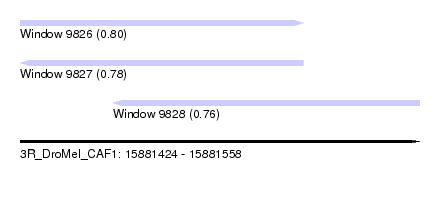
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 15,881,424 – 15,881,558 |
| Length | 134 |
| Max. P | 0.802201 |

| Location | 15,881,424 – 15,881,519 |
|---|---|
| Length | 95 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 80.95 |
| Mean single sequence MFE | -30.43 |
| Consensus MFE | -23.01 |
| Energy contribution | -23.77 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.802201 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15881424 95 + 27905053 AUGGGCCAUAUGGCUAAGGGCUUACCAUGGG------------GC-UUUGUUGUGUGCCAACUGCCAUUAAAUGGCUAAUUAAGAU----UUCAGUCCGCUGGCAGUCGUCC ...(((((((((((...((.....)).(((.------------((-........)).)))...)))))...))))))......(((----(((((....)))).)))).... ( -28.90) >DroPse_CAF1 9224 112 + 1 AUUGGCCAUAUGGCUGAGCGCUUACCUCGGUAGCCAGCUACCAGCUAUUGUUGUCUGCCAACUGCCAUUAAAUGCCUAAUUAAGAUCUAUCUCACUUUACUGGCAGUUGUCC .(..(((....)))..)..((..((..(((((((.........)))))))..))..))(((((((((.((((((........(((....))))).)))).)))))))))... ( -31.20) >DroPer_CAF1 9221 112 + 1 AUUGGCCAUAUGGCUGAGCGCUUACCUCGGUAGCCAGCUACCAGCUAUUGUUGUCUGCCAACUGCCAUUAAAUGCCUAAUUAAGAUCUAUCUCACUUUACUGGCAGUUGUCC .(..(((....)))..)..((..((..(((((((.........)))))))..))..))(((((((((.((((((........(((....))))).)))).)))))))))... ( -31.20) >consensus AUUGGCCAUAUGGCUGAGCGCUUACCUCGGUAGCCAGCUACCAGCUAUUGUUGUCUGCCAACUGCCAUUAAAUGCCUAAUUAAGAUCUAUCUCACUUUACUGGCAGUUGUCC .((((((....))))))..((..((..(((((((.........)))))))..))..))(((((((((((((........))))((......)).......)))))))))... (-23.01 = -23.77 + 0.75)

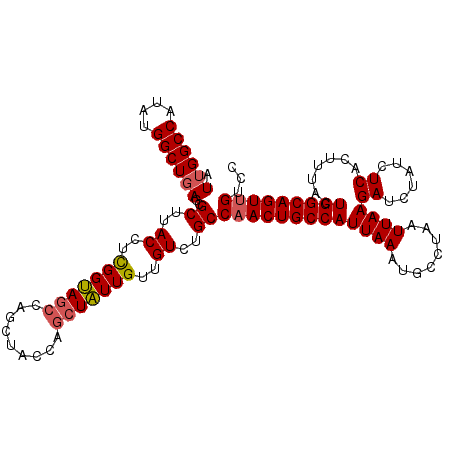

| Location | 15,881,424 – 15,881,519 |
|---|---|
| Length | 95 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 80.95 |
| Mean single sequence MFE | -31.10 |
| Consensus MFE | -23.81 |
| Energy contribution | -25.04 |
| Covariance contribution | 1.23 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.781240 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
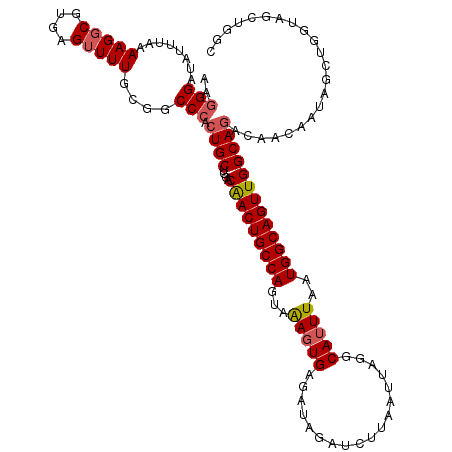
>3R_DroMel_CAF1 15881424 95 - 27905053 GGACGACUGCCAGCGGACUGAA----AUCUUAAUUAGCCAUUUAAUGGCAGUUGGCACACAACAAA-GC------------CCCAUGGUAAGCCCUUAGCCAUAUGGCCCAU ((.(((((((((..((.((((.----.......))))))......)))))))))............-((------------(..(((((.........)))))..))))).. ( -28.30) >DroPse_CAF1 9224 112 - 1 GGACAACUGCCAGUAAAGUGAGAUAGAUCUUAAUUAGGCAUUUAAUGGCAGUUGGCAGACAACAAUAGCUGGUAGCUGGCUACCGAGGUAAGCGCUCAGCCAUAUGGCCAAU ((.(((((((((.((((.(((((....)))))........)))).)))))))((((..............(((((....)))))((((....).))).))))..)).))... ( -32.50) >DroPer_CAF1 9221 112 - 1 GGACAACUGCCAGUAAAGUGAGAUAGAUCUUAAUUAGGCAUUUAAUGGCAGUUGGCAGACAACAAUAGCUGGUAGCUGGCUACCGAGGUAAGCGCUCAGCCAUAUGGCCAAU ((.(((((((((.((((.(((((....)))))........)))).)))))))((((..............(((((....)))))((((....).))).))))..)).))... ( -32.50) >consensus GGACAACUGCCAGUAAAGUGAGAUAGAUCUUAAUUAGGCAUUUAAUGGCAGUUGGCAGACAACAAUAGCUGGUAGCUGGCUACCGAGGUAAGCGCUCAGCCAUAUGGCCAAU ...(((((((((...(((((..................)))))..)))))))))((..((.....(((((((...))))))).....))..)).....(((....))).... (-23.81 = -25.04 + 1.23)



| Location | 15,881,455 – 15,881,558 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.22 |
| Mean single sequence MFE | -34.63 |
| Consensus MFE | -24.01 |
| Energy contribution | -24.90 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.760422 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15881455 103 - 27905053 AAGGGAU-UUGAAAAGGCGUGAGAUUUGCGGCCAGCUGCUGGACGACUGCCAGCGGACUGAA----AUCUUAAUUAGCCAUUUAAUGGCAGUUGGCACACAACAAA-GC----------- ..(((((-((.....(.((..(...)..)).)((((((((((.......))))))).)))))----))))).....(((((...))))).((((.....))))...-..----------- ( -32.10) >DroPse_CAF1 9256 120 - 1 AAGGGAUAUUUAAAAGGCGUGAGUUUUGCGUCCCACUGCUGGACAACUGCCAGUAAAGUGAGAUAGAUCUUAAUUAGGCAUUUAAUGGCAGUUGGCAGACAACAAUAGCUGGUAGCUGGC ..(((((......(((((....)))))..))))).((((....(((((((((.((((.(((((....)))))........)))).))))))))))))).......((((.....)))).. ( -37.10) >DroPer_CAF1 9253 120 - 1 AAGGGAUAUUUAAAAGGCGUGAGUUUUGCAGCCCACUGCUGGACAACUGCCAGUAAAGUGAGAUAGAUCUUAAUUAGGCAUUUAAUGGCAGUUGGCAGACAACAAUAGCUGGUAGCUGGC ..(((........(((((....)))))....))).((((....(((((((((.((((.(((((....)))))........)))).))))))))))))).......((((.....)))).. ( -34.70) >consensus AAGGGAUAUUUAAAAGGCGUGAGUUUUGCGGCCCACUGCUGGACAACUGCCAGUAAAGUGAGAUAGAUCUUAAUUAGGCAUUUAAUGGCAGUUGGCAGACAACAAUAGCUGGUAGCUGGC ..(((........(((((....)))))....))).((((....(((((((((...(((((..................)))))..)))))))))))))...................... (-24.01 = -24.90 + 0.89)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:10:42 2006