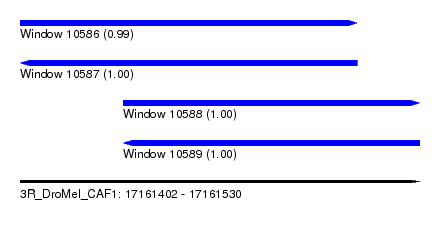

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 17,161,402 – 17,161,530 |
| Length | 128 |
| Max. P | 0.999953 |

| Location | 17,161,402 – 17,161,510 |
|---|---|
| Length | 108 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 91.41 |
| Mean single sequence MFE | -28.93 |
| Consensus MFE | -25.83 |
| Energy contribution | -26.31 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.89 |
| SVM decision value | 2.08 |
| SVM RNA-class probability | 0.987398 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17161402 108 + 27905053 CGACAAAAACUUGUACAGAGCAAAAUGAA-----UGGA-AGCCGGAGGAGAGGGCGUGGCAAAAAACGGAUUAAAUAUGUAUACAAUACCCGCCCUCGCCUCCAGCAGCACAGC ..........((((.....))))......-----....-.((.(((((.(((((((.((......(((.........)))........))))))))).))))).))........ ( -30.14) >DroSec_CAF1 70614 108 + 1 CGACAAAAACUUGUACAGAGCAAAAUGAA-----UGGA-GGCCGGAGGAGAGGGCGUGGCAAAAAACGAAUUAAAUAUGUAUACAAUACCCGCCCUCGCCUCCAGCAGCACAGC ..........((((.....))))......-----(((.-.((.(((((.(((((((.((.............................))))))))).))))).))..).)).. ( -31.05) >DroSim_CAF1 61919 108 + 1 CGACAAAAACUUGUACAGAGCAAAAUGAA-----UGGA-GGCCGGAGGAGAGGGCGUGGCAAAAAACGAAUUAAAUAUGUAUACAAUACCCGCCCUCGCCUCCAGCAGCACAGC ..........((((.....))))......-----(((.-.((.(((((.(((((((.((.............................))))))))).))))).))..).)).. ( -31.05) >DroEre_CAF1 61797 111 + 1 CGACAAAAACUUGUACAGAGCAAAAUGAACUGAAUGGA-AGCCGAAGGAGAGGGCGUGGCAAA-AAUGAAUUAAAUAUGAAUACAGUCCC-GCCCUCGCCUCCAGCAGCACAGC ..........((((.....))))..((..(((..((((-.(((....).(((((((.(((...-.....................))).)-)))))))).)))).)))..)).. ( -26.86) >DroYak_CAF1 89364 113 + 1 CGACAAAAACUUGUACAGAGCAAAAUGAACGGAAUGGAGAGCCAGAGGAGAGGGCGUGGCAAA-AAUGAAUUAAAUAUGAAUAUAACCCCUGCCCUCGCCUCCAGCAGCACAGC ..((((....)))).....((.............(((....)))((((.(((((((.((....-.....((.....)).........)).))))))).))))..))........ ( -25.57) >consensus CGACAAAAACUUGUACAGAGCAAAAUGAA_____UGGA_AGCCGGAGGAGAGGGCGUGGCAAAAAACGAAUUAAAUAUGUAUACAAUACCCGCCCUCGCCUCCAGCAGCACAGC ..((((....)))).....((..............((....))(((((.(((((((.((.............................))))))))).))))).))........ (-25.83 = -26.31 + 0.48)

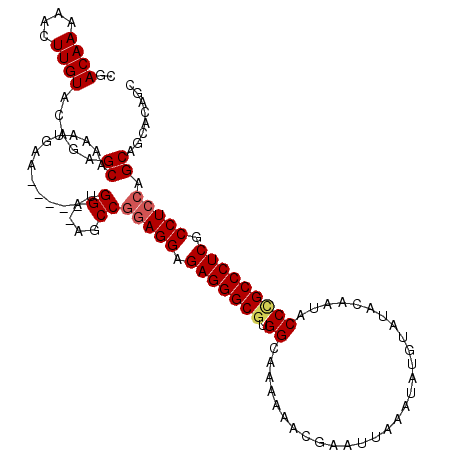

| Location | 17,161,402 – 17,161,510 |
|---|---|
| Length | 108 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 91.41 |
| Mean single sequence MFE | -36.55 |
| Consensus MFE | -32.30 |
| Energy contribution | -32.62 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.12 |
| Mean z-score | -3.08 |
| Structure conservation index | 0.88 |
| SVM decision value | 4.36 |
| SVM RNA-class probability | 0.999879 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17161402 108 - 27905053 GCUGUGCUGCUGGAGGCGAGGGCGGGUAUUGUAUACAUAUUUAAUCCGUUUUUUGCCACGCCCUCUCCUCCGGCU-UCCA-----UUCAUUUUGCUCUGUACAAGUUUUUGUCG ..(((((.((((((((.(((((((((((.........................)))).))))))).)))))))).-..((-----.......))....)))))........... ( -38.81) >DroSec_CAF1 70614 108 - 1 GCUGUGCUGCUGGAGGCGAGGGCGGGUAUUGUAUACAUAUUUAAUUCGUUUUUUGCCACGCCCUCUCCUCCGGCC-UCCA-----UUCAUUUUGCUCUGUACAAGUUUUUGUCG ..(((((.((((((((.(((((((((((.........................)))).))))))).)))))))).-..((-----.......))....)))))........... ( -38.41) >DroSim_CAF1 61919 108 - 1 GCUGUGCUGCUGGAGGCGAGGGCGGGUAUUGUAUACAUAUUUAAUUCGUUUUUUGCCACGCCCUCUCCUCCGGCC-UCCA-----UUCAUUUUGCUCUGUACAAGUUUUUGUCG ..(((((.((((((((.(((((((((((.........................)))).))))))).)))))))).-..((-----.......))....)))))........... ( -38.41) >DroEre_CAF1 61797 111 - 1 GCUGUGCUGCUGGAGGCGAGGGC-GGGACUGUAUUCAUAUUUAAUUCAUU-UUUGCCACGCCCUCUCCUUCGGCU-UCCAUUCAGUUCAUUUUGCUCUGUACAAGUUUUUGUCG ..(((((.((((((((.((((((-(((.(.....................-...))).))))))).)))))))).-.......(((.......)))..)))))........... ( -34.36) >DroYak_CAF1 89364 113 - 1 GCUGUGCUGCUGGAGGCGAGGGCAGGGGUUAUAUUCAUAUUUAAUUCAUU-UUUGCCACGCCCUCUCCUCUGGCUCUCCAUUCCGUUCAUUUUGCUCUGUACAAGUUUUUGUCG ..(((((.((..((((.((((((...(((.....................-...)))..)))))).))))..))..........((.......))...)))))........... ( -32.76) >consensus GCUGUGCUGCUGGAGGCGAGGGCGGGUAUUGUAUACAUAUUUAAUUCGUUUUUUGCCACGCCCUCUCCUCCGGCU_UCCA_____UUCAUUUUGCUCUGUACAAGUUUUUGUCG ..(((((.((((((((.(((((((((((.........................)))).))))))).))))))))........................)))))........... (-32.30 = -32.62 + 0.32)


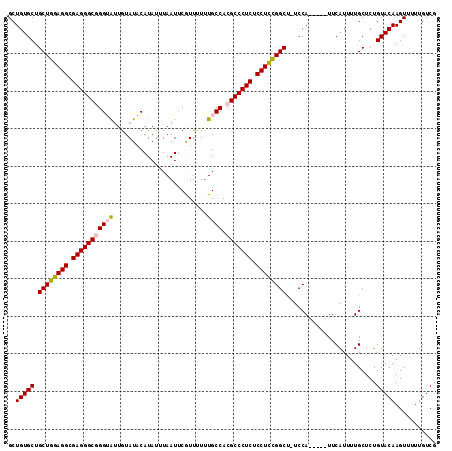
| Location | 17,161,435 – 17,161,530 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 93.18 |
| Mean single sequence MFE | -32.15 |
| Consensus MFE | -28.15 |
| Energy contribution | -28.63 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.05 |
| Mean z-score | -4.29 |
| Structure conservation index | 0.88 |
| SVM decision value | 4.34 |
| SVM RNA-class probability | 0.999874 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17161435 95 + 27905053 -AGCCGGAGGAGAGGGCGUGGCAAAAAACGGAUUAAAUAUGUAUACAAUACCCGCCCUCGCCUCCAGCAGCACAGCAAAAAGGCCAAUACUAUUUA -.((((((((.(((((((.((......(((.........)))........))))))))).))))).((......)).....)))............ ( -32.84) >DroSec_CAF1 70647 95 + 1 -GGCCGGAGGAGAGGGCGUGGCAAAAAACGAAUUAAAUAUGUAUACAAUACCCGCCCUCGCCUCCAGCAGCACAGCAAAAAGGCCAAUACUAUUUA -(((((((((.(((((((.((.............................))))))))).))))).((......)).....))))........... ( -36.75) >DroSim_CAF1 61952 95 + 1 -GGCCGGAGGAGAGGGCGUGGCAAAAAACGAAUUAAAUAUGUAUACAAUACCCGCCCUCGCCUCCAGCAGCACAGCAAAAAGGCCAAUACUAUUUA -(((((((((.(((((((.((.............................))))))))).))))).((......)).....))))........... ( -36.75) >DroEre_CAF1 61835 93 + 1 -AGCCGAAGGAGAGGGCGUGGCAAA-AAUGAAUUAAAUAUGAAUACAGUCCC-GCCCUCGCCUCCAGCAGCACAGCAAAAAGGCCAAUACUAUUUA -.((((.(((.(((((((.(((...-.....................))).)-)))))).))).).((......)).....)))............ ( -27.26) >DroYak_CAF1 89402 95 + 1 GAGCCAGAGGAGAGGGCGUGGCAAA-AAUGAAUUAAAUAUGAAUAUAACCCCUGCCCUCGCCUCCAGCAGCACAGCAAAAAGGCCAAUACUAUUUA (.(((.((((.(((((((.((....-.....((.....)).........)).))))))).))))..((......)).....))))........... ( -27.17) >consensus _AGCCGGAGGAGAGGGCGUGGCAAAAAACGAAUUAAAUAUGUAUACAAUACCCGCCCUCGCCUCCAGCAGCACAGCAAAAAGGCCAAUACUAUUUA ..((((((((.(((((((.((.............................))))))))).))))).((......)).....)))............ (-28.15 = -28.63 + 0.48)


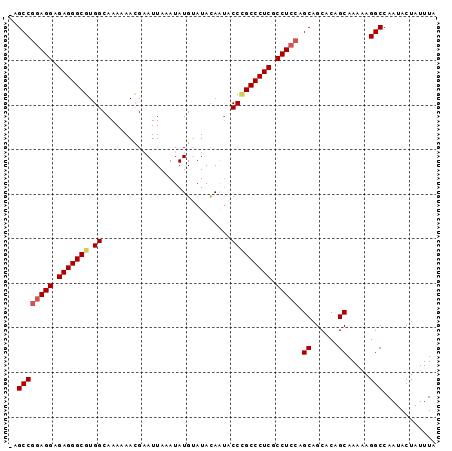
| Location | 17,161,435 – 17,161,530 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 93.18 |
| Mean single sequence MFE | -37.49 |
| Consensus MFE | -34.71 |
| Energy contribution | -35.03 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.11 |
| Mean z-score | -4.57 |
| Structure conservation index | 0.93 |
| SVM decision value | 4.82 |
| SVM RNA-class probability | 0.999953 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17161435 95 - 27905053 UAAAUAGUAUUGGCCUUUUUGCUGUGCUGCUGGAGGCGAGGGCGGGUAUUGUAUACAUAUUUAAUCCGUUUUUUGCCACGCCCUCUCCUCCGGCU- .....(((((.(((......))))))))((((((((.(((((((((((.........................)))).))))))).)))))))).- ( -40.21) >DroSec_CAF1 70647 95 - 1 UAAAUAGUAUUGGCCUUUUUGCUGUGCUGCUGGAGGCGAGGGCGGGUAUUGUAUACAUAUUUAAUUCGUUUUUUGCCACGCCCUCUCCUCCGGCC- .....(((((.(((......))))))))((((((((.(((((((((((.........................)))).))))))).)))))))).- ( -39.81) >DroSim_CAF1 61952 95 - 1 UAAAUAGUAUUGGCCUUUUUGCUGUGCUGCUGGAGGCGAGGGCGGGUAUUGUAUACAUAUUUAAUUCGUUUUUUGCCACGCCCUCUCCUCCGGCC- .....(((((.(((......))))))))((((((((.(((((((((((.........................)))).))))))).)))))))).- ( -39.81) >DroEre_CAF1 61835 93 - 1 UAAAUAGUAUUGGCCUUUUUGCUGUGCUGCUGGAGGCGAGGGC-GGGACUGUAUUCAUAUUUAAUUCAUU-UUUGCCACGCCCUCUCCUUCGGCU- .....(((((.(((......))))))))((((((((.((((((-(((.(.....................-...))).))))))).)))))))).- ( -34.36) >DroYak_CAF1 89402 95 - 1 UAAAUAGUAUUGGCCUUUUUGCUGUGCUGCUGGAGGCGAGGGCAGGGGUUAUAUUCAUAUUUAAUUCAUU-UUUGCCACGCCCUCUCCUCUGGCUC .....(((((.(((......))))))))((..((((.((((((...(((.....................-...)))..)))))).))))..)).. ( -33.26) >consensus UAAAUAGUAUUGGCCUUUUUGCUGUGCUGCUGGAGGCGAGGGCGGGUAUUGUAUACAUAUUUAAUUCGUUUUUUGCCACGCCCUCUCCUCCGGCU_ .....(((((.(((......))))))))((((((((.(((((((((((.........................)))).))))))).)))))))).. (-34.71 = -35.03 + 0.32)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:23:05 2006