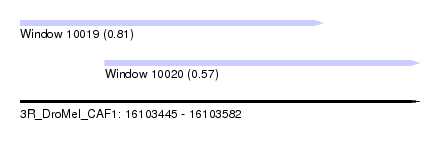

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 16,103,445 – 16,103,582 |
| Length | 137 |
| Max. P | 0.810991 |

| Location | 16,103,445 – 16,103,549 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 77.29 |
| Mean single sequence MFE | -24.38 |
| Consensus MFE | -12.44 |
| Energy contribution | -12.68 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.51 |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.810991 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16103445 104 + 27905053 UUAGCUGAUGACGCUGACCAGCAUA---GC--ACUCAGUGCUCACUUUCUCAUCCUGUG-----AUGCCUCUCAUCCGAUGCUUCGCUAUCGCCAUCGGCAUCGCCAUCCUUGA ......(((((.(((....)))..(---((--((...))))).......)))))..(((-----(((((.......((((((...)).)))).....))))))))......... ( -26.50) >DroSec_CAF1 54072 84 + 1 -UAGCUGAUGACGCUGACCAGCACA---GCACACUCAAUGCUCACUUUCUCAUCCUGCG-----AUGCCUCUC---------------------AUCGGCAUCGCCAUCCUUGA -.....(((((.(((....)))..(---(((.......)))).......)))))..(((-----(((((....---------------------...))))))))......... ( -23.50) >DroSim_CAF1 60170 108 + 1 -UAGCUGAUGACGCUGACCAGCACAGCAGCACACUCAGUGCUCACUUUCUCAUCCUGCG-----AUGCCUCUCGUCCGAUGCUUCGCUAUCGCCAUCGGCAUCGCCAUCCUUGA -.....(((((.((((.......))))(((((.....))))).......)))))..(((-----(((((.......((((((...)).)))).....))))))))......... ( -31.80) >DroEre_CAF1 30161 98 + 1 ---GCUGAUGACGCUGACCAGC----------ACUCAG---UCACUUCCUCAUCCUUCAUCCUCAUGCCUCUCAUCGGAUGCUUCGCUAUCACCAUCGGCAUCGCCAUCCUUGA ---(((((((..(((((.....----------..))))---)...............(((((..(((.....))).)))))............))))))).............. ( -19.10) >DroYak_CAF1 72855 93 + 1 ---GCUGAUGACGCUGACCAGC----------ACUCAG---UCAUUUCCUCAUCCUUCG-----GUGCCUCUCAUCCGAUGCGUCGCUAUCACCAUCAGCAUCGCCAUCCUUGA ---(((((((..(((((.....----------..))))---)...............((-----..((.((......)).))..)).......))))))).............. ( -21.00) >consensus _UAGCUGAUGACGCUGACCAGCA_A___GC__ACUCAGUGCUCACUUUCUCAUCCUGCG_____AUGCCUCUCAUCCGAUGCUUCGCUAUCACCAUCGGCAUCGCCAUCCUUGA ...(((((((..(((....))).......................................................((((......))))..))))))).............. (-12.44 = -12.68 + 0.24)

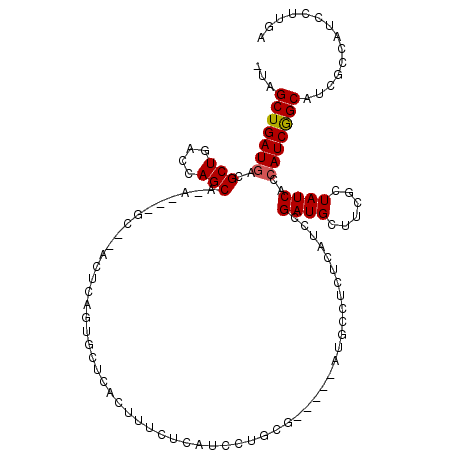

| Location | 16,103,474 – 16,103,582 |
|---|---|
| Length | 108 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 80.71 |
| Mean single sequence MFE | -19.25 |
| Consensus MFE | -12.24 |
| Energy contribution | -13.17 |
| Covariance contribution | 0.94 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.573682 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16103474 108 + 27905053 UCAGUGCUCACUUUCUCAUCCUGUGAUGCCUCUCAUCCGAUGCUUCGCUAUCGCCAUCGGCAUCGCCAUCCUUGAACACACGCUUCAGAUACAAGUAUAUUUGCCCCC ..((((......(((.......((((((((.......((((((...)).)))).....)))))))).......)))....)))).((((((......))))))..... ( -18.84) >DroSec_CAF1 54102 87 + 1 UCAAUGCUCACUUUCUCAUCCUGCGAUGCCUCUC---------------------AUCGGCAUCGCCAUCCUUGAACACACGCUUCAGAUACAAGUAUAUUUGCCCCC .....((.....(((.......((((((((....---------------------...)))))))).......))).....))..((((((......))))))..... ( -17.74) >DroSim_CAF1 60203 108 + 1 UCAGUGCUCACUUUCUCAUCCUGCGAUGCCUCUCGUCCGAUGCUUCGCUAUCGCCAUCGGCAUCGCCAUCCUUGAACACACGCUUCAGAUACAAGUGUAUUUGCCCCC ..((((......(((.......((((((((.......((((((...)).)))).....)))))))).......)))....)))).(((((((....)))))))..... ( -24.44) >DroYak_CAF1 72876 99 + 1 UCAG---UCAUUUCCUCAUCCUUCGGUGCCUCUCAUCCGAUGCGUCGCUAUCACCAUCAGCAUCGCCAUCCUUGAACAGACGCUUCUGAUA------UAUUUGCCCCC ....---.................((.((.....(((.((.((((((((.........))).....((....))....))))).)).))).------.....)))).. ( -16.00) >consensus UCAGUGCUCACUUUCUCAUCCUGCGAUGCCUCUCAUCCGAUGCUUCGCUAUCGCCAUCGGCAUCGCCAUCCUUGAACACACGCUUCAGAUACAAGUAUAUUUGCCCCC ...............(((....((((((((.......(((((......))))).....))))))))......)))..........((((((......))))))..... (-12.24 = -13.17 + 0.94)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:13:52 2006