
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 17,378,362 – 17,378,464 |
| Length | 102 |
| Max. P | 0.942170 |

| Location | 17,378,362 – 17,378,464 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 85.10 |
| Mean single sequence MFE | -21.09 |
| Consensus MFE | -16.29 |
| Energy contribution | -16.10 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.28 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.32 |
| SVM RNA-class probability | 0.942170 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
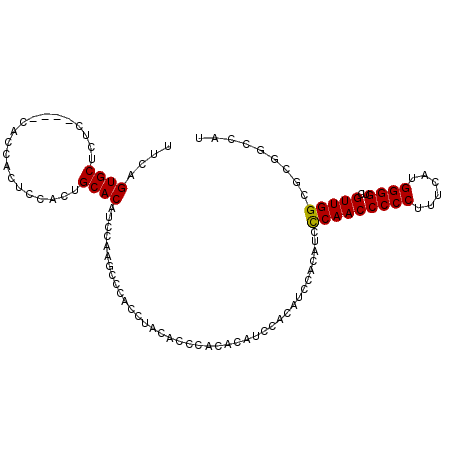
>3R_DroMel_CAF1 17378362 102 + 27905053 UUCAGUGCUCUC----CACCACUCCACUGCACAUCCAAGCCCACCUACACGCACACAUCCACAUCCACAUCCCAACCCCCAUUCAUGGGGUCGUUGGCGCGGCCAU ....((((....----............))))......((((........(....)...............((((((((((....)))))..))))).).)))... ( -19.79) >DroSec_CAF1 27375 94 + 1 UUCAGUGCUCUC------------CACUGCACAUCCAAGCCCACCUACACCCACUCAUCCACAUCGACAUCCCAACCCCCUUUCAUGGGGUCGUUGGCGCGGCCAU ....((((....------------....))))......((((...............((......))....(((((((((......))))..))))).).)))... ( -21.70) >DroSim_CAF1 36099 102 + 1 UUCAGUGCUCUC----CACCACUCCACUGCACAUCCAAGCCCACCUACACCCACACAUCCACAUCGACAUCCCAACCCCCUUUCAUGGGGUCGUUGGCGCGGCCAU ....((((....----............))))......((((...............((......))....(((((((((......))))..))))).).)))... ( -20.29) >DroYak_CAF1 43447 99 + 1 UUCAGUGC----CCACCAGCACUCCACUGCACAUCCAGGCU---CUACACCUCCACCUACACAUCCACAUCUCAACCCCCUUUCAUGGGGUCGUUGGCGCGGCCAU ....(.((----((.(((((......(((......)))...---................................((((......))))..))))).).)))).. ( -22.60) >consensus UUCAGUGCUCUC____CACCACUCCACUGCACAUCCAAGCCCACCUACACCCACACAUCCACAUCCACAUCCCAACCCCCUUUCAUGGGGUCGUUGGCGCGGCCAU ....((((....................)))).......................................(((((((((......))))..)))))......... (-16.29 = -16.10 + -0.19)


| Location | 17,378,362 – 17,378,464 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 85.10 |
| Mean single sequence MFE | -33.95 |
| Consensus MFE | -24.87 |
| Energy contribution | -25.93 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.95 |
| SVM RNA-class probability | 0.888442 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17378362 102 - 27905053 AUGGCCGCGCCAACGACCCCAUGAAUGGGGGUUGGGAUGUGGAUGUGGAUGUGUGCGUGUAGGUGGGCUUGGAUGUGCAGUGGAGUGGUG----GAGAGCACUGAA ....((((((((.(((((((.......))))))).....))).)))))...((..(((.((((....)))).)))..))....((((.(.----...).))))... ( -35.90) >DroSec_CAF1 27375 94 - 1 AUGGCCGCGCCAACGACCCCAUGAAAGGGGGUUGGGAUGUCGAUGUGGAUGAGUGGGUGUAGGUGGGCUUGGAUGUGCAGUG------------GAGAGCACUGAA ..((((.((((..(((((((.......)))))))..((..(.((........)).)..)).))))))))........(((((------------.....))))).. ( -32.10) >DroSim_CAF1 36099 102 - 1 AUGGCCGCGCCAACGACCCCAUGAAAGGGGGUUGGGAUGUCGAUGUGGAUGUGUGGGUGUAGGUGGGCUUGGAUGUGCAGUGGAGUGGUG----GAGAGCACUGAA .(.(((((.(((.(((((((.......)))))))..(((((......)))))(((.((.((((....)))).)).)))..))).))))).----)........... ( -31.90) >DroYak_CAF1 43447 99 - 1 AUGGCCGCGCCAACGACCCCAUGAAAGGGGGUUGAGAUGUGGAUGUGUAGGUGGAGGUGUAG---AGCCUGGAUGUGCAGUGGAGUGCUGGUGG----GCACUGAA ....((((.(((.(((((((.......))))))).....)))...((((.((..((((....---.))))..)).))))))))((((((....)----)))))... ( -35.90) >consensus AUGGCCGCGCCAACGACCCCAUGAAAGGGGGUUGGGAUGUCGAUGUGGAUGUGUGGGUGUAGGUGGGCUUGGAUGUGCAGUGGAGUGGUG____GAGAGCACUGAA ..((((.((((..(((((((.......)))))))..(..((.((........)).))..).))))))))........(((((.................))))).. (-24.87 = -25.93 + 1.06)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:25:40 2006