
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 16,061,199 – 16,061,337 |
| Length | 138 |
| Max. P | 0.994721 |

| Location | 16,061,199 – 16,061,298 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 91.75 |
| Mean single sequence MFE | -32.58 |
| Consensus MFE | -29.14 |
| Energy contribution | -29.45 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.64 |
| SVM RNA-class probability | 0.969138 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16061199 99 + 27905053 ACGGCUGAAAUUCAGCAAUAAAUUGCCGUGAGUUAGUCAGA-GUGGAGCAUGCUUCAUUAAUGCGACAAGCGUGAGAGGCGAACCUCUCACCCCAA-------UCCC ..((((((....(.((((....)))).)....))))))..(-((((((....)))))))...((.....))((((((((....)))))))).....-------.... ( -31.20) >DroSec_CAF1 21559 98 + 1 ACGGCUGAAAUUCAGCAA-AAAUUGCCGUGAGUGAGUCAGA-GUGGAGCAUGCUUCCUUAAUGCGACAAGCGUGAGAGGCUAACCUCUCACCCCAC-------ACCC ..((((((.((((((((.-....)))..)))))...)))).-((((.((((.........)))).......((((((((....)))))))).))))-------.)). ( -33.10) >DroSim_CAF1 19986 99 + 1 ACGGCUGAAAUUCAGCAAUAAAUUGCCGUGAGUGAGUCAGA-GUGGAGCAUGCUUCAUUAAUGCGACAAGCGUGAGAGGCUAACCUCUCACCCCAC-------ACCC ..((((((.(((((((((....))))..)))))...)))).-((((.((((.........)))).......((((((((....)))))))).))))-------.)). ( -34.00) >DroYak_CAF1 27606 107 + 1 ACAGCUGAAAUUCAGCAAUAAAUUGCCGUGAGUGAGUCAGAGGUGGAGCAUGCUUCAUUAAUGCGACAAGCGUGAGGGGCUAACCUCGCACCCCACACCACACACCC ...((((.....))))...........(((.(((.(((...(((((((....))))))).....)))....((((((......))))))......))))))...... ( -32.00) >consensus ACGGCUGAAAUUCAGCAAUAAAUUGCCGUGAGUGAGUCAGA_GUGGAGCAUGCUUCAUUAAUGCGACAAGCGUGAGAGGCUAACCUCUCACCCCAC_______ACCC ....((((.(((((((((....))))..)))))...))))..((((.((((.........)))).......((((((((....)))))))).))))........... (-29.14 = -29.45 + 0.31)



| Location | 16,061,199 – 16,061,298 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 91.75 |
| Mean single sequence MFE | -34.65 |
| Consensus MFE | -29.08 |
| Energy contribution | -30.14 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.824960 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16061199 99 - 27905053 GGGA-------UUGGGGUGAGAGGUUCGCCUCUCACGCUUGUCGCAUUAAUGAAGCAUGCUCCAC-UCUGACUAACUCACGGCAAUUUAUUGCUGAAUUUCAGCCGU .(((-------.((((((((((((....))))))))((.(((..((....))..))).)))))).-))).........(((((((((((....))))))...))))) ( -33.20) >DroSec_CAF1 21559 98 - 1 GGGU-------GUGGGGUGAGAGGUUAGCCUCUCACGCUUGUCGCAUUAAGGAAGCAUGCUCCAC-UCUGACUCACUCACGGCAAUUU-UUGCUGAAUUUCAGCCGU ((((-------(.(((((((((((....))))))))((((.((.......))))))...))))))-))..........((((((((((-.....)))))...))))) ( -34.50) >DroSim_CAF1 19986 99 - 1 GGGU-------GUGGGGUGAGAGGUUAGCCUCUCACGCUUGUCGCAUUAAUGAAGCAUGCUCCAC-UCUGACUCACUCACGGCAAUUUAUUGCUGAAUUUCAGCCGU .(((-------(((((((((((((....))))))))((.(((..((....))..))).)).....-.....)))))...((((((....)))))).......))).. ( -35.70) >DroYak_CAF1 27606 107 - 1 GGGUGUGUGGUGUGGGGUGCGAGGUUAGCCCCUCACGCUUGUCGCAUUAAUGAAGCAUGCUCCACCUCUGACUCACUCACGGCAAUUUAUUGCUGAAUUUCAGCUGU (((((.((((((..(.((((((((......))))..((.....)).........)))).)..))))....)).))))).((((((....))))))............ ( -35.20) >consensus GGGU_______GUGGGGUGAGAGGUUAGCCUCUCACGCUUGUCGCAUUAAUGAAGCAUGCUCCAC_UCUGACUCACUCACGGCAAUUUAUUGCUGAAUUUCAGCCGU ((((.......(((((((((((((....))))))))((.(((..(......)..))).))))))).........))))(((((((((((....))))))...))))) (-29.08 = -30.14 + 1.06)


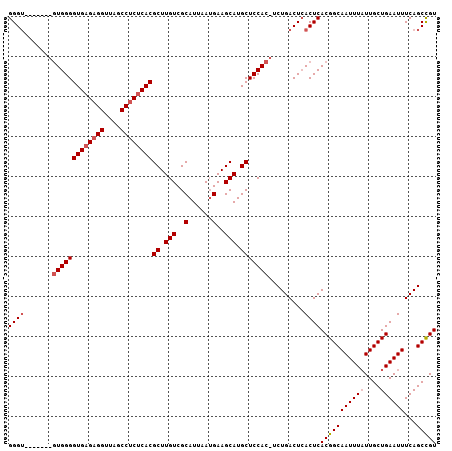
| Location | 16,061,239 – 16,061,337 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 86.44 |
| Mean single sequence MFE | -29.05 |
| Consensus MFE | -24.26 |
| Energy contribution | -24.57 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.05 |
| Mean z-score | -3.11 |
| Structure conservation index | 0.84 |
| SVM decision value | 2.50 |
| SVM RNA-class probability | 0.994721 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16061239 98 + 27905053 A-GUGGAGCAUGCUUCAUUAAUGCGACAAGCGUGAGAGGCGAACCUCUCACCCCAA-------UCCCCUUAACCCUAAAGCGGCCCACCCUUAACCCCCUACCAGC .-((((.((.(((((.......((.....))((((((((....)))))))).....-------..............))))))))))).................. ( -27.80) >DroSec_CAF1 21598 98 + 1 A-GUGGAGCAUGCUUCCUUAAUGCGACAAGCGUGAGAGGCUAACCUCUCACCCCAC-------ACCCCUUAACCCUAAAGCGGGCCACCCUUAACCGCCUGCCAGC .-((((.((((.........)))).......((((((((....)))))))).))))-------................((((((...........)))))).... ( -31.90) >DroSim_CAF1 20026 98 + 1 A-GUGGAGCAUGCUUCAUUAAUGCGACAAGCGUGAGAGGCUAACCUCUCACCCCAC-------ACCCCUUAACCCUAAAGCGGCCCACCCUUAACCCCCUGCCAGC .-((((.((.(((((.......((.....))((((((((....)))))))).....-------..............))))))))))).................. ( -27.80) >DroYak_CAF1 27646 99 + 1 AGGUGGAGCAUGCUUCAUUAAUGCGACAAGCGUGAGGGGCUAACCUCGCACCCCACACCACACACCCCUAAACCCUAAAGCGGCCCACCCUUUACUCCU------- .(((((.((.(((((.......((.....))(((.(((((.......)).)))))).....................))))))))))))..........------- ( -28.70) >consensus A_GUGGAGCAUGCUUCAUUAAUGCGACAAGCGUGAGAGGCUAACCUCUCACCCCAC_______ACCCCUUAACCCUAAAGCGGCCCACCCUUAACCCCCUGCCAGC ..((((.((.(((((.......((.....))((((((((....))))))))..........................))))))))))).................. (-24.26 = -24.57 + 0.31)



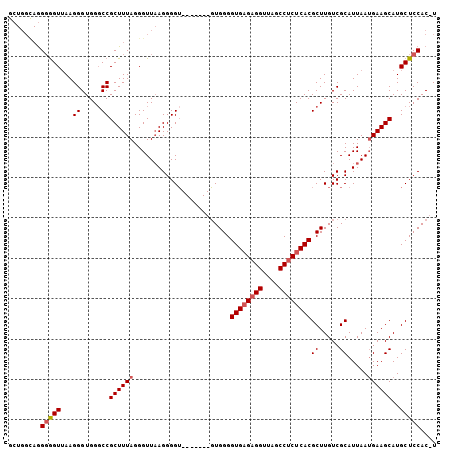
| Location | 16,061,239 – 16,061,337 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 86.44 |
| Mean single sequence MFE | -38.17 |
| Consensus MFE | -26.11 |
| Energy contribution | -26.93 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.761459 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 16061239 98 - 27905053 GCUGGUAGGGGGUUAAGGGUGGGCCGCUUUAGGGUUAAGGGGA-------UUGGGGUGAGAGGUUCGCCUCUCACGCUUGUCGCAUUAAUGAAGCAUGCUCCAC-U .................(((((((.(((((...(((((.(.((-------(.((.((((((((....)))))))).)).))).).))))))))))..)).))))-) ( -38.00) >DroSec_CAF1 21598 98 - 1 GCUGGCAGGCGGUUAAGGGUGGCCCGCUUUAGGGUUAAGGGGU-------GUGGGGUGAGAGGUUAGCCUCUCACGCUUGUCGCAUUAAGGAAGCAUGCUCCAC-U ((.(((((((.........((((((......))))))......-------.....((((((((....))))))))))))))))).....(((.(....))))..-. ( -38.80) >DroSim_CAF1 20026 98 - 1 GCUGGCAGGGGGUUAAGGGUGGGCCGCUUUAGGGUUAAGGGGU-------GUGGGGUGAGAGGUUAGCCUCUCACGCUUGUCGCAUUAAUGAAGCAUGCUCCAC-U ..((((.....))))..(((((((.(((((...(((((...((-------(.((.((((((((....)))))))).))...))).))))))))))..)).))))-) ( -36.20) >DroYak_CAF1 27646 99 - 1 -------AGGAGUAAAGGGUGGGCCGCUUUAGGGUUUAGGGGUGUGUGGUGUGGGGUGCGAGGUUAGCCCCUCACGCUUGUCGCAUUAAUGAAGCAUGCUCCACCU -------.........((((((((.((((((.........((((((.(((((((((.((.......)).)))))))))...))))))..))))))..)).)))))) ( -39.70) >consensus GCUGGCAGGGGGUUAAGGGUGGGCCGCUUUAGGGUUAAGGGGU_______GUGGGGUGAGAGGUUAGCCUCUCACGCUUGUCGCAUUAAUGAAGCAUGCUCCAC_U ........(((((...((.....))((((((........................((((((((....))))))))((.....)).....))))))..))))).... (-26.11 = -26.93 + 0.81)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:13:31 2006