
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 17,115,476 – 17,115,606 |
| Length | 130 |
| Max. P | 0.946998 |

| Location | 17,115,476 – 17,115,584 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 89.10 |
| Mean single sequence MFE | -26.80 |
| Consensus MFE | -22.56 |
| Energy contribution | -23.57 |
| Covariance contribution | 1.01 |
| Combinations/Pair | 1.08 |
| Mean z-score | -3.03 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.37 |
| SVM RNA-class probability | 0.946998 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17115476 108 - 27905053 GUACGCUGGCAAAAUAAGAGUUGUCAACAUACUCGUAAAACAAUAUGUUCGAUUAAUAGGAAAUAGUUUAACAAAUUGUGUUACUACAUUUCUUUGCCACAAGUAUAU ((((..(((((((....((((((....)).))))((((.(((((.((((.(((((........))))).)))).))))).))))........)))))))...)))).. ( -27.40) >DroSec_CAF1 27769 105 - 1 GUACACUGGCAAA---AGAGUUGUCAACAUGCUCGUAAAACAACAUGUUGGAUUAUUAGGAAAUAAUUUAACAAAUUGUGUUGCUACCUUGUCUUGCCACAAGUAUAU ((((..((((((.---.((((((....)).))))((((.((((..((((((((((((....))))))))))))..)))).)))).........))))))...)))).. ( -28.90) >DroSim_CAF1 26751 105 - 1 GUACACUGGCAAA---AGAGUUGUCAACAUGCUCGUAAAACAACAUGUUGGAUUAUUAGGAAAUAGUUUAACAAAUGGUGUUGAUACAUUCCCUUGACACAAGUAUAU ((((..((.(((.---.(((((((((((((....((......)).((((((((((((....))))))))))))....))))))))).))))..))).))...)))).. ( -24.10) >consensus GUACACUGGCAAA___AGAGUUGUCAACAUGCUCGUAAAACAACAUGUUGGAUUAUUAGGAAAUAGUUUAACAAAUUGUGUUGCUACAUUCCCUUGCCACAAGUAUAU ((((..((((((.....((((((....)).))))((((.((((..((((((((((((....))))))))))))..)))).)))).........))))))...)))).. (-22.56 = -23.57 + 1.01)



| Location | 17,115,511 – 17,115,606 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 82.89 |
| Mean single sequence MFE | -17.72 |
| Consensus MFE | -11.96 |
| Energy contribution | -13.02 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.606756 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17115511 95 + 27905053 --UGUUAAACUAUUUCCUAUUAAUCGAACAUAUUGUUUUACGAGUAUGUUGACAACUCUUAUUUUGCCAGCGUACUCGCUGUAUAGCAACAAGAAAA --(((((.((...............((((.....))))..(((((((((((.(((........))).)))))))))))..)).)))))......... ( -20.90) >DroSec_CAF1 27804 92 + 1 --UGUUAAAUUAUUUCCUAAUAAUCCAACAUGUUGUUUUACGAGCAUGUUGACAACUCU---UUUGCCAGUGUACUCGCUGUAUAGCAACAAGAAAA --.(((..((((((....)))))).(((((((((........)))))))))..)))(((---((((((((((....)))))....)))).))))... ( -20.40) >DroSim_CAF1 26786 92 + 1 --UGUUAAACUAUUUCCUAAUAAUCCAACAUGUUGUUUUACGAGCAUGUUGACAACUCU---UUUGCCAGUGUACUCGCUGUAUAGCAACAAGAAAA --.......................(((((((((........))))))))).....(((---((((((((((....)))))....)))).))))... ( -19.80) >DroEre_CAF1 27938 81 + 1 UUUAUUCAACU-----CAAAUAAUUCAACAUUUUGUUUUGCG---CUAUUGAAA----U---UUUGCCAGUGUAC-CGCUGUAUAGCAACAAGAAAA ...........-----..............((((((..((((---(........----.---...))(((((...-)))))....)))))))))... ( -9.80) >consensus __UGUUAAACUAUUUCCUAAUAAUCCAACAUGUUGUUUUACGAGCAUGUUGACAACUCU___UUUGCCAGUGUACUCGCUGUAUAGCAACAAGAAAA .........................(((((((((........)))))))))............(((((((((....)))))....))))........ (-11.96 = -13.02 + 1.06)



| Location | 17,115,511 – 17,115,606 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 82.89 |
| Mean single sequence MFE | -19.78 |
| Consensus MFE | -14.81 |
| Energy contribution | -16.87 |
| Covariance contribution | 2.06 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.777359 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 17115511 95 - 27905053 UUUUCUUGUUGCUAUACAGCGAGUACGCUGGCAAAAUAAGAGUUGUCAACAUACUCGUAAAACAAUAUGUUCGAUUAAUAGGAAAUAGUUUAACA-- .((((((((((.......(((((((.(.((((((........)))))).).)))))))..(((.....)))....))))))))))..........-- ( -21.30) >DroSec_CAF1 27804 92 - 1 UUUUCUUGUUGCUAUACAGCGAGUACACUGGCAAA---AGAGUUGUCAACAUGCUCGUAAAACAACAUGUUGGAUUAUUAGGAAAUAAUUUAACA-- ......(((((.......(((((((...((((((.---....))))))...)))))))....)))))((((((((((((....))))))))))))-- ( -24.10) >DroSim_CAF1 26786 92 - 1 UUUUCUUGUUGCUAUACAGCGAGUACACUGGCAAA---AGAGUUGUCAACAUGCUCGUAAAACAACAUGUUGGAUUAUUAGGAAAUAGUUUAACA-- ......(((((.......(((((((...((((((.---....))))))...)))))))....)))))((((((((((((....))))))))))))-- ( -24.10) >DroEre_CAF1 27938 81 - 1 UUUUCUUGUUGCUAUACAGCG-GUACACUGGCAAA---A----UUUCAAUAG---CGCAAAACAAAAUGUUGAAUUAUUUG-----AGUUGAAUAAA ........(((((((((....-)))...)))))).---.----.(((((((.---............)))))))((((((.-----....)))))). ( -9.62) >consensus UUUUCUUGUUGCUAUACAGCGAGUACACUGGCAAA___AGAGUUGUCAACAUGCUCGUAAAACAACAUGUUGGAUUAUUAGGAAAUAGUUUAACA__ ......(((((.......(((((((...((((((........))))))...)))))))....)))))((((((((((((....)))))))))))).. (-14.81 = -16.87 + 2.06)


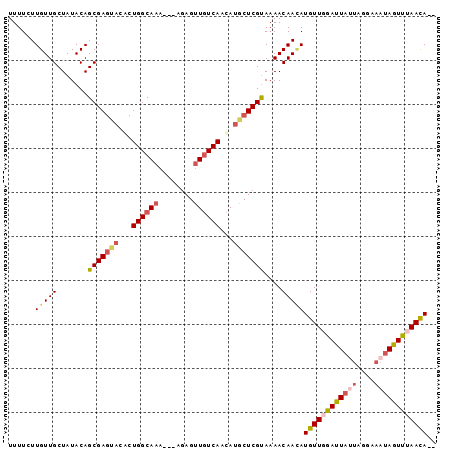
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:22:49 2006