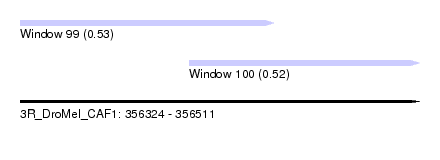
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 356,324 – 356,511 |
| Length | 187 |
| Max. P | 0.534025 |

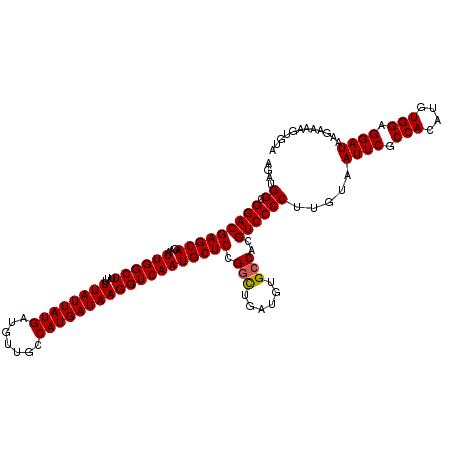
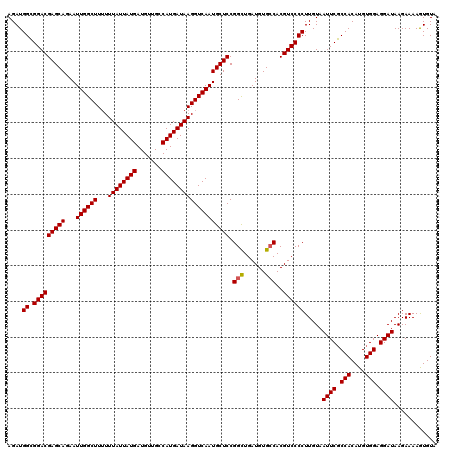
| Location | 356,324 – 356,443 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.67 |
| Mean single sequence MFE | -36.63 |
| Consensus MFE | -33.39 |
| Energy contribution | -33.50 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.00 |
| SVM RNA-class probability | 0.534025 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 356324 119 + 27905053 AGAUGGAGGACGAGCAGAAUUGGCUU-UUUAUUAUGAUGUUGCCAUGAUAAGGUCAAUGCUCCGGUUGAUGUGCCACGUCCCCUUGUAAUUCUCCACAUGUGGAGGAUAAGAAAAAUGUA ....((.(((((((((...((((((.-.((((((((.......))))))))))))))))))).(((......)))..)))))).....((((((((....))))))))............ ( -39.30) >DroSec_CAF1 32339 120 + 1 AGAUGGCGGACGAGCAGAAUUGGCUUUUUUAUUAUGAUGUUGCCAUGAUAAGGUCAAUGCUCCGGCUGAUGUGCCACGUCCCCUUGUAAUUCGCCACAUGUGGAGGAUAAGAAAAGUGUA ....((.(((((((((...((((((...((((((((.......))))))))))))))))))).(((......)))..)))))).....((((.(((....))).))))............ ( -36.60) >DroSim_CAF1 30763 120 + 1 AGAUGGCGGACGAGCAGAAUUGGCUUUUUUAUUAUGAUGUUGCCAUGAUAAGGUCAAUGCUCCGGCUGAUGUGGCACGUCCCCUUGUAAUUCGCCACAUGUGGAGGAUAAGAAAAGUGUA ....((.(((((.((...((..(((...((((((((.......))))))))((........)))))..))...)).))))))).....((((.(((....))).))))............ ( -34.00) >consensus AGAUGGCGGACGAGCAGAAUUGGCUUUUUUAUUAUGAUGUUGCCAUGAUAAGGUCAAUGCUCCGGCUGAUGUGCCACGUCCCCUUGUAAUUCGCCACAUGUGGAGGAUAAGAAAAGUGUA ....((.(((((((((...((((((...((((((((.......))))))))))))))))))).(((......)))..)))))).....((((.(((....))).))))............ (-33.39 = -33.50 + 0.11)

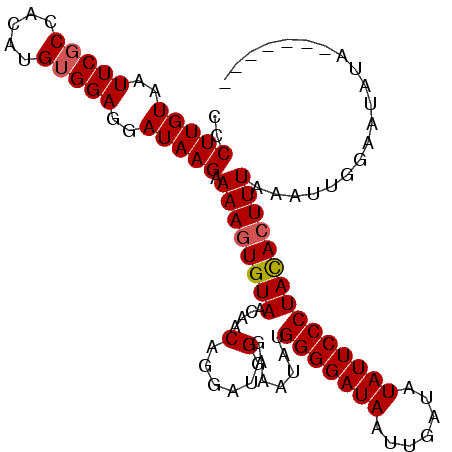
| Location | 356,403 – 356,511 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 89.27 |
| Mean single sequence MFE | -23.70 |
| Consensus MFE | -17.62 |
| Energy contribution | -18.07 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.74 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.523558 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 356403 108 + 27905053 CCCUUGUAAUUCUCCACAUGUGGAGGAUAAGAAAAAUGUAACAACAAAAACGGAAAUAUGGGGAUAAUUGAUAUAUUCCCUAUACUUUAAAGUGACAUAUAUUAGGUU .(((.(((((((((((....)))))))).......((((.((..(......)(((.((((((((............)))))))).)))...)).)))).))).))).. ( -25.10) >DroSec_CAF1 32419 101 + 1 CCCUUGUAAUUCGCCACAUGUGGAGGAUAAGAAAAGUGUAACAACAGGAUGGGAAAUAUGGGGAUAAUUGAUAUAUUCCCUACACUUUAAAUUGGAAUAUA------- (((((((..(((((.....)))))..))))).((((((((.((......))........(((((((.......))))))))))))))).....))......------- ( -23.00) >DroSim_CAF1 30843 101 + 1 CCCUUGUAAUUCGCCACAUGUGGAGGAUAAGAAAAGUGUAACAACAGGAUGGGAAAUAUGGGGAUAAUUGAUAUAUUCCCUACACUUUAAAUUGGAAUAUA------- (((((((..(((((.....)))))..))))).((((((((.((......))........(((((((.......))))))))))))))).....))......------- ( -23.00) >consensus CCCUUGUAAUUCGCCACAUGUGGAGGAUAAGAAAAGUGUAACAACAGGAUGGGAAAUAUGGGGAUAAUUGAUAUAUUCCCUACACUUUAAAUUGGAAUAUA_______ ..(((((..(((((.....)))))..))))).((((((((....(......).......(((((((.......))))))))))))))).................... (-17.62 = -18.07 + 0.45)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:31:44 2006