
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 16,178,640 – 16,178,782 |
| Length | 142 |
| Max. P | 0.671653 |

| Location | 16,178,640 – 16,178,742 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 76.40 |
| Mean single sequence MFE | -34.58 |
| Consensus MFE | -18.38 |
| Energy contribution | -19.72 |
| Covariance contribution | 1.34 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.53 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.605322 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16178640 102 - 27905053 AGGA--AACUUAAAUAAAAGUCGGCCAGAAGUUGUCUGCUCCUGGGAUUGCCCUCGAAAGGGGCGGCACCAGUGGGAAAACUGCGUGGG----------------GUAAACGUGGAGCUA ((..--..))............(((((((.....))))(((((((..((((((.(....)))))))..)))).)))....(..(((...----------------....)))..).))). ( -34.60) >DroPse_CAF1 8203 113 - 1 AGGUAGAACUGAAAUAAAAGGCAAGCAAAAGUGGUCUGCUCUUGGU-----C--CCAGAGAGGCAAGGAUUGGAGGAAAACUGCGUGCGUGUGAAAUAGUUGUGGACUCACAAACGGCAC ........(((.........(((.(((...(..((...(..(..((-----(--(..(.....)..))))..)..)...))..).))).))).......(((((....))))).)))... ( -23.50) >DroSec_CAF1 7912 102 - 1 AGGA--AACUUAAAUAAAAGUCGGCCAGAAGAUGUCUGCUCCUGGGAUUGCCCUCGAAAGGGGCGGGGCCAGUGGGAAAACUGCGUGGG----------------GAAAAUGUGCGGCUG ((..--..))........(((((..((((.....))))(((((((..((((((.(....)))))))..)))(..(.....)..)..)))----------------)........))))). ( -33.60) >DroSim_CAF1 1319 102 - 1 AGGA--AACUUAAAUAAAAGUCGGCCAGAAGUUGUCUGCUCCUGGGAUUGCCCUUGAAAGGGGCGGGGACAGUGGGAAAACUGCGUGGG----------------GUAAACGUGCGGCUG ((..--..))...........(((((...((((.....((((((...(((((((.....)))))))...))).)))..))))(((((..----------------.....)))))))))) ( -31.60) >DroEre_CAF1 6794 103 - 1 AGGA-AAACUUAAAUAAAAGUCGGGCAGAAGUUGCCUGCUCCUGGGAUGGCCCUCGAAAGGGGCGGGGCCAGCGGGAAAACUGCGUGGG----------------GCAAGCUACCGGCGG ....-..............(((((((....)).((.(((((((((..(.((((.(....))))).)..)))((((.....))))..)))----------------))).))..))))).. ( -40.70) >DroYak_CAF1 8756 103 - 1 AGGA-AAACUUAAAUAAAAGUCGGGCAGAAGUUGUCUGCUCCUGGGACUGCCCUCGAAAGGGGCGGGGCCAGCGGGAAAACUGCGUGGG----------------GCGAGCUACCGGCUG ....-.............((((((((....)).(..(((((((((..((((((.(....)))))))..)))((((.....))))..)))----------------)))..)..)))))). ( -43.50) >consensus AGGA__AACUUAAAUAAAAGUCGGCCAGAAGUUGUCUGCUCCUGGGAUUGCCCUCGAAAGGGGCGGGGCCAGUGGGAAAACUGCGUGGG________________GCAAACGAGCGGCUG ...................((((...........((((((((((((....))).....))))))))).(((((((.....)))).)))..........................)))).. (-18.38 = -19.72 + 1.34)



| Location | 16,178,664 – 16,178,782 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 88.88 |
| Mean single sequence MFE | -31.76 |
| Consensus MFE | -23.45 |
| Energy contribution | -23.85 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.74 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16178664 118 + 27905053 UUUUCCCACUGGUGCCGCCCCUUUCGAGGGCAAUCCCAGGAGCAGACAACUUCUGGCCGACUUUUAUUUAAGUU-UCCUUCACCUUUCCAGAACUCAGAACUAAGGACUCGGGACUUAG ...((((...((((..(((((....).))))....(((((((.......)))))))..(((((......)))))-.....))))..(((...............)))...))))..... ( -28.76) >DroSec_CAF1 7936 111 + 1 UUUUCCCACUGGCCCCGCCCCUUUCGAGGGCAAUCCCAGGAGCAGACAUCUUCUGGCCGACUUUUAUUUAAGUU-UCCUUCACCUUUCCAGAA-------CUCAGGACUCGGGACUUAG ...((((.((((....(((((....).))))....))))((((.......((((((..(((((......)))))-............))))))-------.....).)))))))..... ( -27.54) >DroSim_CAF1 1343 118 + 1 UUUUCCCACUGUCCCCGCCCCUUUCAAGGGCAAUCCCAGGAGCAGACAACUUCUGGCCGACUUUUAUUUAAGUU-UCCUUCACCUUUCCAGAACUCAGAACUCAGGACUCGGGACUUAG ...((((...((((..((((.......))))....(((((((.......)))))))..(((((......)))))-.............................))))..))))..... ( -28.70) >DroEre_CAF1 6818 119 + 1 UUUUCCCGCUGGCCCCGCCCCUUUCGAGGGCCAUCCCAGGAGCAGGCAACUUCUGCCCGACUUUUAUUUAAGUUUUCCUUCACCUUGCCGGAAUUCAGAACUGAGGACUAGGGACUUAG ...((((..((((((((.......)).))))))((((((((((.((((.....)))).).)))........(..((((..(.....)..))))..)....))).)))...))))..... ( -35.30) >DroYak_CAF1 8780 112 + 1 UUUUCCCGCUGGCCCCGCCCCUUUCGAGGGCAGUCCCAGGAGCAGACAACUUCUGCCCGACUUUUAUUUAAGUUUUCCUUCACCUU-------UUCAGAACUGAGGACUCGGGGCUUAG ..........(((((((((((....).))))(((((((((.(((((.....))))).)(((((......)))))............-------.......))).))))).))))))... ( -38.50) >consensus UUUUCCCACUGGCCCCGCCCCUUUCGAGGGCAAUCCCAGGAGCAGACAACUUCUGGCCGACUUUUAUUUAAGUU_UCCUUCACCUUUCCAGAACUCAGAACUCAGGACUCGGGACUUAG ...((((.((((....((((.......))))....))))(((((((.....))))...(((((......))))).(((((......................))))))))))))..... (-23.45 = -23.85 + 0.40)

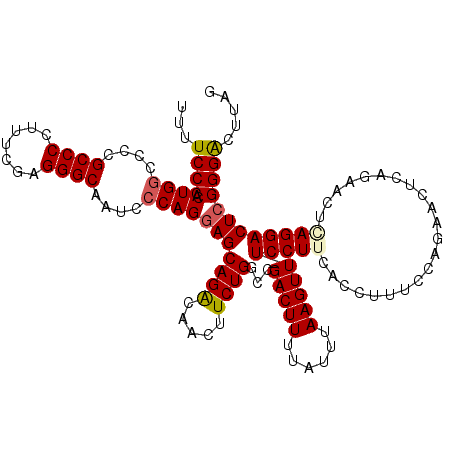

| Location | 16,178,664 – 16,178,782 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 88.88 |
| Mean single sequence MFE | -40.53 |
| Consensus MFE | -27.56 |
| Energy contribution | -27.72 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.671653 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16178664 118 - 27905053 CUAAGUCCCGAGUCCUUAGUUCUGAGUUCUGGAAAGGUGAAGGA-AACUUAAAUAAAAGUCGGCCAGAAGUUGUCUGCUCCUGGGAUUGCCCUCGAAAGGGGCGGCACCAGUGGGAAAA .....((((((((....(.(((((.(((((....))...(((..-..)))...........)))))))).).....))))((((..((((((.(....)))))))..)))).))))... ( -40.50) >DroSec_CAF1 7936 111 - 1 CUAAGUCCCGAGUCCUGAG-------UUCUGGAAAGGUGAAGGA-AACUUAAAUAAAAGUCGGCCAGAAGAUGUCUGCUCCUGGGAUUGCCCUCGAAAGGGGCGGGGCCAGUGGGAAAA .....((((((((......-------((((((...(((.(((..-..)))........)))..)))))).......))))((((..((((((.(....)))))))..)))).))))... ( -38.02) >DroSim_CAF1 1343 118 - 1 CUAAGUCCCGAGUCCUGAGUUCUGAGUUCUGGAAAGGUGAAGGA-AACUUAAAUAAAAGUCGGCCAGAAGUUGUCUGCUCCUGGGAUUGCCCUUGAAAGGGGCGGGGACAGUGGGAAAA (((.(((((.((((((((((..(((.((((((...(((.(((..-..)))........)))..)))))).)))...))))..))))))(((((.....))))).)))))..)))..... ( -40.70) >DroEre_CAF1 6818 119 - 1 CUAAGUCCCUAGUCCUCAGUUCUGAAUUCCGGCAAGGUGAAGGAAAACUUAAAUAAAAGUCGGGCAGAAGUUGCCUGCUCCUGGGAUGGCCCUCGAAAGGGGCGGGGCCAGCGGGAAAA .....((((..........(((((...((((((......(((.....)))........)))))))))))((((((((((((((((....))).....)))))))))..))))))))... ( -39.54) >DroYak_CAF1 8780 112 - 1 CUAAGCCCCGAGUCCUCAGUUCUGAA-------AAGGUGAAGGAAAACUUAAAUAAAAGUCGGGCAGAAGUUGUCUGCUCCUGGGACUGCCCUCGAAAGGGGCGGGGCCAGCGGGAAAA ......((((..((((((..(((...-------.))))).)))).................(((((((.....)))))))((((..((((((.(....)))))))..)))))))).... ( -43.90) >consensus CUAAGUCCCGAGUCCUCAGUUCUGAAUUCUGGAAAGGUGAAGGA_AACUUAAAUAAAAGUCGGCCAGAAGUUGUCUGCUCCUGGGAUUGCCCUCGAAAGGGGCGGGGCCAGUGGGAAAA ....(((((.(((((...................((((........))))..............(((.(((.....))).))))))))((((.(....))))))))))........... (-27.56 = -27.72 + 0.16)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:14:45 2006