| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
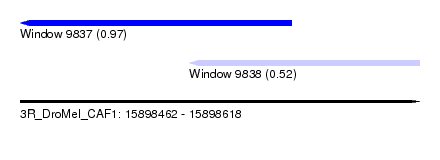
| Location | 15,898,462 – 15,898,618 |
| Length | 156 |
| Max. P | 0.968673 |

| Location | 15,898,462 – 15,898,568 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 86.37 |
| Mean single sequence MFE | -35.30 |
| Consensus MFE | -26.82 |
| Energy contribution | -30.42 |
| Covariance contribution | 3.60 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.54 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.63 |
| SVM RNA-class probability | 0.968673 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15898462 106 - 27905053 GCCGCCAUACAAGAAGAUGCGGCUUCCGUUUCAGACACACGCCUGUCAAUGAAGCGACAACCACUGGGGGCACCAAAAGUGGCAGGAAGUCGCGUUUUUCAAACCG ............(((((((((((((((((((((((((......))))..)))))).....(((((.((....))...)))))..)))))))))))))))....... ( -40.80) >DroSec_CAF1 5715 103 - 1 GCCGCCAUA---GAAGACGCGGCUUCCGUUUCAGACACACGCCUGUCAAUGAAGCGACAACUACUGGGGGCACCAAAAGUGGCAGGAAGUCGCUUUUUUCUAACCU .......((---(((((.(((((((((((((((((((......))))..)))))).....(((((.((....))...)))))..)))))))))..))))))).... ( -37.90) >DroSim_CAF1 5731 103 - 1 GCCGCCAUA---GAAGACGCGGCUUCCGUUUCAGACACACGCCUGUCAAUGAAGCGACAACCACUGGGGGCACCAAAAGUGGCAGGAAGUCGCUUUUUUCUAACCU .......((---(((((.(((((((((((((((((((......))))..)))))).....(((((.((....))...)))))..)))))))))..))))))).... ( -40.20) >DroEre_CAF1 5451 84 - 1 GCCGCCACA---CAAGACGCAGCUUCCGUUUCAGACACACGCCUGUCAAUGAAGCGACAACCACUAGGGGCAC------------------GCGUU-UUCUAACCU .........---.(((((((.(((..(((((((((((......))))..)))))))....((....)))))..------------------)))))-))....... ( -20.90) >DroYak_CAF1 6587 103 - 1 GCCGCCAUA---GAAGACGCAGCUUCCGUUUCAGACACACGCCUGUCAAUGAAGCGACAACCACUGGGGGCACAAAAAGUGGGAGGAAGUCGCGUUUUUCUAUUCU ......(((---((((((((.((((((((((((((((......))))..)))))).....(((((..(....)....)))))..)))))).))).))))))))... ( -36.70) >consensus GCCGCCAUA___GAAGACGCGGCUUCCGUUUCAGACACACGCCUGUCAAUGAAGCGACAACCACUGGGGGCACCAAAAGUGGCAGGAAGUCGCGUUUUUCUAACCU ............(((((((((((((((((((((((((......))))..)))))).....(((((.((....))...)))))..)))))))))))))))....... (-26.82 = -30.42 + 3.60)



| Location | 15,898,528 – 15,898,618 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 81.33 |
| Mean single sequence MFE | -25.54 |
| Consensus MFE | -16.72 |
| Energy contribution | -17.58 |
| Covariance contribution | 0.86 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.42 |
| Structure conservation index | 0.65 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.516998 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15898528 90 - 27905053 ACUGG--------------GGUUUCCACAGCCUGCGAAACAAAGGCGACAUAUCAACAAAUGUUGCCGCCAUACAAGAAGAUGCGGCUUCCGUUUCAGACACAC ....(--------------((((.....)))))..(((((...((((((((........))))))))(((..............)))....)))))........ ( -23.84) >DroSec_CAF1 5781 87 - 1 ACUGG--------------GGUUUCCACAGCCUGCGAAACAAAGGAGACAUAUCAACAAAUGUUGCCGCCAUA---GAAGACGCGGCUUCCGUUUCAGACACAC .((((--------------((((((((.....)).)))))...((((((((........)))))(((((....---......))))).)))..)))))...... ( -21.70) >DroSim_CAF1 5797 87 - 1 ACUGG--------------GGUUUCCACAGCCUGCGAAACAAAGGCGACAUAUCAACAAAUGUUGCCGCCAUA---GAAGACGCGGCUUCCGUUUCAGACACAC .((((--------------((((((((.....)).)))))...((((((((........)))))))).)).))---)..((.((((...)))).))........ ( -24.20) >DroEre_CAF1 5498 87 - 1 ACUGG--------------GGCUUCCAUAGCCUGCGAAACAAAGGCGACAUAUCAACAAAUGUUGCCGCCACA---CAAGACGCAGCUUCCGUUUCAGACACAC .(((.--------------.((......(((.((((.......((((((((........))))))))(.....---)....)))))))...))..)))...... ( -25.70) >DroYak_CAF1 6653 97 - 1 ACAGGCGGCA--AUCGU--GGCACUGGUAGCCUGCGAAACAAAGGCGACAUAUCAACAAAUGUUGCCGCCAUA---GAAGACGCAGCUUCCGUUUCAGACACAC ..(((((((.--.((((--(((.......))).))))......((((((((........)))))))))))...---((((......)))).))))......... ( -28.70) >DroPer_CAF1 12566 100 - 1 GCAGGCUGGAGGAUAGUGGGGGGGCUGGAG-UGGGGAAACAAAAGCGACAUAUCAACAAAUGUUGCCGGCAUA---GAAGACGCAGCUUCCGUGGCAGCCACAC ...(((((..(((.(((.(.(...(((..(-(.((....)....(((((((........)))))))).)).))---)....).).))))))....))))).... ( -29.10) >consensus ACUGG______________GGUUUCCACAGCCUGCGAAACAAAGGCGACAUAUCAACAAAUGUUGCCGCCAUA___GAAGACGCAGCUUCCGUUUCAGACACAC ...................(((.......)))...(((((...((((((((........)))))))).........((((......)))).)))))........ (-16.72 = -17.58 + 0.86)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:10:52 2006