
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 17,598,325 – 17,598,472 |
| Length | 147 |
| Max. P | 0.999936 |

| Location | 17,598,325 – 17,598,432 |
|---|---|
| Length | 107 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 85.55 |
| Mean single sequence MFE | -15.53 |
| Consensus MFE | -13.00 |
| Energy contribution | -13.00 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.652765 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17598325 107 + 27905053 ------UACACGAUUAUUA-UAUGUAACAUAUAACAGCUACAAAAACUACAAAACUUCCUUACUACCAGUCUUACUUUCGGUACGAACACCAAACACAAAGUGGCUGUCGUACA ------...((((...(((-((((...)))))))(((((((..........................((.....))...(((......))).........)))))))))))... ( -17.60) >DroVir_CAF1 822 101 + 1 AAGUAUUAUACGAUUAU----A----U-----AACAGCUACAAAAACUACAAAACUUCCUUACUACCAGUCUUACUUUCGGUACGAACACCAAACACAAAGUGGCUGUCGUACA ................(----(----.-----.((((((((..........................((.....))...(((......))).........))))))))..)).. ( -14.80) >DroGri_CAF1 1135 110 + 1 AAGUACUAUACGACUAU----AUAUAUAUUAAAACAGCUACAAAAACUACAAAACUUCCUUACUACCAGUCUUACUUUCGGUACGAACACCAAACACAAAGUGGCUGUCGUACA ..((((((((......)----))).........((((((((..........................((.....))...(((......))).........)))))))).)))). ( -16.10) >DroEre_CAF1 767 107 + 1 ------UACACGAUUAUC-AUAUGUAACAUAUAACAGCUACAAAAACUACAAAACUUCCUUACUACCAGUCUUACUUUCGGUACGAACACCAAACACAAAGUGGCUGUCGUACA ------...((((.....-(((((...)))))..(((((((..........................((.....))...(((......))).........)))))))))))... ( -15.60) >DroAna_CAF1 698 100 + 1 ------AACGAGUUU--------ACAAUGUUUAACAGCUACAAAAACUACAAAACUUCCUUACUACCAGUCUUACUUUCGGUACGAACACCAAACACAAAGUGGCUGUCGUACA ------.........--------....(((...((((((((..........................((.....))...(((......))).........))))))))...))) ( -14.50) >DroMoj_CAF1 805 105 + 1 AAGUAAUAUACGAUUAU----GUGUUU-----AACAGCUACAAAAACUACAAAACUUCCUUACUACCAGUCUUACUUUCGGUACGAACACCAAACACAAAGUGGCUGUCGUACA ..(((((.....)))))----.(((..-----.((((((((..........................((.....))...(((......))).........))))))))...))) ( -14.60) >consensus ______UACACGAUUAU____AUGUAA__UAUAACAGCUACAAAAACUACAAAACUUCCUUACUACCAGUCUUACUUUCGGUACGAACACCAAACACAAAGUGGCUGUCGUACA .................................((((((((..........................((.....))...(((......))).........))))))))...... (-13.00 = -13.00 + 0.00)


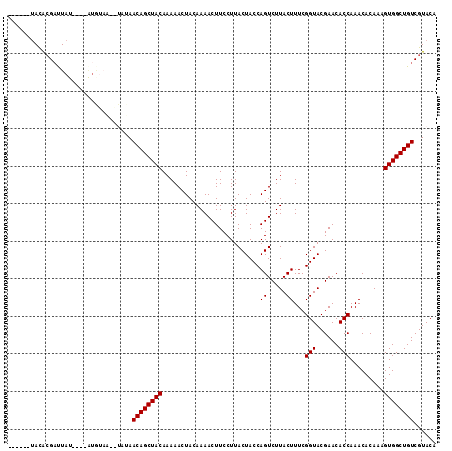
| Location | 17,598,352 – 17,598,472 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.44 |
| Mean single sequence MFE | -25.75 |
| Consensus MFE | -24.57 |
| Energy contribution | -24.73 |
| Covariance contribution | 0.17 |
| Combinations/Pair | 1.00 |
| Mean z-score | -4.96 |
| Structure conservation index | 0.95 |
| SVM decision value | 4.67 |
| SVM RNA-class probability | 0.999936 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17598352 120 + 27905053 CAGCUACAAAAACUACAAAACUUCCUUACUACCAGUCUUACUUUCGGUACGAACACCAAACACAAAGUGGCUGUCGUACAAAAAAUGAUAAAAAACGAGAGUAAGACAAAAACUAAAUCC ..................................((((((((((((((((((.(((((.........))).))))))))......(......)..))))))))))))............. ( -26.00) >DroVir_CAF1 843 120 + 1 CAGCUACAAAAACUACAAAACUUCCUUACUACCAGUCUUACUUUCGGUACGAACACCAAACACAAAGUGGCUGUCGUACAAAAAAUUAUAAAAAACGAGAGUAAGACAAAAACUAAAUCC ..................................((((((((((((((((((.(((((.........))).))))))))......((....))..))))))))))))............. ( -25.80) >DroGri_CAF1 1165 120 + 1 CAGCUACAAAAACUACAAAACUUCCUUACUACCAGUCUUACUUUCGGUACGAACACCAAACACAAAGUGGCUGUCGUACAAAAAAUGAUAAAAAACGAGAGUAAGACAAAAACUAAAUCU ..................................((((((((((((((((((.(((((.........))).))))))))......(......)..))))))))))))............. ( -26.00) >DroWil_CAF1 18271 120 + 1 CAGCUACAGAAACUACAAAACUUCCUUACUACCAGUCUUACUUUCGGUACGAACACCAAACACAAAGUUGCUGUCGUACAAAAAAUUAUAAAAAACGAGAGUAAGACAAAAACUAAGUCC ..................................((((((((((((((((((.((.(((........))).))))))))......((....))..))))))))))))............. ( -24.70) >DroAna_CAF1 718 120 + 1 CAGCUACAAAAACUACAAAACUUCCUUACUACCAGUCUUACUUUCGGUACGAACACCAAACACAAAGUGGCUGUCGUACAAAAAAUGAUAAAAAACGAGAGUAAGACAAAAACUAAAUCC ..................................((((((((((((((((((.(((((.........))).))))))))......(......)..))))))))))))............. ( -26.00) >DroMoj_CAF1 830 120 + 1 CAGCUACAAAAACUACAAAACUUCCUUACUACCAGUCUUACUUUCGGUACGAACACCAAACACAAAGUGGCUGUCGUACAAAAAAUGAUAAAAAACGAGAGUAAGACAAAAACUAAAUCC ..................................((((((((((((((((((.(((((.........))).))))))))......(......)..))))))))))))............. ( -26.00) >consensus CAGCUACAAAAACUACAAAACUUCCUUACUACCAGUCUUACUUUCGGUACGAACACCAAACACAAAGUGGCUGUCGUACAAAAAAUGAUAAAAAACGAGAGUAAGACAAAAACUAAAUCC ..................................((((((((((((((((((.(((((.........))).))))))))......(......)..))))))))))))............. (-24.57 = -24.73 + 0.17)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:27:38 2006