
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 15,893,742 – 15,893,890 |
| Length | 148 |
| Max. P | 0.643816 |

| Location | 15,893,742 – 15,893,861 |
|---|---|
| Length | 119 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 78.42 |
| Mean single sequence MFE | -34.83 |
| Consensus MFE | -14.82 |
| Energy contribution | -17.10 |
| Covariance contribution | 2.28 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.43 |
| SVM decision value | -0.05 |
| SVM RNA-class probability | 0.507619 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
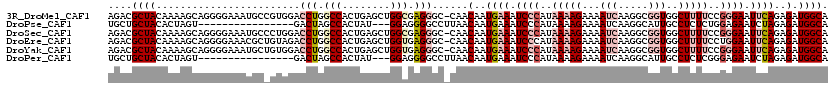
>3R_DroMel_CAF1 15893742 119 + 27905053 AGACGCUACAAAAGCAGGGGAAAUGCCGUGGACCUGGCCACUGAGCUGGCGAGGGC-CAACAAUGAAAUCCCAUAAAAGAAAAUCAAGGCGGUGGCUUUUCCGGGAAUUCAGAGAUGGCA ....(((.....)))........((((((((.(((.((((......)))).))).)-)..(..((((.((((..(((((...(((.....)))..)))))..)))).))))..))))))) ( -38.60) >DroPse_CAF1 7419 101 + 1 UGCUGCUACACUAGU----------------GACUAGCCACUAU---GGAGGGGCCUUAACAAUGAAAUCCCAUAAAAGAAAAUCAAGGCAUUGCCUCUCUGGAGAAUCUAGAGAUGGCA ...((((.((.((((----------------(......))))))---)..((((..(((....)))..))))...............)))).(((((((((((.....))))))).)))) ( -28.70) >DroSec_CAF1 696 119 + 1 AGACGCUACAAAAGCAGGGGAAAUGCCCUGGACCUGGCCACUGAGCUGGCGAGGGC-CAACAAUGAAAUCCCAUAAAAGAAAAUCAAGGCGGUGGCUUUUCCGGGAAUUCAGAGAUGGCA ..............(((((......)))))..(((.((((......)))).)))((-((.(..((((.((((..(((((...(((.....)))..)))))..)))).))))..).)))). ( -42.10) >DroEre_CAF1 704 119 + 1 AGACGCUACAAAAGCAGGGGAAACGCUGUAGACCUGGCCACUGAGCUGGUGAGGGC-CAACAAUGAAAUCCCAUAAAAGAAAAUCAAGGCGGUGGCUUUUCCUGGAAUUCAGAGAUGGCA .............((((.(....).))))...(((.((((......)))).)))((-((.(..((((.(((...(((((...(((.....)))..)))))...))).))))..).)))). ( -37.10) >DroYak_CAF1 664 119 + 1 AGACGCUACAAAAGCAGGGGAAAUGCUGUGGACCUGGCCACUGAGCUGGUGAGGGC-CAACAAUGAAAUCCCAUAAAAGAAAAUCAAGGCGGUGGCUUUUCCGGGAAUUCAGAGAUGGCA .....(((((...(((.......)))))))).(((.((((......)))).)))((-((.(..((((.((((..(((((...(((.....)))..)))))..)))).))))..).)))). ( -38.00) >DroPer_CAF1 7451 101 + 1 UGCUGCUACACUAGU----------------GACUAGCCACUAU---GGAGGGGCCUUAACAAUGAAAUCCCAUAAAAGAAAAUCAAGGCAUUGCCUCUCGGGAGAAUCUAGAGAUGGCA ...((((.((.((((----------------(......))))))---)..((((..(((....)))..))))...............)))).((((((((.((.....)).)))).)))) ( -24.50) >consensus AGACGCUACAAAAGCAGGGGAAAUGC_GUGGACCUGGCCACUGAGCUGGAGAGGGC_CAACAAUGAAAUCCCAUAAAAGAAAAUCAAGGCGGUGGCUUUUCCGGGAAUUCAGAGAUGGCA ....((((........................(((.(((........))).)))......(..((((.((((..(((((...(((.....)))..)))))..)))).))))..).)))). (-14.82 = -17.10 + 2.28)



| Location | 15,893,782 – 15,893,890 |
|---|---|
| Length | 108 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.26 |
| Mean single sequence MFE | -30.56 |
| Consensus MFE | -13.46 |
| Energy contribution | -12.77 |
| Covariance contribution | -0.69 |
| Combinations/Pair | 1.30 |
| Mean z-score | -2.40 |
| Structure conservation index | 0.44 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.643816 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15893782 108 - 27905053 -----------UUUGAGCGCCGGCACUCUGUAUAUCACAUUGCCAUCUCUGAAUUCCCGGAAAAGCCACCGCCUUGAUUUUCUUUUAUGGGAUUUCAUUGUUG-GCCCUCGCCAGCUCAG -----------..(((((...(((....(((.....)))..((((.(..((((.((((((((((..((......)).)))))).....)))).))))..).))-))....))).))))). ( -31.30) >DroPse_CAF1 7443 113 - 1 UCACACACCAGUUUGU----AUGGAGUCCGUAUAUCACAUUGCCAUCUCUAGAUUCUCCAGAGAGGCAAUGCCUUGAUUUUCUUUUAUGGGAUUUCAUUGUUAAGGCCCCUCC---AUAG ...............(----((((((............((((((.(((((.(.....).)))))))))))((((((((..((((....)))).......))))))))..))))---))). ( -32.60) >DroSim_CAF1 730 108 - 1 -----------UUUGAGCGCCGGCACUCAGUAUAUCACAUUGCCAUCUCUGAAUUCCCGGAAAAGCCACCGCCUUGAUUUUCUUUUAUGGGAUUUCAUUGUUG-GCCCUCGCCAGCUCAG -----------..(((((...(((.....((.....))...((((.(..((((.((((((((((..((......)).)))))).....)))).))))..).))-))....))).))))). ( -30.90) >DroEre_CAF1 744 108 - 1 -----------AUUGAGCGCCGGCACUCAGUAUAUCACAUUGCCAUCUCUGAAUUCCAGGAAAAGCCACCGCCUUGAUUUUCUUUUAUGGGAUUUCAUUGUUG-GCCCUCACCAGCUCAG -----------..(((((...((((...............)))).((.(((.....)))))...((((..((..(((..((((.....))))..)))..))))-))........))))). ( -25.56) >DroYak_CAF1 704 108 - 1 -----------UUUGAGCGCCGACACUCAGUAUAUCACAUUGCCAUCUCUGAAUUCCCGGAAAAGCCACCGCCUUGAUUUUCUUUUAUGGGAUUUCAUUGUUG-GCCCUCACCAGCUCAG -----------..((((.(((((((....(((........)))......((((.((((((((((..((......)).)))))).....)))).)))).)))))-)).))))......... ( -27.20) >DroPer_CAF1 7475 113 - 1 UCAGACACCAGUCUGU----AUGGAGUCCGUAUAUCACAUUGCCAUCUCUAGAUUCUCCCGAGAGGCAAUGCCUUGAUUUUCUUUUAUGGGAUUUCAUUGUUAAGGCCCCUCC---AUAG .(((((....)))))(----((((((............((((((.((((..(.....)..))))))))))((((((((..((((....)))).......))))))))..))))---))). ( -35.80) >consensus ___________UUUGAGCGCCGGCACUCAGUAUAUCACAUUGCCAUCUCUGAAUUCCCGGAAAAGCCACCGCCUUGAUUUUCUUUUAUGGGAUUUCAUUGUUG_GCCCUCACCAGCUCAG ............((((((...........(((........)))......((((.(((((.(((((................))))).))))).)))).................)))))) (-13.46 = -12.77 + -0.69)

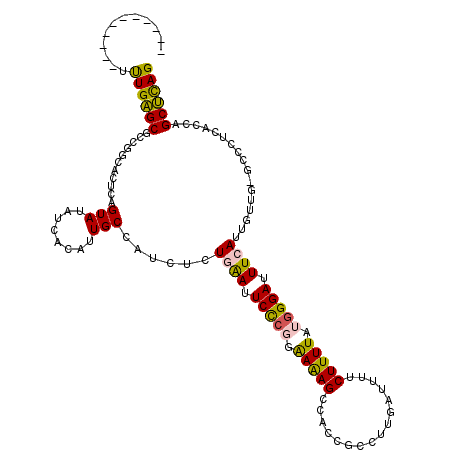

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:10:47 2006