
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 15,716,113 – 15,716,334 |
| Length | 221 |
| Max. P | 0.933380 |

| Location | 15,716,113 – 15,716,228 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 64.68 |
| Mean single sequence MFE | -23.31 |
| Consensus MFE | -12.51 |
| Energy contribution | -13.32 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.54 |
| SVM decision value | 1.23 |
| SVM RNA-class probability | 0.933380 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
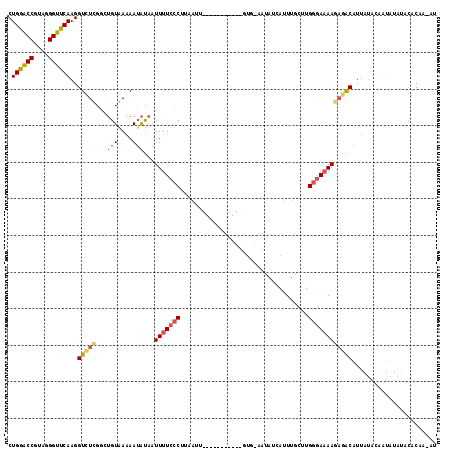
>3R_DroMel_CAF1 15716113 115 - 27905053 CUGGACCGCAGGGUUCAAGGUUUUGGGUGUAAAAAUACAAUUUUCCCUUAAUUUAACGUUAUAUGUGGAAUAUAAUUUGCUUGGGA-AAAAGGCAUUAUAUAAUAUAUACACCA-AU .((((((....))))))........((((((.........(((((((..........(((((((.....)))))))......))))-)))......((((...)))))))))).-.. ( -23.59) >DroSec_CAF1 12348 95 - 1 CUGGACCGUAGGGUUCAAGGUCUCGGCUGUGAAAAUAUAAUUUUCCCUUCAUU-----------AUGUAUCAUCAUUUGCUUGGGAAAAGAGACAUU----------UACACAA-AU .((((((....))))))..(((((...(((....)))...(((((((......-----------(((......)))......))))))))))))...----------.......-.. ( -24.40) >DroSim_CAF1 12662 99 - 1 CUGGACCGUAGGGUUCAAGGUCUCGGCUGUGAAAAUAUAAUUUUCCCUUCAUU-----------------UAACAUUUUUCUGGGAAAAGAGGCAUUAUACAAUAUAUACACAA-AU .((((((....))))))..(((((...(((....)))...(((((((......-----------------............))))))))))))....................-.. ( -21.37) >DroYak_CAF1 14073 117 - 1 CUGACCCUUCUGGGUCAAGGUCUUGCCAGUAAGAAAGUAAUUUUCACAGACCUUACAAAUGUAUGUAAAAUGUCAUUCAGUUGGCAAAAUUAACAUAAUACAAUAUUUAAACAAAAU .((((((....)))))).(((.(((((((...(((((...)))))...(((.(((((......)))))...)))......))))))).))).......................... ( -23.90) >consensus CUGGACCGUAGGGUUCAAGGUCUCGGCUGUAAAAAUAUAAUUUUCCCUUAAUU___________GUG_AAUAUCAUUUGCUUGGGAAAAGAGACAUUAUACAAUAUAUACACAA_AU .((((((....))))))..(((((...(((....)))...(((((((...................................))))))))))))....................... (-12.51 = -13.32 + 0.81)


| Location | 15,716,228 – 15,716,334 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 79.42 |
| Mean single sequence MFE | -33.76 |
| Consensus MFE | -17.55 |
| Energy contribution | -18.42 |
| Covariance contribution | 0.87 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.660931 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
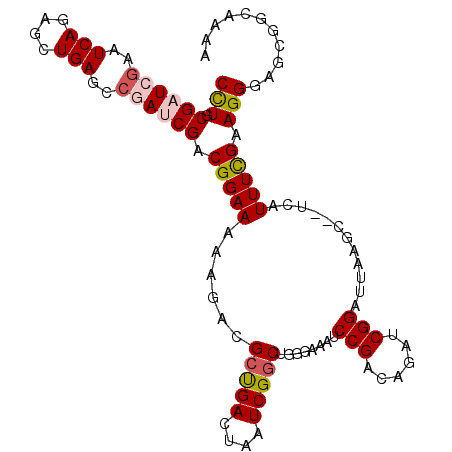
>3R_DroMel_CAF1 15716228 106 - 27905053 CCUGCGAUCGAAUCAGAGCUGAGCCGAUCGACGGAAAAAGACGCUGACUAAUCGGCUGGGAAAUCCGCCAGAUCGGAUUAAGC--UCAUUUCGAAGGGAGCGGCAAUA .((((..(((((...(((((...((((((..((((.......(((((....))))).......))))...))))))....)))--))..))))).....))))..... ( -37.64) >DroSec_CAF1 12443 106 - 1 CCUGCGAUCGAAUCAGAGCUGAGCCGAUCGACGGAAAAAGACGCUGACUAAUCGGCUGGGAAAUCCGACAGAUCGGCUUAAGC--UCAUUUCGAAGGGAGCGGCAAAA .((((..(((((...((((((((((((((..((((.......(((((....))))).......))))...))))))))).)))--))..))))).....))))..... ( -42.84) >DroSim_CAF1 12761 106 - 1 CCUGCGAUCGAAUCAGAGCUGAGCCGAUCGACGGAAAAAGGCGCUGACUAAUCGGCUGGGAAAUCCGACAAAUCGGCUUAAGC--UCAUUUCGAAGGGAGCGGCAAAA .((((..(((((...(((((((((((((...((((.......(((((....))))).......))))....)))))))).)))--))..))))).....))))..... ( -38.24) >DroEre_CAF1 12674 84 - 1 CCUGCGAUCGAAUCAGAGGUGAGCCAAGCGACGGAAA-AAUCGCCGACUGAUCGGCGGGG----------------------CACUCAUUUCGAAGGGG-CGGCAACG (((((...(((.((((.(((((.((.......))...-..)))))..))))))))))))(----------------------(.(((........))))-)(....). ( -27.20) >DroYak_CAF1 14190 108 - 1 CUUGCGAUCAAAUCAGAGGUGAGCCAAGCGACGGAAAAAACCGCUGACAAAUCGACUGGGAAGUCCGACAGAUCGGAUUAAGCAAACAUUUUGAAGGGGGCGAUUAUA .((((..((...((((.(((...((.......)).....))).))))((((.....((...(((((((....)))))))...)).....))))..))..))))..... ( -22.90) >consensus CCUGCGAUCGAAUCAGAGCUGAGCCGAUCGACGGAAAAAGACGCUGACUAAUCGGCUGGGAAAUCCGACAGAUCGGAUUAAGC__UCAUUUCGAAGGGAGCGGCAAAA (((.((((((..(((....)))..)))))).(((((......(((((....)))))........(((......)))............))))).)))........... (-17.55 = -18.42 + 0.87)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:08:52 2006