
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 16,821,565 – 16,821,707 |
| Length | 142 |
| Max. P | 0.863023 |

| Location | 16,821,565 – 16,821,667 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 70.90 |
| Mean single sequence MFE | -22.80 |
| Consensus MFE | -5.58 |
| Energy contribution | -8.66 |
| Covariance contribution | 3.08 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.24 |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.863023 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16821565 102 - 27905053 GACAAUUAGAAAUAAGUUUAUGCUGUAUA-GCUACUGUACAAUUUUAGUUGCUAAACAGCUUAUAUCUUUGAGAGUAUAA-ACUUGUUCCAUAGAUAACCA----------------AAA ........(.((((((((((((((((.((-((.((((........)))).)))).))..((((......)))))))))))-))))))).)...........----------------... ( -25.20) >DroSec_CAF1 5485 102 - 1 GAUAAUUAGAAAUAAGUUGAAGCUGUGUA-GCUACUGUGCAAUUUUAGUUGCUAAACAGCUAAAAUCUUUGAGAGUAUAA-ACUUGUUCCAGAGAUGGCCA----------------AAC .....(((....))).(((.((((((.((-((.((((........)))).)))).))))))...(((((((.(((((...-...))))))))))))...))----------------).. ( -25.80) >DroSim_CAF1 3352 102 - 1 GAUAAUUAGAAAUAAGUUGAAGCUGUAUA-GCUACUGUGCAAUUUUAGUUGCUAAACAGCUUAUAUCUUUGAGAGUAUAA-ACUUGUUCCAGAGAUGGCCA----------------AAA .....(((....))).((((((((((.((-((.((((........)))).)))).))))))).((((((((.(((((...-...)))))))))))))..))----------------).. ( -27.90) >DroEre_CAF1 8696 106 - 1 AAUAAUUAGAAAUUACUUU-----GUAUA-GCUACUGUAAAAUAUGAGUUGCUAAACAGCUUACGCUUUG-AGCGUAUAAAACUUGUUUCAGAGACAACCAAUACGU-------UAGCAU ...................-----.....-((((.((((.....(((((((.....))))))).((((((-((((.........)).))))))).)......)))).-------)))).. ( -18.80) >DroYak_CAF1 8786 115 - 1 AAUAAUUAUAAAUUAGUUU-----UUGUUGGAAAGUGGAAAGUGUGAGUAGCUAAACAGCCGACGUCUUGCAGAGUAUAAAACUUGUUUCAGAAGCCACAAUAACAUACAAGUUUCAAAA ...............(((.-----((((.((....(((((..((..((..(((....)))......))..))((((.....)))).)))))....)))))).)))............... ( -16.30) >consensus GAUAAUUAGAAAUAAGUUUA_GCUGUAUA_GCUACUGUACAAUUUUAGUUGCUAAACAGCUUACAUCUUUGAGAGUAUAA_ACUUGUUCCAGAGAUAACCA________________AAA .....................(((((....((.((((........)))).))...)))))....(((((((.((((.....))))....)))))))........................ ( -5.58 = -8.66 + 3.08)



| Location | 16,821,588 – 16,821,707 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 89.20 |
| Mean single sequence MFE | -24.90 |
| Consensus MFE | -15.32 |
| Energy contribution | -14.88 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.11 |
| Mean z-score | -3.46 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.584010 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16821588 119 + 27905053 UUAUACUCUCAAAGAUAUAAGCUGUUUAGCAACUAAAAUUGUACAGUAGCUAUACAGCAUAAACUUAUUUCUAAUUGUCUGCCAAUUGUUUGCUAUAGAAUAGUAUCUAUGUAGAUAUG .((((.(((((.((((((..(((((.((((.(((..........))).)))).)))))..........(((((...((..((.....))..))..)))))..)))))).)).))))))) ( -25.40) >DroSec_CAF1 5508 114 + 1 UUAUACUCUCAAAGAUUUUAGCUGUUUAGCAACUAAAAUUGCACAGUAGCUACACAGCUUCAACUUAUUUCUAAUUAUCUACCAAUUGUUUG-UAUACUCUUG----UAUAUAGAUAUA ...........(((.((..((((((.((((.(((..........))).)))).))))))..)))))....................((((((-(((((....)----)))))))))).. ( -24.20) >DroSim_CAF1 3375 114 + 1 UUAUACUCUCAAAGAUAUAAGCUGUUUAGCAACUAAAAUUGCACAGUAGCUAUACAGCUUCAACUUAUUUCUAAUUAUCUACCAAUUGUUUG-UAUACUCUUG----UAUAUAGAUAUA ............(((((.(((((((.((((.(((..........))).)))).)))))))...............)))))......((((((-(((((....)----)))))))))).. ( -25.09) >consensus UUAUACUCUCAAAGAUAUAAGCUGUUUAGCAACUAAAAUUGCACAGUAGCUAUACAGCUUCAACUUAUUUCUAAUUAUCUACCAAUUGUUUG_UAUACUCUUG____UAUAUAGAUAUA ....................(((((.((((.(((..........))).)))).))))).................((((((..............................)))))).. (-15.32 = -14.88 + -0.44)

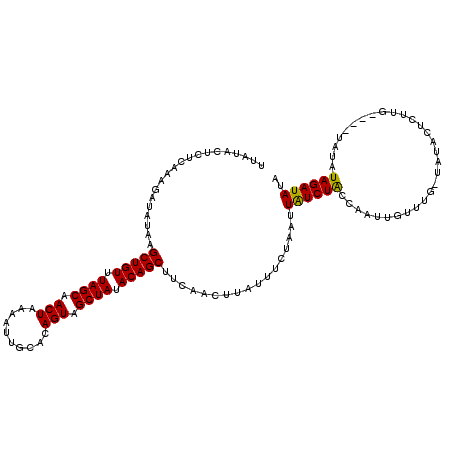

| Location | 16,821,588 – 16,821,707 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 89.20 |
| Mean single sequence MFE | -25.00 |
| Consensus MFE | -19.17 |
| Energy contribution | -19.83 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.717797 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16821588 119 - 27905053 CAUAUCUACAUAGAUACUAUUCUAUAGCAAACAAUUGGCAGACAAUUAGAAAUAAGUUUAUGCUGUAUAGCUACUGUACAAUUUUAGUUGCUAAACAGCUUAUAUCUUUGAGAGUAUAA ....(((.((.(((((...(((((.........((((.....)))))))))..........(((((.((((.((((........)))).)))).)))))...))))).)))))...... ( -25.10) >DroSec_CAF1 5508 114 - 1 UAUAUCUAUAUA----CAAGAGUAUA-CAAACAAUUGGUAGAUAAUUAGAAAUAAGUUGAAGCUGUGUAGCUACUGUGCAAUUUUAGUUGCUAAACAGCUAAAAUCUUUGAGAGUAUAA ..((((((((((----(....)))).-(((....)))))))))).(((....))).....((((((.((((.((((........)))).)))).))))))................... ( -24.10) >DroSim_CAF1 3375 114 - 1 UAUAUCUAUAUA----CAAGAGUAUA-CAAACAAUUGGUAGAUAAUUAGAAAUAAGUUGAAGCUGUAUAGCUACUGUGCAAUUUUAGUUGCUAAACAGCUUAUAUCUUUGAGAGUAUAA ........((((----(....)))))-...((..(..(.(((((.(((....)))....(((((((.((((.((((........)))).)))).))))))).))))))..)..)).... ( -25.80) >consensus UAUAUCUAUAUA____CAAGAGUAUA_CAAACAAUUGGUAGAUAAUUAGAAAUAAGUUGAAGCUGUAUAGCUACUGUGCAAUUUUAGUUGCUAAACAGCUUAUAUCUUUGAGAGUAUAA ..............................((..(..(.(((((.(((....)))......(((((.((((.((((........)))).)))).)))))...))))))..)..)).... (-19.17 = -19.83 + 0.67)


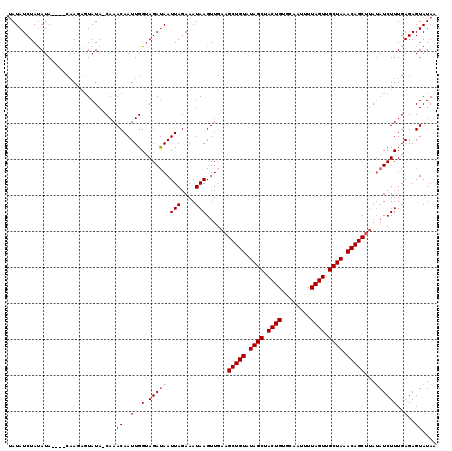
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:21:11 2006