
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 15,739,447 – 15,739,538 |
| Length | 91 |
| Max. P | 0.805717 |

| Location | 15,739,447 – 15,739,538 |
|---|---|
| Length | 91 |
| Sequences | 3 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 84.01 |
| Mean single sequence MFE | -18.23 |
| Consensus MFE | -12.17 |
| Energy contribution | -11.73 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.707635 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
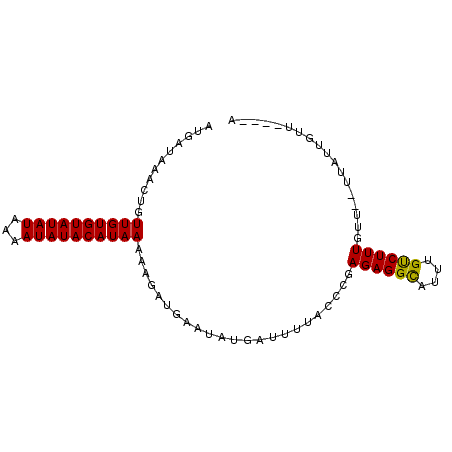
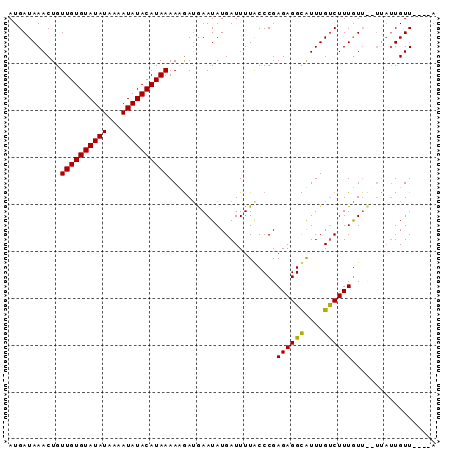
>3R_DroMel_CAF1 15739447 91 + 27905053 AUGAUAAACUGUUGUGUAUAUAAAAUAUACAUAAAAAGAUAAAUAUGAUUUUGCCCGAGAGGCAUUUGUCUUUGUAUUUUAUUGUUUGUGA ...((((((..((((((((((...)))))))))).((((((.....(((..((((.....))))...)))....))))))...)))))).. ( -19.90) >DroSim_CAF1 8089 87 + 1 AUGAUAAACUGUUGUGUAUAUAAAAUAUACAUAAAAUGAAGAAUAUGAUUUUACCCGAGAGGCAUUUGUCUUUAUUCGUUAUUGUU----A ......(((..((((((((((...))))))))))(((((.(((((.(((.....((....)).....)))..))))).))))))))----. ( -17.50) >DroEre_CAF1 7974 81 + 1 AUGAUAAACUGUUGUGUAUAUAAAAUAUACAUAAAACAAUGAAUAUGAUUUUACCCGAGAGGUGUUUGCCUUUGUU------UGUU----A .((((((((..((((((((((...)))))))))).......................((((((....)))))))))------))))----) ( -17.30) >consensus AUGAUAAACUGUUGUGUAUAUAAAAUAUACAUAAAAAGAUGAAUAUGAUUUUACCCGAGAGGCAUUUGUCUUUGUU__UUAUUGUU____A ...........((((((((((...)))))))))).......................((((((....)))))).................. (-12.17 = -11.73 + -0.44)

| Location | 15,739,447 – 15,739,538 |
|---|---|
| Length | 91 |
| Sequences | 3 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 84.01 |
| Mean single sequence MFE | -10.13 |
| Consensus MFE | -9.28 |
| Energy contribution | -8.40 |
| Covariance contribution | -0.88 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.805717 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
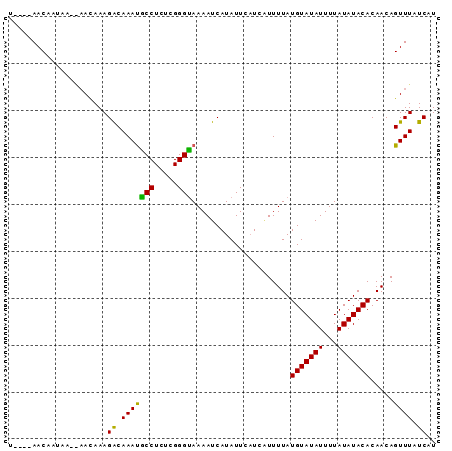
>3R_DroMel_CAF1 15739447 91 - 27905053 UCACAAACAAUAAAAUACAAAGACAAAUGCCUCUCGGGCAAAAUCAUAUUUAUCUUUUUAUGUAUAUUUUAUAUACACAACAGUUUAUCAU ....((((......(((.(((((.(((((((.....))))........))).))))).)))((((((...))))))......))))..... ( -11.20) >DroSim_CAF1 8089 87 - 1 U----AACAAUAACGAAUAAAGACAAAUGCCUCUCGGGUAAAAUCAUAUUCUUCAUUUUAUGUAUAUUUUAUAUACACAACAGUUUAUCAU .----......((((((((..((....((((.....))))...)).))))).........(((((((...))))))).....)))...... ( -8.70) >DroEre_CAF1 7974 81 - 1 U----AACA------AACAAAGGCAAACACCUCUCGGGUAAAAUCAUAUUCAUUGUUUUAUGUAUAUUUUAUAUACACAACAGUUUAUCAU .----....------......(..(((((((.....)))..............((((...(((((((...))))))).))))))))..).. ( -10.50) >consensus U____AACAAUAA__AACAAAGACAAAUGCCUCUCGGGUAAAAUCAUAUUCAUCAUUUUAUGUAUAUUUUAUAUACACAACAGUUUAUCAU .....................((.(((((((.....))).....................(((((((...))))))).....)))).)).. ( -9.28 = -8.40 + -0.88)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:09:05 2006