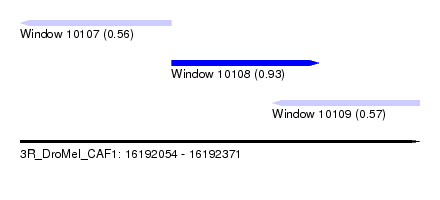
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 16,192,054 – 16,192,371 |
| Length | 317 |
| Max. P | 0.930056 |

| Location | 16,192,054 – 16,192,174 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.58 |
| Mean single sequence MFE | -31.76 |
| Consensus MFE | -28.68 |
| Energy contribution | -29.32 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.16 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.562943 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16192054 120 - 27905053 AAAAGGACAAAUAUUGCAUGCUGACUUCGGAAAAUGCCAAAGGUACUUGCUAUCAGAAUGUCAGCAGGGUCCCUUUAAACUGAAUCGAAUGUGCGAUUUAUACAGCCGCUUCACGGCCUU ....((((..........((((((((((((......))...((((.....)))).))).)))))))..))))........(((((((......)))))))....((((.....))))... ( -30.60) >DroSec_CAF1 21359 120 - 1 AAAAGGACAUAUAUUGCAUGCUGACUUCGGAAAAUGCCAAAGGUACUUGCUAUCAGAAUGUCAGCAGGGUCCCUUUAAAGUGAAUCGAAUGUGCGAUUUAUACAGCCGCUUCACGGCCUU ....((((..........((((((((((((......))...((((.....)))).))).)))))))..)))).......((((((((......))))))))...((((.....))))... ( -32.30) >DroSim_CAF1 14883 120 - 1 AAAAGGACAAAUAUUGCAUGCUGACUUCGGAAAAUGCCAAAGGUACUUGCUAUCAGAAUGUCAGCAGGGUCCCUUUAAAGUGAAUCGAAUGUGCGAUUUAUACAGCCGCUUCACGGCCUU ....((((..........((((((((((((......))...((((.....)))).))).)))))))..)))).......((((((((......))))))))...((((.....))))... ( -32.30) >DroEre_CAF1 20627 119 - 1 AAAAGGACA-AUAUUGAAUGCUGACUUCGGAAAAUGCCGAAGGUACUCGCUAUCAGAAUGUCAGCAGGGCGCCUUUAAAGUGAAUCGAAUGUGCGAUGCAUACAGCCGCUUCACGGCCCU .........-........((((((((((((......)))))((((.....)))).....)))))))((((.........(((((.((..((((.......))))..)).))))).)))). ( -32.30) >DroYak_CAF1 22052 120 - 1 AAAAGGACAAAUAUUGAAUGCUGACUUCGGAAAAUGCCGAAGGUACUCGCUAUCAGAAUGUCAGCAGGGUCCCUUUAAAGUGAAUCGAAUGUGCAAUUUAUACAGCCGCUUCACGGCCUU ....((((..........((((((((((((......)))))((((.....)))).....)))))))..)))).......(((((.((..((((.......))))..)).)))))...... ( -31.30) >consensus AAAAGGACAAAUAUUGCAUGCUGACUUCGGAAAAUGCCAAAGGUACUUGCUAUCAGAAUGUCAGCAGGGUCCCUUUAAAGUGAAUCGAAUGUGCGAUUUAUACAGCCGCUUCACGGCCUU .(((((((..........((((((((((((......))...((((.....)))).))).)))))))..)).)))))...((((((((......))))))))...((((.....))))... (-28.68 = -29.32 + 0.64)

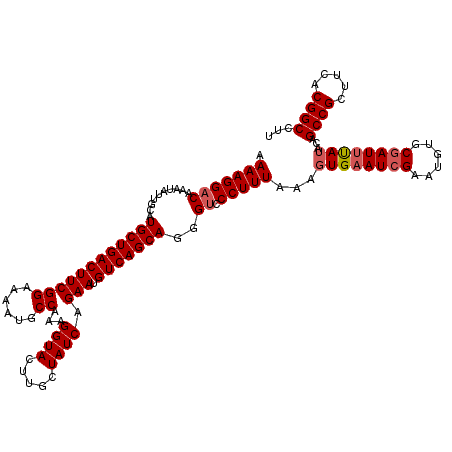
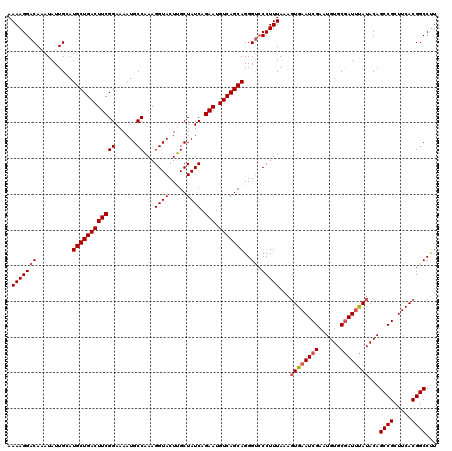
| Location | 16,192,174 – 16,192,291 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 91.49 |
| Mean single sequence MFE | -35.62 |
| Consensus MFE | -28.55 |
| Energy contribution | -29.55 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.50 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.930056 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
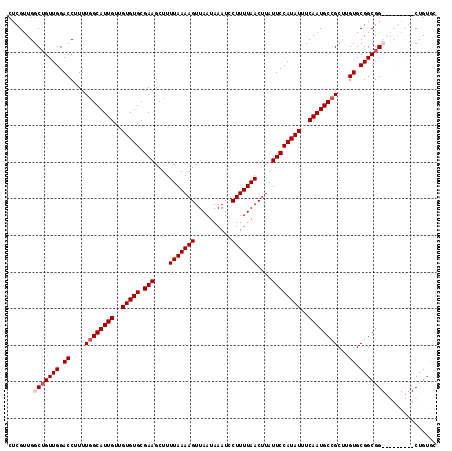
>3R_DroMel_CAF1 16192174 117 + 27905053 CUCGUUGGCUGUUGGACCUUUUGGCAUUGUUGUGUGCGAAGCUUUUAAAAGUUAAUAAAUCCUUUUAACUUAUUCCAUAUUUCAAUGCCGCUUGUGCGGCGGCGGCGGCGGCUGUGC .(((((((((((((.((....((((((((..(((((.(((....(((((((..........)))))))....))))))))..))))))))...)).)))))))..))))))...... ( -38.30) >DroSec_CAF1 21479 117 + 1 CUCGUUGGCUGUUGGACCUUUUGGCAUUGUUGUGUGCGAAGCUUUUAAAAGUUAAUAAAUCCUUUUAACUUAUUCCAUAUUUCAAUGCCGCUUGUGCGGCGGCGGCGGCGGCUGUGC .(((((((((((((.((....((((((((..(((((.(((....(((((((..........)))))))....))))))))..))))))))...)).)))))))..))))))...... ( -38.30) >DroEre_CAF1 20746 106 + 1 CUCGCUGGCUGUUGGACCUUUU-GCAUUGUUGUGUGCGAAGCUUUUAAAAGUUAAUAAAUCCUUUUAACUUAUUCCAUAUUUCAAUGCCGC-UGUGCGGCGG---------CUGUGU ..(((.((((((((.((.....-((((((..(((((.(((....(((((((..........)))))))....))))))))..))))))...-.)).))))))---------))))). ( -32.80) >DroYak_CAF1 22172 105 + 1 CUCGUUGGCGGUUGGACCUUUUGGCAUUGUUGUGUGCGAAGCUUUUAAAAGUUAAUAAAUCCUUUUAACUUAUUCCAUAUUUCAAUGCCGCUUGUGCGGCGG---------CUG--- ......(((.((((.((....((((((((..(((((.(((....(((((((..........)))))))....))))))))..))))))))...)).)))).)---------)).--- ( -33.10) >consensus CUCGUUGGCUGUUGGACCUUUUGGCAUUGUUGUGUGCGAAGCUUUUAAAAGUUAAUAAAUCCUUUUAACUUAUUCCAUAUUUCAAUGCCGCUUGUGCGGCGG_________CUGUGC .......(((((((.((....((((((((..(((((.(((....(((((((..........)))))))....))))))))..))))))))...)).))))))).............. (-28.55 = -29.55 + 1.00)

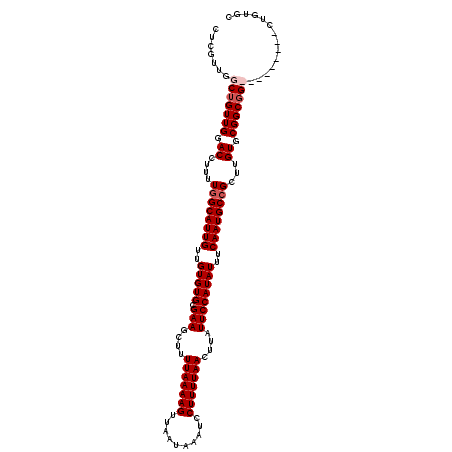
| Location | 16,192,254 – 16,192,371 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 86.72 |
| Mean single sequence MFE | -34.25 |
| Consensus MFE | -24.23 |
| Energy contribution | -25.35 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.566444 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
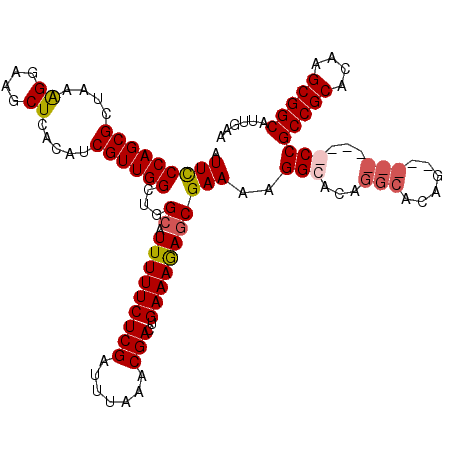
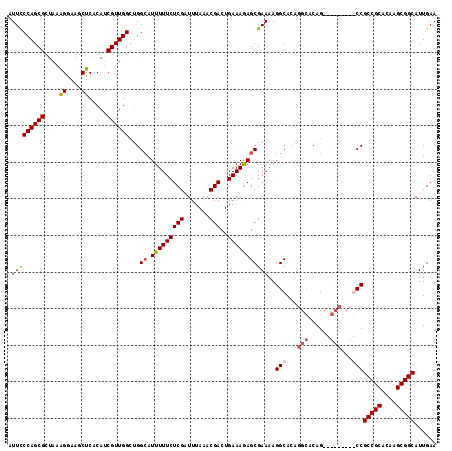
>3R_DroMel_CAF1 16192254 117 - 27905053 AUUCCCAGCGCUAAGGGAAGCUCACAUCGUUGGCUGGAAUUUUUCUCGAUUUAAACGACUGAAAGAGCGAAAAGGCACAGGCACAGCCGCCGCCGCCGCCGCACAAGCGGCAUUGAA .(((((........)))))((((...((((((....(((((......)))))...)))).))..)))).....(((...(((......)))...)))(((((....)))))...... ( -38.10) >DroSec_CAF1 21559 117 - 1 AUUCCCAGCGCUAAAGAAAGCUCACAUCGUUGGGUGGCAUUUUUCUCGAUUUAAACGACUGAAAAAGCGAAAAGGCACAGGCACAGCCGCCGCCGCCGCCGCACAAGCGGCAUUGAA ...(((((((.....((....))....))))))).(((.(((((((((.......)))..)))))).......(((...(((...)))...))))))(((((....)))))...... ( -37.50) >DroEre_CAF1 20825 107 - 1 AUUUCCAGCGCCAAAGGAACCUCACAUCGUUGGCUGGCAUUUUUCUCGAUUUAAACGACUGAAAGAGCGAAAAGGCACAGACACAG---------CCGCCGCACA-GCGGCAUUGAA ....((((((....((....)).....))))))(((((.((((((((..((((......)))).))).))))).)).))).....(---------((((......-)))))...... ( -27.20) >DroYak_CAF1 22252 102 - 1 AUUCCCAGCGCUAAAGGAAGCUCACAGCGUUGGCAGGCAUUUUUCUCGAUUUAAGCGACUGAAAGAGCGAAAAGGCA------CAG---------CCGCCGCACAAGCGGCAUUGAA ....((((((((..((....))...))))))))..(((.(((((((((.......)))..))))))((......)).------..)---------))(((((....)))))...... ( -34.20) >consensus AUUCCCAGCGCUAAAGGAAGCUCACAUCGUUGGCUGGCAUUUUUCUCGAUUUAAACGACUGAAAGAGCGAAAAGGCACAGGCACAG_________CCGCCGCACAAGCGGCAUUGAA .(((((((((....((....)).....))))))...((.(((((((((.......)))..)))))))))))..(((...(((......)))...)))(((((....)))))...... (-24.23 = -25.35 + 1.12)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:15:19 2006