
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 15,934,065 – 15,934,195 |
| Length | 130 |
| Max. P | 0.998617 |

| Location | 15,934,065 – 15,934,180 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.29 |
| Mean single sequence MFE | -26.34 |
| Consensus MFE | -21.53 |
| Energy contribution | -21.57 |
| Covariance contribution | 0.03 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.654170 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15934065 115 - 27905053 CUCGAAAGCAGGACAGCCACUGGCGAAUGGGGAUCAAAUGAUUUAUUCAUAAAUUCCGCAUACGCUAAGGGCAC-----AAAAAAUGCUUGUAAGCCGCAAAUAAUUAUUUACAACAAGU .......((.((.(.(((..(((((.(((.(((....((((.....))))....))).))).)))))..)))((-----((.......))))..)))))..................... ( -28.60) >DroSec_CAF1 63705 115 - 1 CUCGAAAGCAGGACAGCGACUGGCGAAUGGGGAUCAAAUGAUUUAUUCAUAAAUUCCGCAUACGCUAAGGGCAC-----AAAAAAUGCUUGUAAGCCGCAAAUAAUUAUUUACAACAAGU (((....).))....((...(((((.(((.(((....((((.....))))....))).))).)))))..(((((-----((.......))))..)))))..................... ( -26.10) >DroSim_CAF1 41220 115 - 1 CUCGAAAGCAGGACAGCCACUGGCGAAUGGGGAUCAAAUGAUUUAUUCAUAAAUUCCGCAUACGCUAAGGGCAC-----AAAAAAUGCUUGUAAGCCGCAAAUAAUUAUUUACAACAAGU .......((.((.(.(((..(((((.(((.(((....((((.....))))....))).))).)))))..)))((-----((.......))))..)))))..................... ( -28.60) >DroEre_CAF1 40610 115 - 1 CUCGAAAGCAGUACAGCCACUGGCGAAUGCGGAUCAAAUGAUUUAUUCAUAAAUUCAGCAUACGCUAAGGGCAC-----AAAAAAUGCUUGUAAGCCGCAAAUAAUUAUUUACAACAAGU .......((......(((..(((((.((((.((....((((.....))))....)).)))).)))))..)))..-----.......(((....))).))..................... ( -25.90) >DroYak_CAF1 58562 115 - 1 CUCGAAAGCAGAACAGCCACUGGCGAAUGCGGAUCAAAUGAUUUAUUCAUAAAUUCCGCAUACGCUAAGGGCAC-----AAAAAAUGCUUGUAAGCCGCCAAUAAUUAUUUACAACAAGU (((....).))....(((..(((((.(((((((....((((.....))))....))))))).)))))..)))..-----.......((((((......................)))))) ( -32.35) >DroAna_CAF1 34909 116 - 1 CCCGAAAG----GCAGUCACAGGUGAAUCCGGAUCAAAUGAUUUAUUCAUAAAUUCUGCAUACACUAAGAGCACGCAAAAAAAAAUACUUGUAAGCCGCAAAUAAUUAUUCACAAUAAGU ...(((..----((((......((((((..(((((....))))))))))).....)))).........(.((..((((..........))))..)))...........)))......... ( -16.50) >consensus CUCGAAAGCAGGACAGCCACUGGCGAAUGCGGAUCAAAUGAUUUAUUCAUAAAUUCCGCAUACGCUAAGGGCAC_____AAAAAAUGCUUGUAAGCCGCAAAUAAUUAUUUACAACAAGU ..(....).......(((..(((((.(((.(((....((((.....))))....))).))).)))))..)))..............((((((......................)))))) (-21.53 = -21.57 + 0.03)



| Location | 15,934,105 – 15,934,195 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 90 |
| Reading direction | reverse |
| Mean pairwise identity | 97.67 |
| Mean single sequence MFE | -27.16 |
| Consensus MFE | -24.20 |
| Energy contribution | -24.60 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.70 |
| Structure conservation index | 0.89 |
| SVM decision value | 3.16 |
| SVM RNA-class probability | 0.998617 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15934105 90 - 27905053 CGAAGAGGUCAUCCACUCGAAAGCAGGACAGCCACUGGCGAAUGGGGAUCAAAUGAUUUAUUCAUAAAUUCCGCAUACGCUAAGGGCACA ...........(((...(....)..)))..(((..(((((.(((.(((....((((.....))))....))).))).)))))..)))... ( -27.50) >DroSec_CAF1 63745 90 - 1 CGAAGAGGUCAUCCACUCGAAAGCAGGACAGCGACUGGCGAAUGGGGAUCAAAUGAUUUAUUCAUAAAUUCCGCAUACGCUAAGGGCACA .......(((.(((...(....)..))).......(((((.(((.(((....((((.....))))....))).))).)))))..)))... ( -23.00) >DroSim_CAF1 41260 90 - 1 CGAAGAGGUCAUCCACUCGAAAGCAGGACAGCCACUGGCGAAUGGGGAUCAAAUGAUUUAUUCAUAAAUUCCGCAUACGCUAAGGGCACA ...........(((...(....)..)))..(((..(((((.(((.(((....((((.....))))....))).))).)))))..)))... ( -27.50) >DroEre_CAF1 40650 90 - 1 CGAAGAGGUCAUCCACUCGAAAGCAGUACAGCCACUGGCGAAUGCGGAUCAAAUGAUUUAUUCAUAAAUUCAGCAUACGCUAAGGGCACA ......((....))((((....).)))...(((..(((((.((((.((....((((.....))))....)).)))).)))))..)))... ( -26.30) >DroYak_CAF1 58602 90 - 1 CGAAGAGGUCAUCCACUCGAAAGCAGAACAGCCACUGGCGAAUGCGGAUCAAAUGAUUUAUUCAUAAAUUCCGCAUACGCUAAGGGCACA ....(((........)))............(((..(((((.(((((((....((((.....))))....))))))).)))))..)))... ( -31.50) >consensus CGAAGAGGUCAUCCACUCGAAAGCAGGACAGCCACUGGCGAAUGGGGAUCAAAUGAUUUAUUCAUAAAUUCCGCAUACGCUAAGGGCACA ....(((........)))............(((..(((((.(((.(((....((((.....))))....))).))).)))))..)))... (-24.20 = -24.60 + 0.40)

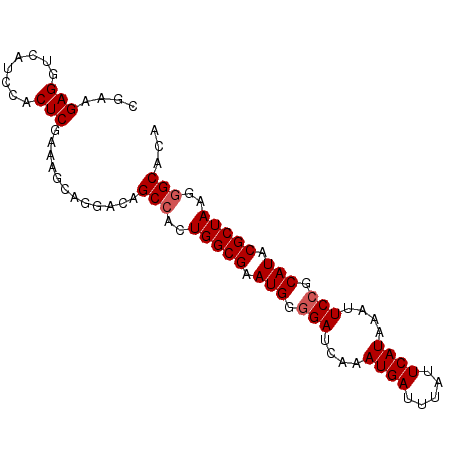

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:11:51 2006