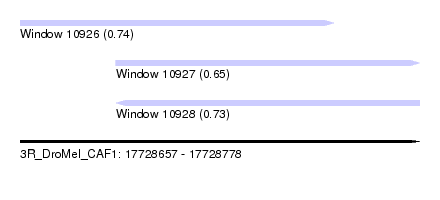
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 17,728,657 – 17,728,778 |
| Length | 121 |
| Max. P | 0.740211 |

| Location | 17,728,657 – 17,728,752 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 71.01 |
| Mean single sequence MFE | -23.30 |
| Consensus MFE | -7.09 |
| Energy contribution | -8.82 |
| Covariance contribution | 1.73 |
| Combinations/Pair | 1.32 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.30 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.740211 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17728657 95 + 27905053 UAAAAAAC--UGUCAGGCUGGCAGAGUUCUUU-UGGCCAGGCGAAUUGGAUUUAUU-UAGCCAAUGUUU-UUCCAAUUCAUAUAU----GAAUGUAUUGGCCAA- .......(--((((.....))))).......(-(((((((..((((((((..((((-.....))))...-.))))))))((((..----..)))).))))))))- ( -25.90) >DroVir_CAF1 20152 88 + 1 --AAAAACGCUGGCAUACUGACAAAAGGCAGCCCGGCUG------CUGCACAUAUUUUAGCCAAC--UU-UU------AAAAUAUGUUGCCAUUUGUUGGCCAAG --........((((.....((((((.((((((...))))------))(((((((((((((.....--..-))------)))))))).)))..)))))).)))).. ( -26.10) >DroSec_CAF1 17771 91 + 1 UAAAAAAC--UGACAGGCUGGUGGAGUUCUUU-UGGCUAGUCGAAUUGGAUUUAUU-UAGCCAAUGUUU-UUCCAAUUUAUAUAU--------GUAUUGGCCAA- ........--.....(((..((((((.....(-(((((((..((((...))))..)-))))))).....-)))))...(((....--------))))..)))..- ( -21.30) >DroSim_CAF1 18045 91 + 1 UAAAAAAC--UAGCAGGCUGGUGGAGUUCUUU-UGGCCAGUCGAAUUGGAUUUAUU-UAGCCAAUGUUU-UUCCAAUUCAUAUAU--------GUAUUGGCCAA- ........--..((.(((((((.(((....))-).)))))))((((((((..((((-.....))))...-.))))))))......--------......))...- ( -24.80) >DroEre_CAF1 20188 81 + 1 ------------GCAGGCUGGUAGAGUUCUUU-UGGCCAGGCAAAUUGGAUUUAUU-UAGCCCAUGUUU-UGCCAAUUCACAUAU--------GUAUUGGCCAA- ------------...(((..(((.(......(-((((.(((((...(((.......-....))))))))-.)))))........)--------.)))..)))..- ( -22.14) >DroYak_CAF1 19312 92 + 1 UAAAAAAC--UGGCAGGCUGUUAGAGUUCCUU-UAACCAGGCGAAUUGAAUUUAUU-UAGUCCAUGUUUUUUCCAAUUCAUACAU--------GUACUGGCCAA- ........--((((.(.(((((((((...)))-))).))).)..............-((((.(((((..............))))--------).)))))))).- ( -19.54) >consensus UAAAAAAC__UGGCAGGCUGGUAGAGUUCUUU_UGGCCAGGCGAAUUGGAUUUAUU_UAGCCAAUGUUU_UUCCAAUUCAUAUAU________GUAUUGGCCAA_ ...............((((((((..(((......))).....((((((((.....................))))))))...............))))))))... ( -7.09 = -8.82 + 1.73)

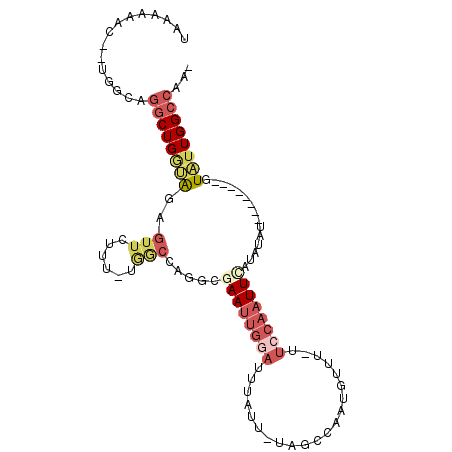
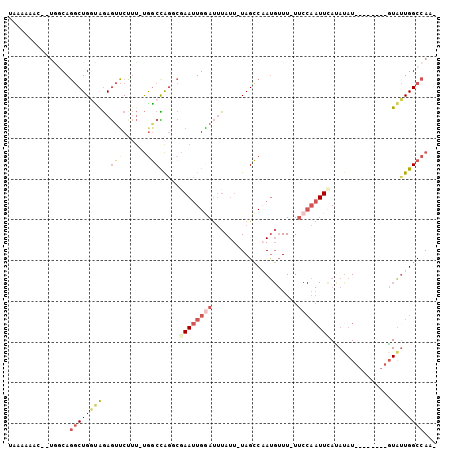
| Location | 17,728,686 – 17,728,778 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 84.56 |
| Mean single sequence MFE | -23.23 |
| Consensus MFE | -15.31 |
| Energy contribution | -17.71 |
| Covariance contribution | 2.40 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.646250 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17728686 92 + 27905053 UUGGCCAGGCGAAUUGGAUUUAUUUAGCCAAUGUUU-UUCCAAUUCAUAUAUGAAUGUAUUGGCCAACGGGCCACAGCUGCUGUUGCACUUGC .(((((.(((((((((((..((((.....))))...-.))))))))(((((....)))))..)))....)))))(((.(((....))).))). ( -26.70) >DroSec_CAF1 17800 88 + 1 UUGGCUAGUCGAAUUGGAUUUAUUUAGCCAAUGUUU-UUCCAAUUUAUAUAU----GUAUUGGCCAACGGGCAACAGCUGCUGCUGCACUUGC ((((((((((((((((((..((((.....))))...-.))))))))......----).)))))))))...((((((((....))))...)))) ( -21.70) >DroSim_CAF1 18074 88 + 1 UUGGCCAGUCGAAUUGGAUUUAUUUAGCCAAUGUUU-UUCCAAUUCAUAUAU----GUAUUGGCCAACGGGCAACAGCUGCUGUUGCACUUGC ((((((((((((((((((..((((.....))))...-.))))))))......----).)))))))))...(((((((...)))))))...... ( -27.70) >DroEre_CAF1 20207 74 + 1 UUGGCCAGGCAAAUUGGAUUUAUUUAGCCCAUGUUU-UGCCAAUUCACAUAU----GUAUUGGCCAACGGGCAACAGCU-------------- (((((((((((((.(((...........)))...))-)))).....((....----))..)))))))..(((....)))-------------- ( -21.30) >DroYak_CAF1 19341 89 + 1 UUAACCAGGCGAAUUGAAUUUAUUUAGUCCAUGUUUUUUCCAAUUCAUACAU----GUACUGGCCAACGGGCAACAGCUGCUGUUGCACUUGC ....((.(((((((...))))...((((.(((((..............))))----).)))))))...))(((((((...)))))))...... ( -18.74) >consensus UUGGCCAGGCGAAUUGGAUUUAUUUAGCCAAUGUUU_UUCCAAUUCAUAUAU____GUAUUGGCCAACGGGCAACAGCUGCUGUUGCACUUGC ((((((((..((((((((....................)))))))).(((......)))))))))))...(((((((...)))))))...... (-15.31 = -17.71 + 2.40)

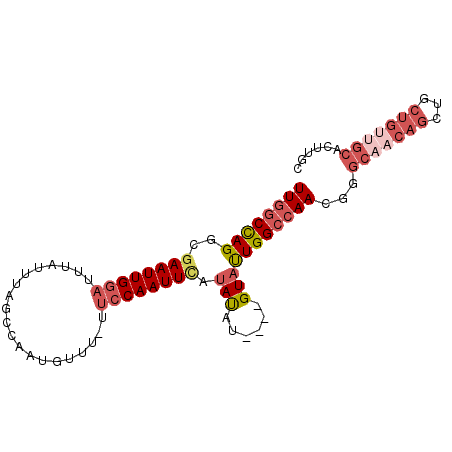
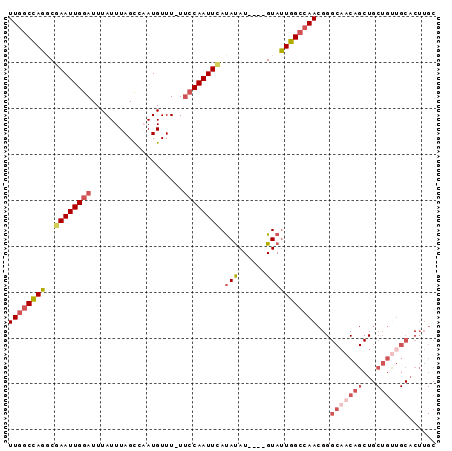
| Location | 17,728,686 – 17,728,778 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 84.56 |
| Mean single sequence MFE | -21.64 |
| Consensus MFE | -14.85 |
| Energy contribution | -16.01 |
| Covariance contribution | 1.16 |
| Combinations/Pair | 1.28 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.728088 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17728686 92 - 27905053 GCAAGUGCAACAGCAGCUGUGGCCCGUUGGCCAAUACAUUCAUAUAUGAAUUGGAA-AAACAUUGGCUAAAUAAAUCCAAUUCGCCUGGCCAA (((..(((....)))..)))((((.(((..(((((.(((......))).)))))..-.)))...(((................))).)))).. ( -23.69) >DroSec_CAF1 17800 88 - 1 GCAAGUGCAGCAGCAGCUGUUGCCCGUUGGCCAAUAC----AUAUAUAAAUUGGAA-AAACAUUGGCUAAAUAAAUCCAAUUCGACUAGCCAA ....(.(((((((...))))))).)(((..(((((..----........)))))..-.))).(((((((.................))))))) ( -20.53) >DroSim_CAF1 18074 88 - 1 GCAAGUGCAACAGCAGCUGUUGCCCGUUGGCCAAUAC----AUAUAUGAAUUGGAA-AAACAUUGGCUAAAUAAAUCCAAUUCGACUGGCCAA ....(.(((((((...))))))).).((((((((((.----...)))((((((((.-..................))))))))...))))))) ( -24.41) >DroEre_CAF1 20207 74 - 1 --------------AGCUGUUGCCCGUUGGCCAAUAC----AUAUGUGAAUUGGCA-AAACAUGGGCUAAAUAAAUCCAAUUUGCCUGGCCAA --------------.((....))...(((((((.(((----....)))....((((-((...(((..(.....)..))).))))))))))))) ( -20.20) >DroYak_CAF1 19341 89 - 1 GCAAGUGCAACAGCAGCUGUUGCCCGUUGGCCAGUAC----AUGUAUGAAUUGGAAAAAACAUGGACUAAAUAAAUUCAAUUCGCCUGGUUAA ....(.(((((((...))))))).).((((((((((.----...)).((((((((....................))))))))..)))))))) ( -19.35) >consensus GCAAGUGCAACAGCAGCUGUUGCCCGUUGGCCAAUAC____AUAUAUGAAUUGGAA_AAACAUUGGCUAAAUAAAUCCAAUUCGCCUGGCCAA ......(((((((...)))))))...((((((((((........)))((((((((....................))))))))...))))))) (-14.85 = -16.01 + 1.16)

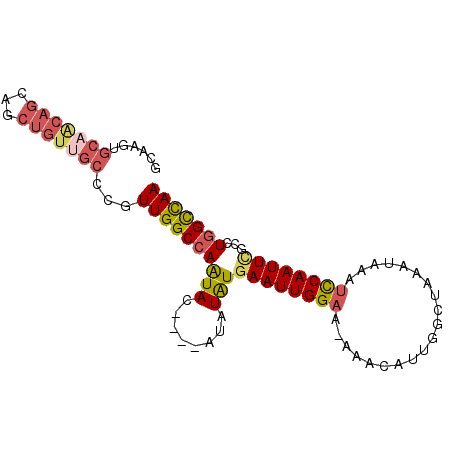

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:28:29 2006