCGAUGGGUGUAAAGCCUUCCAUGUGGAAAUUCAGCGGAAAUGGCUGUGUAGCCAUUUUGUUUGGGGUAACAAAUGUUCGGAAUCU ......(((((((.....(((((((((((...((((....(((.....)))((((....)))).....))))...(((((....)))))))))))))))).......)))))))...... ( -37.40) >DroAna_CAF1 14796 120 - 1 GAAACGCGAAUAUAAUAAUCCAAACAAAAGCGCCGAGAUGGGUGUAAAGCCUUCCAUGUGGAAAUUUGGCGGAAAUGGCUAUGUAGCCAUUUUGUUUGGGGUAACAAAUGUUUGGUCUGC ...((.(((((((.....(((((((.....(((((((...(((.....)))((((....)))).)))))))(((((((((....)))))))))))))))).......))))))))).... ( -38.60) >consensus GAAACGCGAAUAUAAUAAUCCAAACAAAAGCGCCGAGAUGGGUGUAAAGCCUUCCAUGUGGAAAUUCGGCGGAAAUGGCUGUGUAGCCAUUUUGUUUGGGGUAACAAAUGUUCGGAAUCU ......(((((((.....(((((((.....(((((((...(((.....)))((((....)))).)))))))(((((((((....)))))))))))))))).......)))))))...... (-38.24 = -38.52 + 0.28)

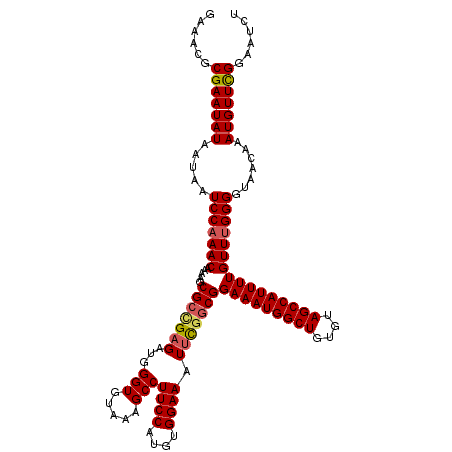
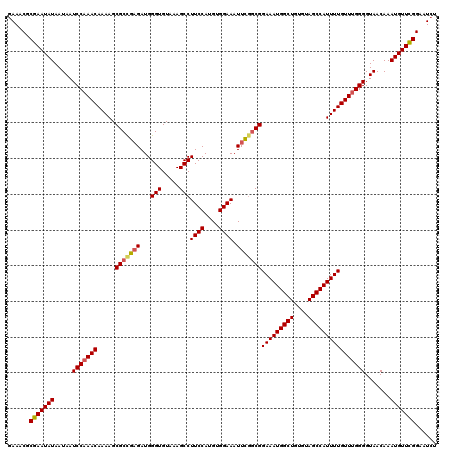
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:49:07 2006