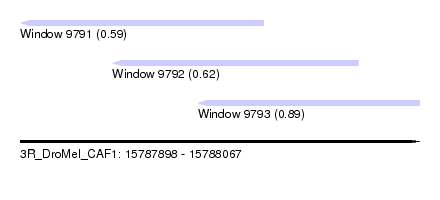
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 15,787,898 – 15,788,067 |
| Length | 169 |
| Max. P | 0.891735 |

| Location | 15,787,898 – 15,788,001 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 94.29 |
| Mean single sequence MFE | -27.26 |
| Consensus MFE | -21.58 |
| Energy contribution | -22.42 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.594341 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15787898 103 - 27905053 CCUGCUGCAAGGAAAAGGUUAAGAGGCAAAGAAACGCUAGGUAGUCCU---GCUAAAUCGGCGGAAAAACAAGCGGCAGGAAGCGAACAGCAUGCAAAUCAGUUCG ((((((((........(..((...(((........)))...))..)((---(((.....)))))........))))))))...(((((.............))))) ( -27.62) >DroSec_CAF1 9161 106 - 1 CCUGCUGCAAGGAAAAGGUUAAUAGGCAAAGAAACGCUAGGUAGUCCUGCUGCUAAAUCGGCGGAAAAACAAGCGGCAGGAAGCGAACAGCAUGCAAAUCAGUUCG ((((((((.......((((((...(((........)))...))).))).(((((.....)))))........))))))))...(((((.............))))) ( -29.62) >DroSim_CAF1 13628 106 - 1 CCUGCUGCAAGGAAAAGGUUAAGAGGCAAAGAAACGCUAGGUAGUCCUGCUGCUAAAUCGGCGGAAAAACAAGCGGCAGGAAGCGAACAGCAUGCAAAUCAGUUCG ((((((((.......((((((...(((........)))...))).))).(((((.....)))))........))))))))...(((((.............))))) ( -29.62) >DroEre_CAF1 12969 103 - 1 CCCGCUGCAAGGAAAAGGUUAAGAGCCAAAGAAAAGCUAGGUAGUCCU---GCUAAAUCGGCGGAAAAACAAGCGGCAGAAAGCGAACGGCAUGCAAAUCAGUUCG .((((((.........(((.....))).......(((.(((....)))---)))....))))))........((........))((((.............)))). ( -23.02) >DroYak_CAF1 30329 103 - 1 CCUGCUGCAAAGAAAAAGUUAAGAGCCAAAGAAACGCCAGGUAGUCUU---GCUAAAUCGGCGGAAAAACAAGCGGCAGGAAGCGAACAGCAUGCAAAUCAGUUCG ((((((((.........((.....))........(((((..(((....---.)))..).)))).........))))))))...(((((.............))))) ( -26.42) >consensus CCUGCUGCAAGGAAAAGGUUAAGAGGCAAAGAAACGCUAGGUAGUCCU___GCUAAAUCGGCGGAAAAACAAGCGGCAGGAAGCGAACAGCAUGCAAAUCAGUUCG ((((((((.........(((....))).......((((...((((......))))....)))).........))))))))...(((((.............))))) (-21.58 = -22.42 + 0.84)



| Location | 15,787,937 – 15,788,041 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 93.59 |
| Mean single sequence MFE | -26.94 |
| Consensus MFE | -23.82 |
| Energy contribution | -23.74 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.21 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.622851 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15787937 104 - 27905053 UGAACUCAAUUUUCGGGCAGCAGAGCAAAUUCGUUAGAGUCCUGCUGCAAGGAAAAGGUUAAGAGGCAAAGAAACGCUAGGUAGUCCU---GCUAAAUCGGCGGAAA .........(((((..(((((((..(....((....)))..)))))))..))))).(..((...(((........)))...))..)((---(((.....)))))... ( -26.60) >DroSec_CAF1 9200 107 - 1 UGAACUCAAUUUUCGGGCAGCAGAGCAAAUUCGUUAGAGUCCUGCUGCAAGGAAAAGGUUAAUAGGCAAAGAAACGCUAGGUAGUCCUGCUGCUAAAUCGGCGGAAA .........(((((..(((((((..(....((....)))..)))))))..)))))((((((...(((........)))...))).))).(((((.....)))))... ( -28.60) >DroSim_CAF1 13667 107 - 1 UGAACUCAAUUUUCGGGCAGCAGAGCAAAUUCGUUAGAGUCCUGCUGCAAGGAAAAGGUUAAGAGGCAAAGAAACGCUAGGUAGUCCUGCUGCUAAAUCGGCGGAAA .........(((((..(((((((..(....((....)))..)))))))..)))))((((((...(((........)))...))).))).(((((.....)))))... ( -28.60) >DroEre_CAF1 13008 104 - 1 UGAACUCAAUUUUCAGGCAGCAGAGCAAAUUCGUUAGAGUCCCGCUGCAAGGAAAAGGUUAAGAGCCAAAGAAAAGCUAGGUAGUCCU---GCUAAAUCGGCGGAAA ..((((...(((((..(((((.(..(....((....)))..).)))))...)))))))))....(((.......(((.(((....)))---))).....)))..... ( -23.40) >DroYak_CAF1 30368 104 - 1 UGAACUACAUUUUCGGGCAGCAGAGCAAAUUCGUUAGAAUCCUGCUGCAAAGAAAAAGUUAAGAGCCAAAGAAACGCCAGGUAGUCUU---GCUAAAUCGGCGGAAA ..((((...(((((..(((((((.(...((((....))))))))))))...)))))))))..............(((((..(((....---.)))..).)))).... ( -27.50) >consensus UGAACUCAAUUUUCGGGCAGCAGAGCAAAUUCGUUAGAGUCCUGCUGCAAGGAAAAGGUUAAGAGGCAAAGAAACGCUAGGUAGUCCU___GCUAAAUCGGCGGAAA ..((((...(((((..(((((((.(...((((....))))))))))))...)))))))))..............((((...((((......))))....)))).... (-23.82 = -23.74 + -0.08)

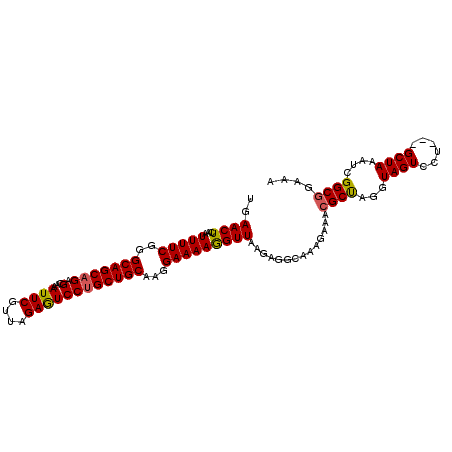

| Location | 15,787,973 – 15,788,067 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 93.62 |
| Mean single sequence MFE | -22.72 |
| Consensus MFE | -19.46 |
| Energy contribution | -19.38 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.891735 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15787973 94 - 27905053 CAGAUACUAAAACGAUGGAAACGAGUUGAACUCAAUUUUCGGGCAGCAGAGCAAAUUCGUUAGAGUCCUGCUGCAAGGAAAAGGUUAAGAGGCA ......((..(((...(....)(((.....)))..(((((..(((((((..(....((....)))..)))))))..)))))..)))...))... ( -23.40) >DroSec_CAF1 9239 94 - 1 CAGAUACUAAAACGAUGGAAACGAGUUGAACUCAAUUUUCGGGCAGCAGAGCAAAUUCGUUAGAGUCCUGCUGCAAGGAAAAGGUUAAUAGGCA ......(((.(((...(....)(((.....)))..(((((..(((((((..(....((....)))..)))))))..)))))..)))..)))... ( -23.40) >DroSim_CAF1 13706 94 - 1 CAGAUACUAAAACGAUGGAAACGAGUUGAACUCAAUUUUCGGGCAGCAGAGCAAAUUCGUUAGAGUCCUGCUGCAAGGAAAAGGUUAAGAGGCA ......((..(((...(....)(((.....)))..(((((..(((((((..(....((....)))..)))))))..)))))..)))...))... ( -23.40) >DroEre_CAF1 13044 94 - 1 CGGAUACCAAAAGGAUGGAAACGAGCUGAACUCAAUUUUCAGGCAGCAGAGCAAAUUCGUUAGAGUCCCGCUGCAAGGAAAAGGUUAAGAGCCA .((..(((........(....)(((.....)))..(((((..(((((.(..(....((....)))..).)))))...))))))))......)). ( -21.10) >DroYak_CAF1 30404 94 - 1 CAGUUACUAAAAGGAUGGAAACGAGUUGAACUACAUUUUCGGGCAGCAGAGCAAAUUCGUUAGAAUCCUGCUGCAAAGAAAAAGUUAAGAGCCA ............((.((....)).....((((...(((((..(((((((.(...((((....))))))))))))...))))))))).....)). ( -22.30) >consensus CAGAUACUAAAACGAUGGAAACGAGUUGAACUCAAUUUUCGGGCAGCAGAGCAAAUUCGUUAGAGUCCUGCUGCAAGGAAAAGGUUAAGAGGCA .....(((........(....)(((.....)))..(((((..(((((((.(...((((....))))))))))))...))))))))......... (-19.46 = -19.38 + -0.08)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:10:08 2006