
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 15,949,027 – 15,949,122 |
| Length | 95 |
| Max. P | 0.902180 |

| Location | 15,949,027 – 15,949,122 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 87.73 |
| Mean single sequence MFE | -28.66 |
| Consensus MFE | -22.68 |
| Energy contribution | -23.40 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.02 |
| SVM RNA-class probability | 0.902180 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
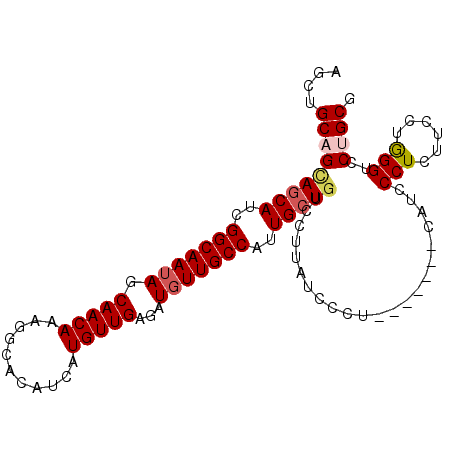
>3R_DroMel_CAF1 15949027 95 + 27905053 AGCUGCAGCAGCAUCGGCAAUAGCAACAAAGGCACAUCAUGUUGAGAUGUUGCCAUUGCUGUCCUUAUCCCU-------CAUCCCUUUUCCUGGGUCCUGCG ....(((((((((..(((((((.(((((...........)))))...)))))))..)))))...........-------...(((.......)))..)))). ( -29.90) >DroSec_CAF1 75386 95 + 1 AGCAGCAGAAGCAUUGGCAAUAGCAACAAAGGCACAUCAUGUUGAGAUGUUGCCAUUGCUGUCCUUAUCCCU-------CAUCCCUCUUCCUGGGUCCAGCG ....((...((((.((((((((.(((((...........)))))...)))))))).))))............-------...(((.......)))....)). ( -26.90) >DroSim_CAF1 50730 95 + 1 AGCUGCAGCAGCAUUGGCAAUAGCAACAAAGGCACAUCAUGUUGAGAUGUUGCCAUUGCUGUCCUUAUCCCU-------CAUCCCUCUUCCUGGGUCCCGCG ....((.((((((.((((((((.(((((...........)))))...)))))))).))))))..........-------...(((.......)))....)). ( -30.80) >DroEre_CAF1 50367 102 + 1 AGCUGCAGUAGCAUCGGCAAUAGCAACAAAGGCACAUCAUGUUGAGAUGUUGCCAUUGCUGUCCUCAUCCCUUAUUCCCCAUCCCUCUUCCUGGGAGCUGCG ....(((((((((..(((((((.(((((...........)))))...)))))))..)))).....................((((.......))))))))). ( -32.00) >DroYak_CAF1 79130 88 + 1 AGCUGCAGCAUCAUCGGCAAAAGCAACAAAGGCACAUCAUGUUGAGAUGUUGCCAUUGCUGUCCUCAUC--------------CCUCUUUCAAGGACCUGCG ....((((((((.((((((...((.......))......))))))))))))))....((.(((((....--------------.........)))))..)). ( -23.72) >consensus AGCUGCAGCAGCAUCGGCAAUAGCAACAAAGGCACAUCAUGUUGAGAUGUUGCCAUUGCUGUCCUUAUCCCU_______CAUCCCUCUUCCUGGGUCCUGCG ....(((((((((..(((((((.(((((...........)))))...)))))))..)))))......................(((......)))..)))). (-22.68 = -23.40 + 0.72)


| Location | 15,949,027 – 15,949,122 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 87.73 |
| Mean single sequence MFE | -31.36 |
| Consensus MFE | -26.88 |
| Energy contribution | -27.20 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.790282 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15949027 95 - 27905053 CGCAGGACCCAGGAAAAGGGAUG-------AGGGAUAAGGACAGCAAUGGCAACAUCUCAACAUGAUGUGCCUUUGUUGCUAUUGCCGAUGCUGCUGCAGCU .((((..(((.......)))...-------.((.(...((.((((((.((((.((((.......)))))))).))))))))..).)).......)))).... ( -31.20) >DroSec_CAF1 75386 95 - 1 CGCUGGACCCAGGAAGAGGGAUG-------AGGGAUAAGGACAGCAAUGGCAACAUCUCAACAUGAUGUGCCUUUGUUGCUAUUGCCAAUGCUUCUGCUGCU ..(((....)))..(((((.((.-------..((....((.((((((.((((.((((.......)))))))).))))))))....)).)).)))))...... ( -30.70) >DroSim_CAF1 50730 95 - 1 CGCGGGACCCAGGAAGAGGGAUG-------AGGGAUAAGGACAGCAAUGGCAACAUCUCAACAUGAUGUGCCUUUGUUGCUAUUGCCAAUGCUGCUGCAGCU .((((..(((.......)))...-------.((.(...((.((((((.((((.((((.......)))))))).))))))))..).)).......)))).... ( -30.50) >DroEre_CAF1 50367 102 - 1 CGCAGCUCCCAGGAAGAGGGAUGGGGAAUAAGGGAUGAGGACAGCAAUGGCAACAUCUCAACAUGAUGUGCCUUUGUUGCUAUUGCCGAUGCUACUGCAGCU .((((.(((((..........))))).....((.(...((.((((((.((((.((((.......)))))))).))))))))..).)).......)))).... ( -32.00) >DroYak_CAF1 79130 88 - 1 CGCAGGUCCUUGAAAGAGG--------------GAUGAGGACAGCAAUGGCAACAUCUCAACAUGAUGUGCCUUUGUUGCUUUUGCCGAUGAUGCUGCAGCU .((((((((((....)))(--------------(..((((.((((((.((((.((((.......)))))))).))))))))))..))...))).)))).... ( -32.40) >consensus CGCAGGACCCAGGAAGAGGGAUG_______AGGGAUAAGGACAGCAAUGGCAACAUCUCAACAUGAUGUGCCUUUGUUGCUAUUGCCGAUGCUGCUGCAGCU .((((..(((.......)))............((....((.((((((.((((.((((.......)))))))).))))))))....)).......)))).... (-26.88 = -27.20 + 0.32)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:11:57 2006