
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 16,190,166 – 16,190,326 |
| Length | 160 |
| Max. P | 0.999241 |

| Location | 16,190,166 – 16,190,286 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.67 |
| Mean single sequence MFE | -33.76 |
| Consensus MFE | -31.18 |
| Energy contribution | -31.78 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.38 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.809988 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
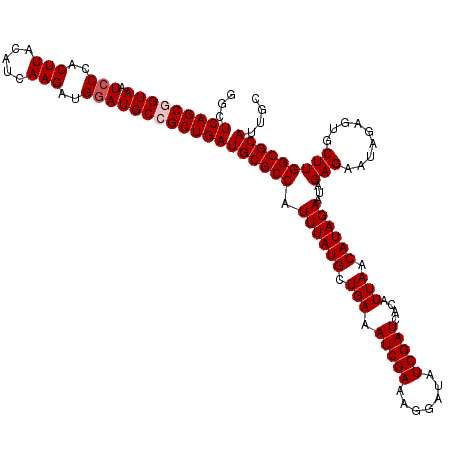
>3R_DroMel_CAF1 16190166 120 + 27905053 GGCUCAGCGGUAAUCCCACUUACAUCAAGAUGAAUGCCGCUGAUGCGCCAUUUAUGCUGAAAUCGAAAGGAUAUCGAUCACAUUAACAUAGAAUAGAGAAUAGAGUGCUUGGCGCAUUGC ...(((((((((.((...(((.....)))..)).)))))))))(((((((..(((.(((....(....)(((....))).....................))).)))..))))))).... ( -32.80) >DroSec_CAF1 19365 120 + 1 GGCUCAGCGGUAAUCCCACUUACAUCAAGAUGGAUGCCGCUGAUGCGCCAUUUAUGCUGAAAUCGAAAGGAUAUCGAUCACAUUAACAUAGAAUAGAGAAUAGAGUGCUUGGCGCAUUGC ...(((((((((.(((..(((.....)))..))))))))))))(((((((..(((.(((....(....)(((....))).....................))).)))..))))))).... ( -35.40) >DroSim_CAF1 12928 120 + 1 GGCUCAGCGGUAAUCCCACUUACAUCAAGAUGGAUGCCGCUGAUGCGCCAUUUAUGCUGAAAUCGAAAGGAUAUCGAUCACAUUAACAUAGAAUAGAGAAUAGAGUGCUUGGCGCAUUGC ...(((((((((.(((..(((.....)))..))))))))))))(((((((..(((.(((....(....)(((....))).....................))).)))..))))))).... ( -35.40) >DroEre_CAF1 18725 120 + 1 GGCUCAGCCGUAAUCCCACUUACAUCAAGAUGGAUGCCGCUGAUGCGCCAUUUAUGCUGAAAUCGAAAGGAUAUCGAUCACAUUAACAUAGAAUAGAGAACAGGGCGCUUGGCGCAUUGC ...(((((.(((.(((..(((.....)))..)))))).)))))(((((((....((((.....(....)(((....)))........................))))..))))))).... ( -29.50) >DroYak_CAF1 20065 120 + 1 GGCUCAGCCGUAAUCCCACUUACAUCAAGAUGGAUGCCGCUGAUGCGCCAUUUAUGCUGAAAUCGAAAGGAUAUCGAUCACAUUAGCAUAGAAUAGAGAAUAGAGCGCUUGGCGCAUUGC ...(((((.(((.(((..(((.....)))..)))))).)))))((((((.((((((((((.(((((.......)))))....))))))))))...(((.........))))))))).... ( -35.70) >consensus GGCUCAGCGGUAAUCCCACUUACAUCAAGAUGGAUGCCGCUGAUGCGCCAUUUAUGCUGAAAUCGAAAGGAUAUCGAUCACAUUAACAUAGAAUAGAGAAUAGAGUGCUUGGCGCAUUGC ...(((((((((.(((..(((.....)))..))))))))))))((((((.((((((.(((.(((((.......)))))....))).))))))...(((.........))))))))).... (-31.18 = -31.78 + 0.60)


| Location | 16,190,166 – 16,190,286 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.67 |
| Mean single sequence MFE | -36.10 |
| Consensus MFE | -35.18 |
| Energy contribution | -35.26 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.97 |
| SVM decision value | 3.46 |
| SVM RNA-class probability | 0.999241 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
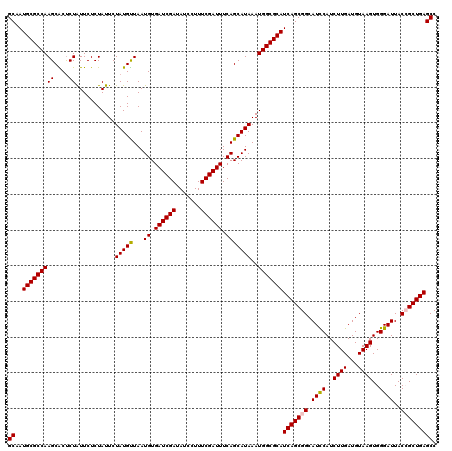
>3R_DroMel_CAF1 16190166 120 - 27905053 GCAAUGCGCCAAGCACUCUAUUCUCUAUUCUAUGUUAAUGUGAUCGAUAUCCUUUCGAUUUCAGCAUAAAUGGCGCAUCAGCGGCAUUCAUCUUGAUGUAAGUGGGAUUACCGCUGAGCC ((..(((((((((....))...........(((((...((.((((((.......)))))).)))))))..)))))))(((((((...((..((((...))))...))...))))))))). ( -35.10) >DroSec_CAF1 19365 120 - 1 GCAAUGCGCCAAGCACUCUAUUCUCUAUUCUAUGUUAAUGUGAUCGAUAUCCUUUCGAUUUCAGCAUAAAUGGCGCAUCAGCGGCAUCCAUCUUGAUGUAAGUGGGAUUACCGCUGAGCC ((..(((((((((....))...........(((((...((.((((((.......)))))).)))))))..)))))))(((((((.((((..((((...))))..))))..))))))))). ( -37.80) >DroSim_CAF1 12928 120 - 1 GCAAUGCGCCAAGCACUCUAUUCUCUAUUCUAUGUUAAUGUGAUCGAUAUCCUUUCGAUUUCAGCAUAAAUGGCGCAUCAGCGGCAUCCAUCUUGAUGUAAGUGGGAUUACCGCUGAGCC ((..(((((((((....))...........(((((...((.((((((.......)))))).)))))))..)))))))(((((((.((((..((((...))))..))))..))))))))). ( -37.80) >DroEre_CAF1 18725 120 - 1 GCAAUGCGCCAAGCGCCCUGUUCUCUAUUCUAUGUUAAUGUGAUCGAUAUCCUUUCGAUUUCAGCAUAAAUGGCGCAUCAGCGGCAUCCAUCUUGAUGUAAGUGGGAUUACGGCUGAGCC ((..((((((((((.....)))........(((((...((.((((((.......)))))).)))))))..)))))))(((((.(.((((..((((...))))..))))..).))))))). ( -34.50) >DroYak_CAF1 20065 120 - 1 GCAAUGCGCCAAGCGCUCUAUUCUCUAUUCUAUGCUAAUGUGAUCGAUAUCCUUUCGAUUUCAGCAUAAAUGGCGCAUCAGCGGCAUCCAUCUUGAUGUAAGUGGGAUUACGGCUGAGCC ((..(((((((((....))...........((((((.....((((((.......))))))..))))))..)))))))(((((.(.((((..((((...))))..))))..).))))))). ( -35.30) >consensus GCAAUGCGCCAAGCACUCUAUUCUCUAUUCUAUGUUAAUGUGAUCGAUAUCCUUUCGAUUUCAGCAUAAAUGGCGCAUCAGCGGCAUCCAUCUUGAUGUAAGUGGGAUUACCGCUGAGCC ((..(((((((((....))...........(((((...((.((((((.......)))))).)))))))..)))))))(((((((.((((..((((...))))..))))..))))))))). (-35.18 = -35.26 + 0.08)


| Location | 16,190,206 – 16,190,326 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.50 |
| Mean single sequence MFE | -23.06 |
| Consensus MFE | -22.98 |
| Energy contribution | -22.82 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.42 |
| Structure conservation index | 1.00 |
| SVM decision value | 1.08 |
| SVM RNA-class probability | 0.911684 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16190206 120 - 27905053 UUAACAAUCUAGUAUCUGUAGCUCUAUGCAAAUCAAAUCUGCAAUGCGCCAAGCACUCUAUUCUCUAUUCUAUGUUAAUGUGAUCGAUAUCCUUUCGAUUUCAGCAUAAAUGGCGCAUCA ..........................((((.........))))((((((((((....))...........(((((...((.((((((.......)))))).)))))))..)))))))).. ( -22.50) >DroSec_CAF1 19405 120 - 1 UUAACAAUCUAGUAUCUGUAGCUCUAUGCAAAUCAAAUCUGCAAUGCGCCAAGCACUCUAUUCUCUAUUCUAUGUUAAUGUGAUCGAUAUCCUUUCGAUUUCAGCAUAAAUGGCGCAUCA ..........................((((.........))))((((((((((....))...........(((((...((.((((((.......)))))).)))))))..)))))))).. ( -22.50) >DroSim_CAF1 12968 120 - 1 UUAACAAUCUAGUAUCUGUAGCUCUAUGCAAAUCAAAUCUGCAAUGCGCCAAGCACUCUAUUCUCUAUUCUAUGUUAAUGUGAUCGAUAUCCUUUCGAUUUCAGCAUAAAUGGCGCAUCA ..........................((((.........))))((((((((((....))...........(((((...((.((((((.......)))))).)))))))..)))))))).. ( -22.50) >DroEre_CAF1 18765 120 - 1 UUAACAAUCUAGUAUCUGUAGCUCUAUGCAAAUCAAAUCUGCAAUGCGCCAAGCGCCCUGUUCUCUAUUCUAUGUUAAUGUGAUCGAUAUCCUUUCGAUUUCAGCAUAAAUGGCGCAUCA ..........................((((.........))))(((((((((((.....)))........(((((...((.((((((.......)))))).)))))))..)))))))).. ( -23.50) >DroYak_CAF1 20105 120 - 1 UUAACAAUCUAGUAUCUGUAGCUCUAUGCAAAUCAAAUCUGCAAUGCGCCAAGCGCUCUAUUCUCUAUUCUAUGCUAAUGUGAUCGAUAUCCUUUCGAUUUCAGCAUAAAUGGCGCAUCA ..........................((((.........))))((((((((((....))...........((((((.....((((((.......))))))..))))))..)))))))).. ( -24.30) >consensus UUAACAAUCUAGUAUCUGUAGCUCUAUGCAAAUCAAAUCUGCAAUGCGCCAAGCACUCUAUUCUCUAUUCUAUGUUAAUGUGAUCGAUAUCCUUUCGAUUUCAGCAUAAAUGGCGCAUCA ..........................((((.........))))((((((((((....))...........(((((...((.((((((.......)))))).)))))))..)))))))).. (-22.98 = -22.82 + -0.16)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:15:08 2006