
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 16,767,439 – 16,767,531 |
| Length | 92 |
| Max. P | 0.989347 |

| Location | 16,767,439 – 16,767,531 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 86.04 |
| Mean single sequence MFE | -29.14 |
| Consensus MFE | -22.00 |
| Energy contribution | -23.20 |
| Covariance contribution | 1.20 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.771727 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16767439 92 + 27905053 AGAGGAGGGUGUGUGGCAGGC--GCUUGCCACAAGGGCGUGGCAUUUGCCUGUUGAAAUAUUGCAACCGCUGCCAAUAAGCAAAGAAAGGAUAA ...(((((((.((((((((((--(..((((((......))))))..))))))))........)))))).)).)).................... ( -30.00) >DroSec_CAF1 70207 89 + 1 ---GGGGGAUGUGUGGCAGGC--GCUUGCCACAAGGGCGUGGCAUUUGCCUGUUGAAAUAUUGCAACCGCUGCCAAUAAGCGAAGAAAGGAUAA ---..(.(((((..(((((((--(..((((((......))))))..))))))))...))))).)...((((.......))))............ ( -32.80) >DroSim_CAF1 64882 89 + 1 ---GGGGGAUGUGUGGCAGGC--GCUUGCCACAAGGGCGUGGCAUUUGCCUGUUGAAAUAUUGCAACCGCUGCCAAUAAGCGAAGAAAGGAUAA ---..(.(((((..(((((((--(..((((((......))))))..))))))))...))))).)...((((.......))))............ ( -32.80) >DroEre_CAF1 61942 88 + 1 ---GGGAGAGGGGCGGCAGGC--GCUUGCCACCAGGGCGUGGCAUUUGCCUGUUGAAAUAUUGCAACCGCUGCCAAUAAGCGAGG-AAGGAUAA ---........((((((((((--(..((((((......))))))..)))))((((........)))).))))))...........-........ ( -32.30) >DroAna_CAF1 41415 83 + 1 -----------UGUGGGAGGCUGACUGGGUUAUGGGGCGUGGCAUUUGCCUGUCGAAAUAUUGCAACCGCUGCCAAUAAGCGAAGAAAGGAUAA -----------........(((.....(((..(((((((.(((....)))))))(........)..)))..)))....)))............. ( -17.80) >consensus ___GGGGGAUGUGUGGCAGGC__GCUUGCCACAAGGGCGUGGCAUUUGCCUGUUGAAAUAUUGCAACCGCUGCCAAUAAGCGAAGAAAGGAUAA ...........(((((((((....)))))))))...((.(((((...((..((((........)))).)))))))....))............. (-22.00 = -23.20 + 1.20)

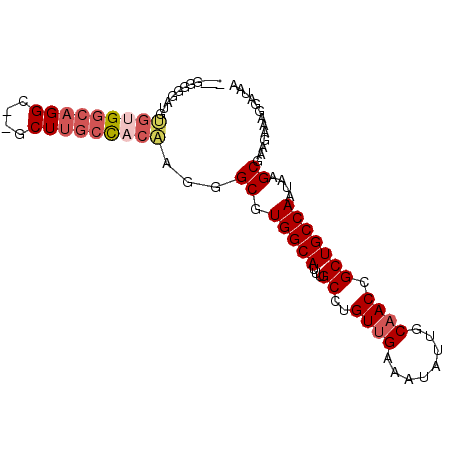

| Location | 16,767,439 – 16,767,531 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 86.04 |
| Mean single sequence MFE | -23.84 |
| Consensus MFE | -20.46 |
| Energy contribution | -21.34 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.86 |
| SVM decision value | 2.16 |
| SVM RNA-class probability | 0.989347 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16767439 92 - 27905053 UUAUCCUUUCUUUGCUUAUUGGCAGCGGUUGCAAUAUUUCAACAGGCAAAUGCCACGCCCUUGUGGCAAGC--GCCUGCCACACACCCUCCUCU ...................((((((((.((((............(((....))).(((....))))))).)--).))))))............. ( -25.40) >DroSec_CAF1 70207 89 - 1 UUAUCCUUUCUUCGCUUAUUGGCAGCGGUUGCAAUAUUUCAACAGGCAAAUGCCACGCCCUUGUGGCAAGC--GCCUGCCACACAUCCCCC--- ...................((((((((.((((............(((....))).(((....))))))).)--).))))))..........--- ( -25.40) >DroSim_CAF1 64882 89 - 1 UUAUCCUUUCUUCGCUUAUUGGCAGCGGUUGCAAUAUUUCAACAGGCAAAUGCCACGCCCUUGUGGCAAGC--GCCUGCCACACAUCCCCC--- ...................((((((((.((((............(((....))).(((....))))))).)--).))))))..........--- ( -25.40) >DroEre_CAF1 61942 88 - 1 UUAUCCUU-CCUCGCUUAUUGGCAGCGGUUGCAAUAUUUCAACAGGCAAAUGCCACGCCCUGGUGGCAAGC--GCCUGCCGCCCCUCUCCC--- ........-...........(((((((.((((............(((....))).(((....))))))).)--).)))))...........--- ( -26.90) >DroAna_CAF1 41415 83 - 1 UUAUCCUUUCUUCGCUUAUUGGCAGCGGUUGCAAUAUUUCGACAGGCAAAUGCCACGCCCCAUAACCCAGUCAGCCUCCCACA----------- .............(((.(((((..(((((((........)))).(((....))).)))........))))).)))........----------- ( -16.10) >consensus UUAUCCUUUCUUCGCUUAUUGGCAGCGGUUGCAAUAUUUCAACAGGCAAAUGCCACGCCCUUGUGGCAAGC__GCCUGCCACACAUCCCCC___ .............(((....))).(((((((........)))).(((....))).)))...(((((((.(....).)))))))........... (-20.46 = -21.34 + 0.88)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:20:13 2006