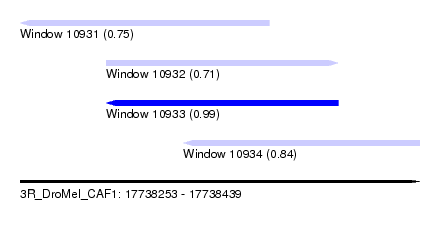
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 17,738,253 – 17,738,439 |
| Length | 186 |
| Max. P | 0.992343 |

| Location | 17,738,253 – 17,738,369 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 74.66 |
| Mean single sequence MFE | -41.55 |
| Consensus MFE | -19.86 |
| Energy contribution | -18.54 |
| Covariance contribution | -1.33 |
| Combinations/Pair | 1.48 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.48 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.751439 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17738253 116 - 27905053 UGGGCAUACUUGUGUGCCCAUUUUGAUGCAGAAGCCCAGCAACGCUCUCCACGUAAGCAGAGCUACUUUUACCGGCAGGCUUGAUUAGUUGGGCUUUACCAGAUCUAUUUAGUAGG ((((((((....))))))))..........((((((((((...(((((...(....).)))))........((....))........))))))))))................... ( -37.40) >DroSec_CAF1 27338 116 - 1 UGGGCAUACUUGUGUGCCCAUUUUGAUGCAGAAGCCCAGCAACGCUUUCCACGUAAGCAGAGCUACUUUCACCGAUAGGCUUGAUUAGUUGGGCUUUACCAGAUUUAUUUAGAAGG ((((((((....))))))))..........((((((((((...((((.......)))).(((((.............))))).....))))))))))................... ( -36.32) >DroSim_CAF1 27639 116 - 1 UGGGCAUACUUGUGUGCCCAUUUUGAUGCAGAAGCCCAGCAACGCUUUCCACGUAAGCAGAGCUACUUCCACCGACAGGCUUGAUUAGUUGGGCUUUACCAGAUUUAUCUAGAAGG ((((((((....))))))))..........((((((((((...((((.......)))).(((((...((....))..))))).....))))))))))................... ( -36.60) >DroEre_CAF1 29680 116 - 1 UGGGCACUCUUGUGUGCCCAUUUUGCUGCAGGAACCCAGCCACGCUGUGCACGUAAGCAGCGCUACUUUGACCGACAUUCCGGAGUGGUUGGACUGCUCCAGCUCUAUUCAAAAGG (((((((......)))))))....((((.((.(..(((((((((((((........)))))..........(((......))).))))))))..).)).))))............. ( -46.30) >DroYak_CAF1 29255 116 - 1 UGGGCAUACUUGGGUGCCCAUUUCGCUCCAGAAACGCAGCAACGCUGUUCACGUAGGCAGAGCUACUUUCACCUUUGGGCCUGAUUGGUUGGACUCUUCCAGAUCUAUUCAAAAGG (((((((......)))))))....((.(((((...(((((...)))))....((((......)))).......))))))).(((.((((((((....)))))..))).)))..... ( -31.90) >DroPer_CAF1 31137 116 - 1 GGGGCACGGGAGUGUGGCCCUUUUGGCCGAGAUACUCGGCAAUGCUGGCCGCAUAAGCCUGGCUCGUUUCUUGGGCGGCCUUCAGCGCCUGGACGUUGCCGGCCAAGUCCAAGAUG .((((.((((.((((((((......((((((...))))))......))))))))...))))))))...((((((((((((..(((((......)))))..))))..)))))))).. ( -60.80) >consensus UGGGCAUACUUGUGUGCCCAUUUUGAUGCAGAAACCCAGCAACGCUGUCCACGUAAGCAGAGCUACUUUCACCGACAGGCUUGAUUAGUUGGACUUUACCAGAUCUAUUCAAAAGG ((((((((....)))))))).................(((...(((.........)))...))).......((.........((((...(((......))))))).........)) (-19.86 = -18.54 + -1.33)

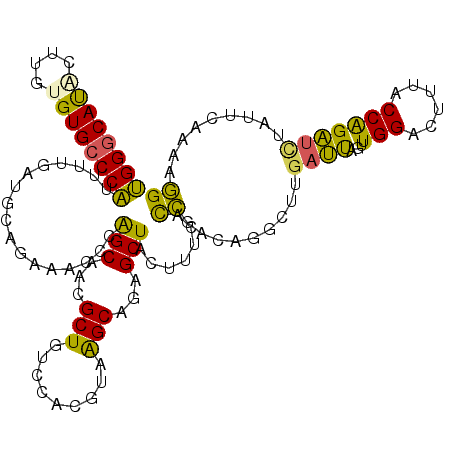

| Location | 17,738,293 – 17,738,401 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 74.52 |
| Mean single sequence MFE | -35.25 |
| Consensus MFE | -18.22 |
| Energy contribution | -17.87 |
| Covariance contribution | -0.35 |
| Combinations/Pair | 1.43 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.707335 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17738293 108 + 27905053 GCCGGUAAAAGUAGCUCUGCUUACGUGGAGAGCGUUGCUGGGCUUCUGCAUCAAAAUGGGCACACAAGUAUGCCCACAUUGCCACC---UGCAGAACCCGAAUAUAUCCUU ...(((((.....(((((.(......).))))).)))))(((.(((((((.(((..((((((........))))))..))).....---))))))))))............ ( -36.30) >DroSec_CAF1 27378 108 + 1 AUCGGUGAAAGUAGCUCUGCUUACGUGGAAAGCGUUGCUGGGCUUCUGCAUCAAAAUGGGCACACAAGUAUGCCCACAUUGUCACC---UGCAGAACCCGAAUAUAACCUU ...(((...((((((((..(....)..))....))))))(((.(((((((.(((..((((((........))))))..))).....---)))))))))).......))).. ( -35.50) >DroSim_CAF1 27679 108 + 1 GUCGGUGGAAGUAGCUCUGCUUACGUGGAAAGCGUUGCUGGGCUUCUGCAUCAAAAUGGGCACACAAGUAUGCCCACAUUGCCACC---AGCAGAACCCGAAUAUAACCUU ((.(((((.((.(((((.((..((((.....)))).)).))))).))....(((..((((((........))))))..))))))))---.))................... ( -34.30) >DroEre_CAF1 29720 111 + 1 GUCGGUCAAAGUAGCGCUGCUUACGUGCACAGCGUGGCUGGGUUCCUGCAGCAAAAUGGGCACACAAGAGUGCCCACAUUGCCAACAACACCAGCUCCAGAAAAUAACCUU ...(((.......((((((..........))))))(((((((((..((..((((..(((((((......)))))))..))))))..))).))))))..........))).. ( -38.40) >DroYak_CAF1 29295 111 + 1 AAAGGUGAAAGUAGCUCUGCCUACGUGAACAGCGUUGCUGCGUUUCUGGAGCGAAAUGGGCACCCAAGUAUGCCCACAUGGCCAAUAGCUCCAGCGCCCGAAAAUAACCUU .(((((...((((((.(((..........))).))))))(((...(((((((....((((((........)))))).((.....)).)))))))))).........))))) ( -34.70) >DroPer_CAF1 31177 105 + 1 GCCCAAGAAACGAGCCAGGCUUAUGCGGCCAGCAUUGCCGAGUAUCUCGGCCAAAAGGGCCACACUCCCGUGCCCCCACUGCCCACGACUGUGGCACCG-AAAGUG----- ((((.........((..((((.....)))).))...((((((...)))))).....))))..((((..((((((...((...........)))))).))-..))))----- ( -32.30) >consensus GUCGGUGAAAGUAGCUCUGCUUACGUGGACAGCGUUGCUGGGCUUCUGCAUCAAAAUGGGCACACAAGUAUGCCCACAUUGCCAAC___UGCAGAACCCGAAAAUAACCUU ...(((((..((((....((((((((.....))))....))))..)))).......((((((........))))))..)))))............................ (-18.22 = -17.87 + -0.35)
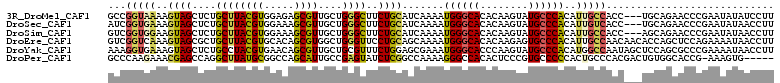



| Location | 17,738,293 – 17,738,401 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 74.52 |
| Mean single sequence MFE | -42.19 |
| Consensus MFE | -23.27 |
| Energy contribution | -24.64 |
| Covariance contribution | 1.37 |
| Combinations/Pair | 1.34 |
| Mean z-score | -2.56 |
| Structure conservation index | 0.55 |
| SVM decision value | 2.32 |
| SVM RNA-class probability | 0.992343 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17738293 108 - 27905053 AAGGAUAUAUUCGGGUUCUGCA---GGUGGCAAUGUGGGCAUACUUGUGUGCCCAUUUUGAUGCAGAAGCCCAGCAACGCUCUCCACGUAAGCAGAGCUACUUUUACCGGC ..((........((((((((((---.....(((.(((((((((....))))))))).))).))))).)))))......(((((...(....).)))))........))... ( -41.60) >DroSec_CAF1 27378 108 - 1 AAGGUUAUAUUCGGGUUCUGCA---GGUGACAAUGUGGGCAUACUUGUGUGCCCAUUUUGAUGCAGAAGCCCAGCAACGCUUUCCACGUAAGCAGAGCUACUUUCACCGAU ..(((.......((((((((((---.....(((.(((((((((....))))))))).))).))))).)))))(((...((((.......))))...)))......)))... ( -39.60) >DroSim_CAF1 27679 108 - 1 AAGGUUAUAUUCGGGUUCUGCU---GGUGGCAAUGUGGGCAUACUUGUGUGCCCAUUUUGAUGCAGAAGCCCAGCAACGCUUUCCACGUAAGCAGAGCUACUUCCACCGAC ..(((........(((((((((---.(((((((.(((((((((....))))))))).))).....(((((........)))))))))...)))))))))......)))... ( -41.14) >DroEre_CAF1 29720 111 - 1 AAGGUUAUUUUCUGGAGCUGGUGUUGUUGGCAAUGUGGGCACUCUUGUGUGCCCAUUUUGCUGCAGGAACCCAGCCACGCUGUGCACGUAAGCAGCGCUACUUUGACCGAC ..(((((.......(.(((((.(((.(((((((.((((((((......)))))))).))))..))).))))))))).((((((........))))))......)))))... ( -48.10) >DroYak_CAF1 29295 111 - 1 AAGGUUAUUUUCGGGCGCUGGAGCUAUUGGCCAUGUGGGCAUACUUGGGUGCCCAUUUCGCUCCAGAAACGCAGCAACGCUGUUCACGUAGGCAGAGCUACUUUCACCUUU (((((.........((((((((((..........((((((((......))))))))...)))))))...)))(((....(((((......))))).)))......))))). ( -35.32) >DroPer_CAF1 31177 105 - 1 -----CACUUU-CGGUGCCACAGUCGUGGGCAGUGGGGGCACGGGAGUGUGGCCCUUUUGGCCGAGAUACUCGGCAAUGCUGGCCGCAUAAGCCUGGCUCGUUUCUUGGGC -----..((..-((.((((((....)).)))).))..))..((((.((((((((......((((((...))))))......))))))))...))))(((((.....))))) ( -47.40) >consensus AAGGUUAUAUUCGGGUGCUGCA___GGUGGCAAUGUGGGCAUACUUGUGUGCCCAUUUUGAUGCAGAAACCCAGCAACGCUGUCCACGUAAGCAGAGCUACUUUCACCGAC ..(((........((((((((.....(((.(((.(((((((((....))))))))).))).)))............(((.......)))..))))))))......)))... (-23.27 = -24.64 + 1.37)

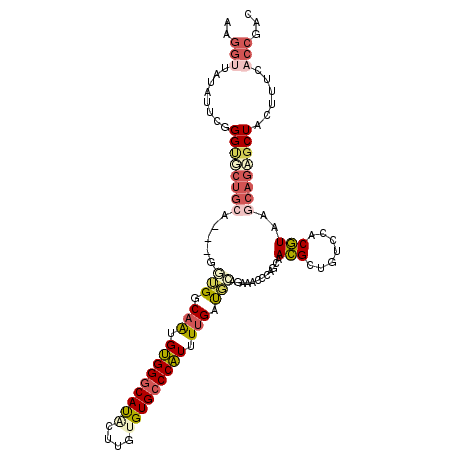

| Location | 17,738,329 – 17,738,439 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 71.30 |
| Mean single sequence MFE | -37.17 |
| Consensus MFE | -15.13 |
| Energy contribution | -15.58 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.41 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.41 |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.840797 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17738329 110 - 27905053 AGC--ACAUAGUCCACAAUAAAAGCAUCGUAUCCUUCCGAAAGGAUAUAUUC-----GGGUUCUGCA---GGUGGCAAUGUGGGCAUACUUGUGUGCCCAUUUUGAUGCAGAAGCCCAGC ...--..................((...((((((((....))))))))....-----((((((((((---.....(((.(((((((((....))))))))).))).))))).))))).)) ( -41.20) >DroSec_CAF1 27414 110 - 1 UGC--ACGUAGUCCACAAUAAAAGCAUCGUAUCCCUCCGAAAGGUUAUAUUC-----GGGUUCUGCA---GGUGACAAUGUGGGCAUACUUGUGUGCCCAUUUUGAUGCAGAAGCCCAGC ...--..................((...........((....))........-----((((((((((---.....(((.(((((((((....))))))))).))).))))).))))).)) ( -36.10) >DroSim_CAF1 27715 110 - 1 UGC--ACAUAGUCCACAAUAAAAGCAUCGUGUCCCUCCGAAAGGUUAUAUUC-----GGGUUCUGCU---GGUGGCAAUGUGGGCAUACUUGUGUGCCCAUUUUGAUGCAGAAGCCCAGC .((--.....(..(((.((......)).)))..)..((....))........-----(((((((((.---.....(((.(((((((((....))))))))).)))..)))).))))).)) ( -35.70) >DroEre_CAF1 29756 113 - 1 AGC--ACAUAGUCCACAAUGAAAGCGUCGUAUCCUUGCGAAAGGUUAUUUUC-----UGGAGCUGGUGUUGUUGGCAAUGUGGGCACUCUUGUGUGCCCAUUUUGCUGCAGGAACCCAGC ...--......((((.(((((.....(((((....)))))....)))))...-----))))(((((.(((.(((((((.((((((((......)))))))).))))..))).)))))))) ( -43.30) >DroYak_CAF1 29331 113 - 1 AGC--ACAUAGUCCACAAUGAUGGCAUCGUAUUCAUCCGAAAGGUUAUUUUC-----GGGCGCUGGAGCUAUUGGCCAUGUGGGCAUACUUGGGUGCCCAUUUCGCUCCAGAAACGCAGC .((--.....((((.....(((((........))))).(((((....)))))-----)))).(((((((..........((((((((......))))))))...)))))))....))... ( -37.02) >DroAna_CAF1 29609 117 - 1 ACCCGAAAUUAUACGUUCUGAAAUGGCCUGAUUCA---CAAAAUUUUCAUUCUCACUAGGGGCCACCGUGGUGGGUAGAGUGGGCAAAGGUGUGUGACCGCCAAAUUCGAGGAAAUUAGA .((((((..................((((.((((.---.............(((((((.((....)).)))))))..))))))))...((((......))))...)))).))........ ( -29.69) >consensus AGC__ACAUAGUCCACAAUAAAAGCAUCGUAUCCAUCCGAAAGGUUAUAUUC_____GGGUGCUGCA___GGUGGCAAUGUGGGCAUACUUGUGUGCCCAUUUUGAUGCAGAAACCCAGC .......................(((..........((....))...............................(((.(((((((((....))))))))).))).)))........... (-15.13 = -15.58 + 0.45)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:28:35 2006