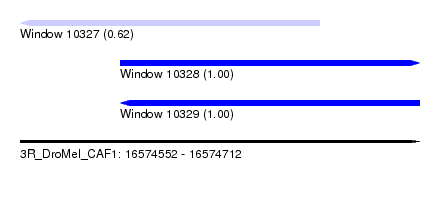
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 16,574,552 – 16,574,712 |
| Length | 160 |
| Max. P | 0.999891 |

| Location | 16,574,552 – 16,574,672 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.75 |
| Mean single sequence MFE | -48.90 |
| Consensus MFE | -41.32 |
| Energy contribution | -42.24 |
| Covariance contribution | 0.92 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.31 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.622266 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16574552 120 - 27905053 CGCCCAGGAAGCUGUUCGAAAGGGCUCCACAGCGGUCGGCGUUCGUGGAGCCAAUUGUGUGGUGCUGGGUGUGGAGAAGAGCUCGGUGUCCGAGAUGCAGGAGGAUCGGACGGUGCGCAA (((((((...((..(.(((...((((((((((((.....)))).))))))))..))).)..)).))))))).........((.((.(((((((..(......)..))))))).)).)).. ( -53.00) >DroSec_CAF1 130767 120 - 1 CGCCCAGGAAGCUGUUCGAAAGGGUUCCACAGCGGUUGGCGUUCGUGGAGCCAAUUGCGUGGUGCUGGGCGUGGAGAAGAGCUCGGUGUCCCCGAUGCAGGAGGAUCGAACGGUGCAAAA (((((((...((..(.(((...((((((((((((.....)))).))))))))..))).)..)).))))))).........((.(.((((((((......)).))))...)).).)).... ( -47.90) >DroSim_CAF1 135117 120 - 1 CGCCCAGGAAGCUGUUCGAAAGGGUUCCACAGCGGUUGGCGUUCGUGGAGCCAAUUGCGUGGUGCUGGGCGUGGAGAAGAGCUCGGUGUCCCCGAUGCAGGAGGAUCGAACGGUGCAAAA (((((((...((..(.(((...((((((((((((.....)))).))))))))..))).)..)).))))))).........((.(.((((((((......)).))))...)).).)).... ( -47.90) >DroEre_CAF1 134881 120 - 1 CGCCCAGGAAGCUGUUCGCAAGGGCUCCACAGCGGUUGGGGUUCGUGGAGCCAAUUGCGUGGUGCUAGGCGUGGAGAAGAGCUCGUUGUCCAAGAUGCAGGAGGACCGCACGGUGCGCAA (((((............((((.(((((((((((.......))).))))))))..))))(((((.((..(((((((.(((....).)).))))...)))...)).)))))..)).)))... ( -46.40) >DroYak_CAF1 130884 120 - 1 CGCCCAGGAAGCUGUUCGAAAGGGCUCCACGGCGGUUGGGGUGCGUGGAGCCAAUUGCGUGGUGCUGGGCGUGGAGAAGAGCUCGGUGUCCAAGAUGCAGGAGUACCGAACGGUGCGCAA .((((((...((..(.(((...(((((((((((.......)).)))))))))..))).)..)).))))))(((.(.....(.(((((((((........))).)))))).)..).))).. ( -49.30) >consensus CGCCCAGGAAGCUGUUCGAAAGGGCUCCACAGCGGUUGGCGUUCGUGGAGCCAAUUGCGUGGUGCUGGGCGUGGAGAAGAGCUCGGUGUCCAAGAUGCAGGAGGAUCGAACGGUGCGCAA .((((((....))((((((...((((((((((((.....)))).))))))))..(((((((..(((((((..........)))))))..)....)))))).....)))))))).)).... (-41.32 = -42.24 + 0.92)

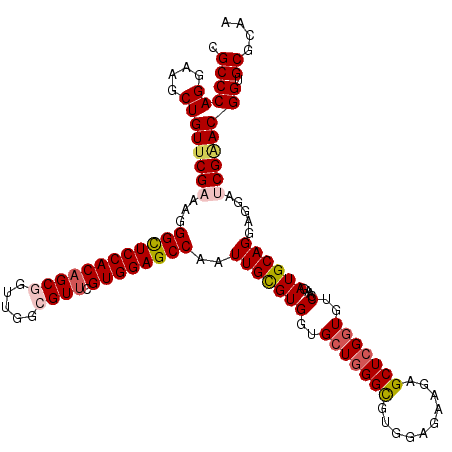
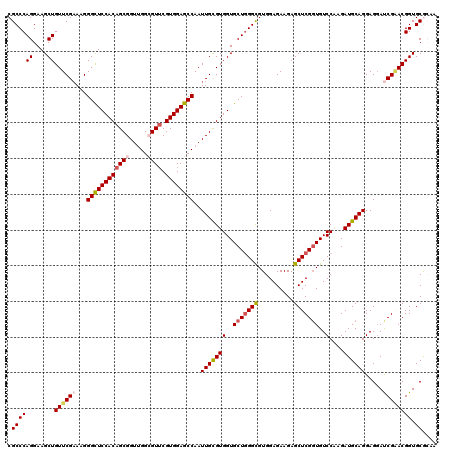
| Location | 16,574,592 – 16,574,712 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.33 |
| Mean single sequence MFE | -43.79 |
| Consensus MFE | -40.16 |
| Energy contribution | -40.64 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.78 |
| Structure conservation index | 0.92 |
| SVM decision value | 4.15 |
| SVM RNA-class probability | 0.999816 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16574592 120 + 27905053 UCUUCUCCACACCCAGCACCACACAAUUGGCUCCACGAACGCCGACCGCUGUGGAGCCCUUUCGAACAGCUUCCUGGGCGUACUCCACCUGCAACAGGUGGCCAUCCGGCGAGAAUAUGG ..(((((.((.(((((............(((((((((...((.....))))))))))).....(((....)))))))).))....((((((...))))))(((....))))))))..... ( -41.10) >DroSec_CAF1 130807 120 + 1 UCUUCUCCACGCCCAGCACCACGCAAUUGGCUCCACGAACGCCAACCGCUGUGGAACCCUUUCGAACAGCUUCCUGGGCGUACUCCACCUGCAACAGGUGGCCAUCCGGCGAGAAUAUGG ..(((((.((((((((..((((((..(((((.........)))))..)).)))).........(((....)))))))))))....((((((...))))))(((....))))))))..... ( -44.30) >DroSim_CAF1 135157 120 + 1 UCUUCUCCACGCCCAGCACCACGCAAUUGGCUCCACGAACGCCAACCGCUGUGGAACCCUUUCGAACAGCUUCCUGGGCGUACUCCACCUGCAACAGGUGGCCAUCCGGCGAGAAUAUGG ..(((((.((((((((..((((((..(((((.........)))))..)).)))).........(((....)))))))))))....((((((...))))))(((....))))))))..... ( -44.30) >DroEre_CAF1 134921 120 + 1 UCUUCUCCACGCCUAGCACCACGCAAUUGGCUCCACGAACCCCAACCGCUGUGGAGCCCUUGCGAACAGCUUCCUGGGCGUACUCCACCUGCAACAAGUGGCCAUCCGGCGAGAAUAUGG ..(((((.(((((((((....(((((..(((((((((............))))))))).)))))....)))....))))))....(((.((...)).)))(((....))))))))..... ( -41.90) >DroYak_CAF1 130924 120 + 1 UCUUCUCCACGCCCAGCACCACGCAAUUGGCUCCACGCACCCCAACCGCCGUGGAGCCCUUUCGAACAGCUUCCUGGGCGUACUCCACCUGCAACAGGUGGCCAUCCGGCGAGAAUAUGG ..(((((.((((((((.....((.((..((((((((((.........).)))))))))..)))).........))))))))....((((((...))))))(((....))))))))..... ( -47.34) >consensus UCUUCUCCACGCCCAGCACCACGCAAUUGGCUCCACGAACGCCAACCGCUGUGGAGCCCUUUCGAACAGCUUCCUGGGCGUACUCCACCUGCAACAGGUGGCCAUCCGGCGAGAAUAUGG ..(((((.((((((((............(((((((((............))))))))).....(((....)))))))))))....((((((...))))))(((....))))))))..... (-40.16 = -40.64 + 0.48)


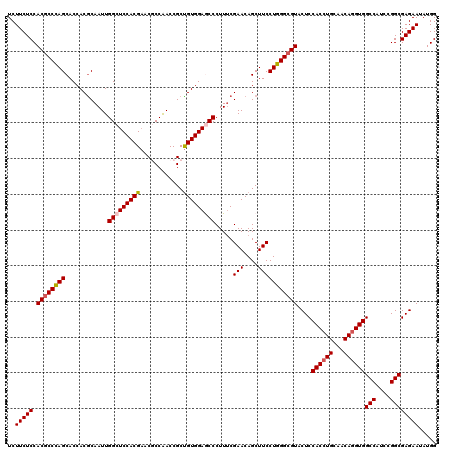
| Location | 16,574,592 – 16,574,712 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.33 |
| Mean single sequence MFE | -54.54 |
| Consensus MFE | -52.14 |
| Energy contribution | -52.58 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.55 |
| Structure conservation index | 0.96 |
| SVM decision value | 4.41 |
| SVM RNA-class probability | 0.999891 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
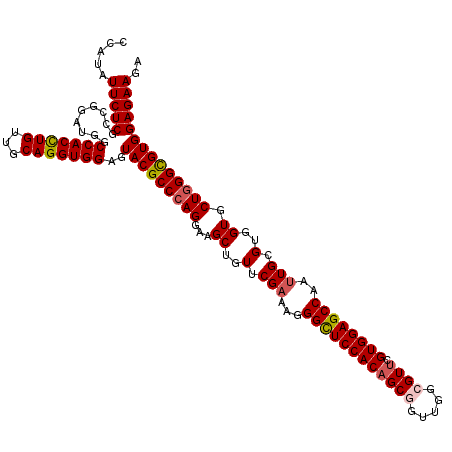
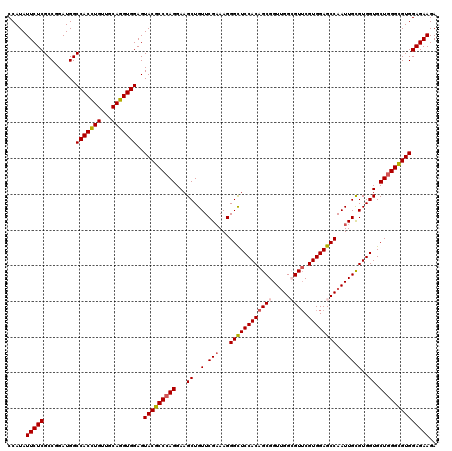
>3R_DroMel_CAF1 16574592 120 - 27905053 CCAUAUUCUCGCCGGAUGGCCACCUGUUGCAGGUGGAGUACGCCCAGGAAGCUGUUCGAAAGGGCUCCACAGCGGUCGGCGUUCGUGGAGCCAAUUGUGUGGUGCUGGGUGUGGAGAAGA .....(((((.........(((((((...)))))))..(((((((((...((..(.(((...((((((((((((.....)))).))))))))..))).)..)).)))))))))))))).. ( -56.00) >DroSec_CAF1 130807 120 - 1 CCAUAUUCUCGCCGGAUGGCCACCUGUUGCAGGUGGAGUACGCCCAGGAAGCUGUUCGAAAGGGUUCCACAGCGGUUGGCGUUCGUGGAGCCAAUUGCGUGGUGCUGGGCGUGGAGAAGA .....(((((.........(((((((...)))))))..(((((((((...((..(.(((...((((((((((((.....)))).))))))))..))).)..)).)))))))))))))).. ( -55.50) >DroSim_CAF1 135157 120 - 1 CCAUAUUCUCGCCGGAUGGCCACCUGUUGCAGGUGGAGUACGCCCAGGAAGCUGUUCGAAAGGGUUCCACAGCGGUUGGCGUUCGUGGAGCCAAUUGCGUGGUGCUGGGCGUGGAGAAGA .....(((((.........(((((((...)))))))..(((((((((...((..(.(((...((((((((((((.....)))).))))))))..))).)..)).)))))))))))))).. ( -55.50) >DroEre_CAF1 134921 120 - 1 CCAUAUUCUCGCCGGAUGGCCACUUGUUGCAGGUGGAGUACGCCCAGGAAGCUGUUCGCAAGGGCUCCACAGCGGUUGGGGUUCGUGGAGCCAAUUGCGUGGUGCUAGGCGUGGAGAAGA .....(((((.........(((((((...)))))))..((((((.((...(((...(((((.(((((((((((.......))).))))))))..))))).))).)).))))))))))).. ( -49.10) >DroYak_CAF1 130924 120 - 1 CCAUAUUCUCGCCGGAUGGCCACCUGUUGCAGGUGGAGUACGCCCAGGAAGCUGUUCGAAAGGGCUCCACGGCGGUUGGGGUGCGUGGAGCCAAUUGCGUGGUGCUGGGCGUGGAGAAGA .....(((((.........(((((((...)))))))..(((((((((...((..(.(((...(((((((((((.......)).)))))))))..))).)..)).)))))))))))))).. ( -56.60) >consensus CCAUAUUCUCGCCGGAUGGCCACCUGUUGCAGGUGGAGUACGCCCAGGAAGCUGUUCGAAAGGGCUCCACAGCGGUUGGCGUUCGUGGAGCCAAUUGCGUGGUGCUGGGCGUGGAGAAGA .....(((((.........(((((((...)))))))..(((((((((...((..(.(((...((((((((((((.....)))).))))))))..))).)..)).)))))))))))))).. (-52.14 = -52.58 + 0.44)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:18:56 2006