
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 16,389,703 – 16,389,823 |
| Length | 120 |
| Max. P | 0.911180 |

| Location | 16,389,703 – 16,389,823 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.28 |
| Mean single sequence MFE | -41.11 |
| Consensus MFE | -39.95 |
| Energy contribution | -39.84 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.08 |
| SVM RNA-class probability | 0.911180 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
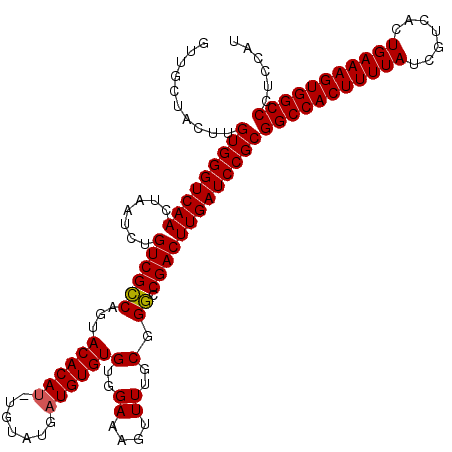
>3R_DroMel_CAF1 16389703 120 + 27905053 GUUGCUACUUGUGGGUCAACUAAUCUGUCGCCAGUACACAUGUGUAUGUUGUGUGUGGAAAGUUUUGCGGGCCGACUUGAUCCGCGGCCACUUUUAUCGUCACUGAAAGUGGCCCUCCAU ..........(((((((((.......((((((...(((((.........)))))(..((....))..).)).)))))))))))))(((((((((((.......)))))))))))...... ( -40.41) >DroSec_CAF1 10837 120 + 1 GUUGCUACUUGUGGGUCAACUAAUCUGUCGCCAGUACACAUAUGUAUGAUGUGUGUGGAAAGUUUUGCGGGCCGACUUGAUCCGCGGCCACUUUUAUCGUCACUGAAAGUGGCCCUCCAU ..........(((((((((.......((((((..(((((((((.....)))))))))....(.....).)).)))))))))))))(((((((((((.......)))))))))))...... ( -42.31) >DroEre_CAF1 11377 116 + 1 GUUGCUACUUGUGGGUCAACUAAUCUGUCGGCAGUACACAU----AUGAUGUGUGUGGAAAGUUUUGCGGUCCGACUUGAUCCGCGGCCACUUUUAUCGUCACUGAAAGUGGCCCUCCGU ..........(((((((((.......((((((...((((((----...))))))(..((....))..).).))))))))))))))(((((((((((.......)))))))))))...... ( -40.61) >consensus GUUGCUACUUGUGGGUCAACUAAUCUGUCGCCAGUACACAU_UGUAUGAUGUGUGUGGAAAGUUUUGCGGGCCGACUUGAUCCGCGGCCACUUUUAUCGUCACUGAAAGUGGCCCUCCAU ..........(((((((((.......((((((...((((((.......))))))(..((....))..).)).)))))))))))))(((((((((((.......)))))))))))...... (-39.95 = -39.84 + -0.11)


| Location | 16,389,703 – 16,389,823 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.28 |
| Mean single sequence MFE | -35.20 |
| Consensus MFE | -33.53 |
| Energy contribution | -33.87 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.779957 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16389703 120 - 27905053 AUGGAGGGCCACUUUCAGUGACGAUAAAAGUGGCCGCGGAUCAAGUCGGCCCGCAAAACUUUCCACACACAACAUACACAUGUGUACUGGCGACAGAUUAGUUGACCCACAAGUAGCAAC .((((((((((((((...........)))))))))((((..(.....)..))))......)))))....((((.((((....))))(((....)))....))))................ ( -36.70) >DroSec_CAF1 10837 120 - 1 AUGGAGGGCCACUUUCAGUGACGAUAAAAGUGGCCGCGGAUCAAGUCGGCCCGCAAAACUUUCCACACACAUCAUACAUAUGUGUACUGGCGACAGAUUAGUUGACCCACAAGUAGCAAC .((((((((((((((...........)))))))))((((..(.....)..))))......))))).((((((.......)))))).(((....))).(((.(((.....))).))).... ( -36.20) >DroEre_CAF1 11377 116 - 1 ACGGAGGGCCACUUUCAGUGACGAUAAAAGUGGCCGCGGAUCAAGUCGGACCGCAAAACUUUCCACACACAUCAU----AUGUGUACUGCCGACAGAUUAGUUGACCCACAAGUAGCAAC ..(((((((((((((...........)))))))))((((.((......))))))......))))..((((((...----))))))..(((((((......))))((......)).))).. ( -32.70) >consensus AUGGAGGGCCACUUUCAGUGACGAUAAAAGUGGCCGCGGAUCAAGUCGGCCCGCAAAACUUUCCACACACAUCAUACA_AUGUGUACUGGCGACAGAUUAGUUGACCCACAAGUAGCAAC ..(((((((((((((...........)))))))))((((..(.....)..))))......))))..((((((.......)))))).(((....))).(((.(((.....))).))).... (-33.53 = -33.87 + 0.33)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:17:29 2006