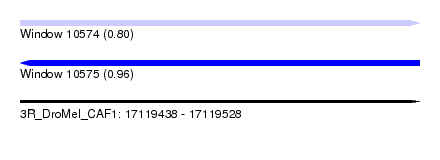

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 17,119,438 – 17,119,528 |
| Length | 90 |
| Max. P | 0.958966 |

| Location | 17,119,438 – 17,119,528 |
|---|---|
| Length | 90 |
| Sequences | 3 |
| Columns | 90 |
| Reading direction | forward |
| Mean pairwise identity | 82.16 |
| Mean single sequence MFE | -24.83 |
| Consensus MFE | -17.56 |
| Energy contribution | -18.22 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.800301 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
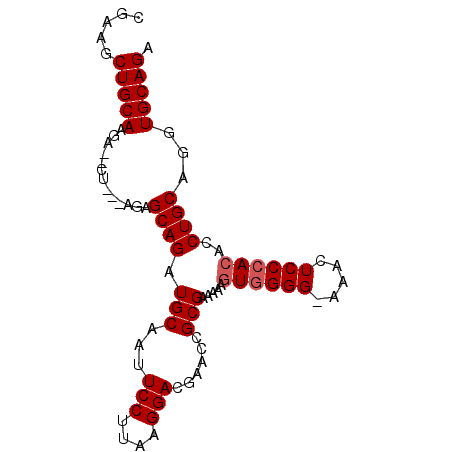
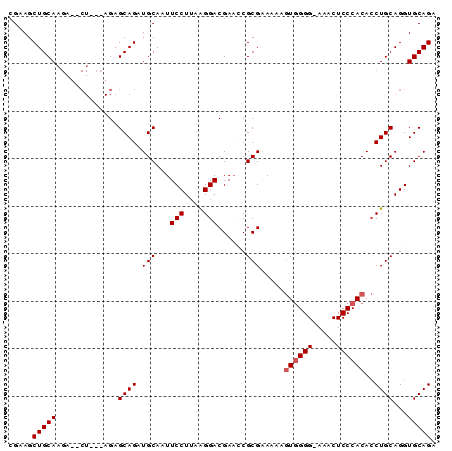
>3R_DroMel_CAF1 17119438 90 + 27905053 CGAAGCUGCAAGAUACUACCAGAGCAGAUGCAAUUCCUAAGGGACGAUGCGCGAAAAAAUGGGGGAAACUCCCACAACUGCAGGUGCAGA .....(((((.....(.....).((((..(((..(((....)))...))).........(((((....).))))...))))...))))). ( -24.80) >DroEre_CAF1 31856 89 + 1 GGAAGCUGCAAGAAGCUGAAAGAGCAGAUGCAAUUCCUUAAGGACGAAUCGCGAAAAAGUGGGG-AAACUCCCACACCUGCAGGUGCAGA ...((((......))))......(((..((((..(((....)))..............((((((-....))))))...))))..)))... ( -25.30) >DroYak_CAF1 44268 78 + 1 CGAAGCUGCAA----------GAGCAGAUGCAAUUCCUUAAGGACGAACCGCGGAAAAGUAGGG-A-ACUCCCACACCUGCAGGUGCAGA .....(((((.----------..((((.(((..((((....((.....))..))))..)))(((-.-...)))....))))...))))). ( -24.40) >consensus CGAAGCUGCAAGA__CU___AGAGCAGAUGCAAUUCCUUAAGGACGAACCGCGAAAAAGUGGGG_AAACUCCCACACCUGCAGGUGCAGA .....(((((.............((((.(((...(((....)))......))).....((((((.....))))))..))))...))))). (-17.56 = -18.22 + 0.67)

| Location | 17,119,438 – 17,119,528 |
|---|---|
| Length | 90 |
| Sequences | 3 |
| Columns | 90 |
| Reading direction | reverse |
| Mean pairwise identity | 82.16 |
| Mean single sequence MFE | -26.13 |
| Consensus MFE | -22.14 |
| Energy contribution | -22.59 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.50 |
| SVM RNA-class probability | 0.958966 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
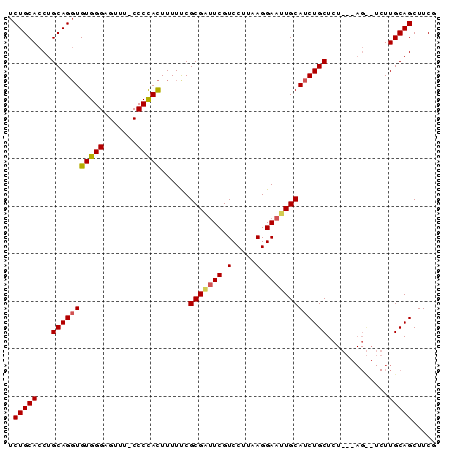
>3R_DroMel_CAF1 17119438 90 - 27905053 UCUGCACCUGCAGUUGUGGGAGUUUCCCCCAUUUUUUCGCGCAUCGUCCCUUAGGAAUUGCAUCUGCUCUGGUAGUAUCUUGCAGCUUCG .(((((((.((((..(((((.......)))))........(((...(((....)))..)))..))))...)))........))))..... ( -23.00) >DroEre_CAF1 31856 89 - 1 UCUGCACCUGCAGGUGUGGGAGUUU-CCCCACUUUUUCGCGAUUCGUCCUUAAGGAAUUGCAUCUGCUCUUUCAGCUUCUUGCAGCUUCC ...((....(((((((((((.....-.)))))......(((((((.(.....).))))))))))))).......((.....)).)).... ( -27.50) >DroYak_CAF1 44268 78 - 1 UCUGCACCUGCAGGUGUGGGAGU-U-CCCUACUUUUCCGCGGUUCGUCCUUAAGGAAUUGCAUCUGCUC----------UUGCAGCUUCG .(((((...(((((((((((...-.-.)))))......(((((((.(.....).)))))))))))))..----------.)))))..... ( -27.90) >consensus UCUGCACCUGCAGGUGUGGGAGUUU_CCCCACUUUUUCGCGAUUCGUCCUUAAGGAAUUGCAUCUGCUCU___AG__UCUUGCAGCUUCG .(((((...(((((((((((.......)))))......(((((((.(.....).))))))))))))).............)))))..... (-22.14 = -22.59 + 0.45)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:22:52 2006