
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 15,480,780 – 15,480,900 |
| Length | 120 |
| Max. P | 0.966565 |

| Location | 15,480,780 – 15,480,900 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 77.50 |
| Mean single sequence MFE | -46.77 |
| Consensus MFE | -24.00 |
| Energy contribution | -25.42 |
| Covariance contribution | 1.42 |
| Combinations/Pair | 1.39 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.51 |
| SVM decision value | 0.76 |
| SVM RNA-class probability | 0.843758 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15480780 120 + 27905053 AAGACAUUCGCCACGCUCCACGAUUGCCAGAGAAUGGAUCCGUCCAGUCCGUUCAAGGCCACCACGCCGCCCUCGCAGGGCACCUCCUCCCCUUGCUGGGCGCACUGGCAGCGCGGCAAG .........(((.((((........(((((.((((((((.......))))))))..(((......)))((((..((((((..........)))))).))))...))))))))).)))... ( -48.80) >DroVir_CAF1 85316 120 + 1 AACACAUUCGCCACACUCCACGUUUGCCACAAUAGGGAACCAUCGCGUCCAUUGAGCGCCACGACGCCACCCUCGCAGGGCACAUCAUCGCCCUGCUGGGCCGAUUUGCAGCGUGGCAGC .........(((((.((.((.(((.(((.((..((((.......(((((...((.....)).)))))..)))).(((((((........)))))))))))).))).)).)).)))))... ( -44.10) >DroGri_CAF1 24800 120 + 1 AAUACAUUGGCCACGCUCCACGUCUGCCACAGAAUGGAACCGUCGCGACCAUUGAGAGCGACGACGCCACCCUCGCAGGGCACAUCGUCGCCGCCUUGGGCCGACUUGCAGCGCGGCAGU .........(((.((((.((.(((.(((..((..(((...((((((...........))))))...)))..))..((((((...........))))))))).))).)).)))).)))... ( -45.80) >DroWil_CAF1 4215 120 + 1 AAAACAUUCGCCACACUCCACGAUUGCCACAACGUGGAGCCAUCUCGUCCGUUGAGGGCUACAACGCCUCCUUCACAUGGUACCUCGUCAUUUUGCUGGGCAGAUUUGCAACGCGGCAGG .........(((.(.(((((((.(((...))))))))))..((((.((((...((((((......))).((.......))...)))((......)).)))))))).......).)))... ( -31.60) >DroMoj_CAF1 90178 120 + 1 AAGACAUUCGCCACGCUCCACGUUUGCCACAGCGUGGAGCCGUCGCGCCCGUUGAGCGCCACGACUCCGCCCUCGCAGGGCACGUCGUCGCCCUGCUGGGCGGACUUGCAGCGUGGCAGC .........(((((((((((((((......))))))))(((((.((((.......)))).)))..(((((((..(((((((........))))))).)))))))...)).)))))))... ( -66.90) >DroAna_CAF1 4519 120 + 1 AAAACGUUGGCCACGCUCCAGGACUGCCACAUAAUGGAGCCAUCAAUUCCAUUGAGAGCCACCACCCCGCCCUCGCAGGGUACUUCUUCCCCCUGCAUGGCAGCCUGACAUCGCGGCAAC .........(((.((...((((.((((((...(((((((.......)))))))(((.((.........)).)))((((((..........)))))).))))))))))....)).)))... ( -43.40) >consensus AAAACAUUCGCCACGCUCCACGAUUGCCACAGAAUGGAGCCAUCGCGUCCAUUGAGAGCCACGACGCCGCCCUCGCAGGGCACAUCGUCGCCCUGCUGGGCAGACUUGCAGCGCGGCAGC .........(((.((((.((.(.(((((.(((..(((...........))))))....................(((((((........)))))))..))))).).)).)))).)))... (-24.00 = -25.42 + 1.42)


| Location | 15,480,780 – 15,480,900 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 77.50 |
| Mean single sequence MFE | -57.55 |
| Consensus MFE | -30.61 |
| Energy contribution | -31.62 |
| Covariance contribution | 1.01 |
| Combinations/Pair | 1.40 |
| Mean z-score | -2.55 |
| Structure conservation index | 0.53 |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.966565 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
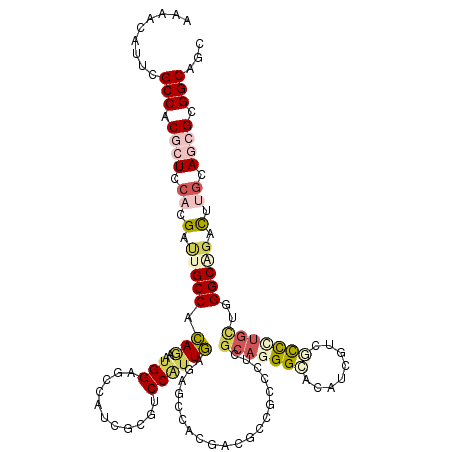
>3R_DroMel_CAF1 15480780 120 - 27905053 CUUGCCGCGCUGCCAGUGCGCCCAGCAAGGGGAGGAGGUGCCCUGCGAGGGCGGCGUGGUGGCCUUGAACGGACUGGACGGAUCCAUUCUCUGGCAAUCGUGGAGCGUGGCGAAUGUCUU .((((((((((.(((((.(((((.(((.(((.(.....).))))))..)))))))..(((.(((..(((.(((((....)).))).)))...))).))).)))))))))))))....... ( -54.30) >DroVir_CAF1 85316 120 - 1 GCUGCCACGCUGCAAAUCGGCCCAGCAGGGCGAUGAUGUGCCCUGCGAGGGUGGCGUCGUGGCGCUCAAUGGACGCGAUGGUUCCCUAUUGUGGCAAACGUGGAGUGUGGCGAAUGUGUU ..(((((((((.((.....(((((((((((((......)))))))).((((.(.(((((((...((....)).))))))).).))))..)).))).....)).)))))))))........ ( -57.60) >DroGri_CAF1 24800 120 - 1 ACUGCCGCGCUGCAAGUCGGCCCAAGGCGGCGACGAUGUGCCCUGCGAGGGUGGCGUCGUCGCUCUCAAUGGUCGCGACGGUUCCAUUCUGUGGCAGACGUGGAGCGUGGCCAAUGUAUU ...((((((((.((.(((((((...((.(((((((((((((((.....)))).))))))))))).))...))))((.((((.......)))).)).))).)).))))))))......... ( -61.70) >DroWil_CAF1 4215 120 - 1 CCUGCCGCGUUGCAAAUCUGCCCAGCAAAAUGACGAGGUACCAUGUGAAGGAGGCGUUGUAGCCCUCAACGGACGAGAUGGCUCCACGUUGUGGCAAUCGUGGAGUGUGGCGAAUGUUUU ..(((((((((.((....((((((((....(.(((.(....).))).).((((.((((....((......))....)))).))))..)))).))))....)).)))))))))........ ( -39.20) >DroMoj_CAF1 90178 120 - 1 GCUGCCACGCUGCAAGUCCGCCCAGCAGGGCGACGACGUGCCCUGCGAGGGCGGAGUCGUGGCGCUCAACGGGCGCGACGGCUCCACGCUGUGGCAAACGUGGAGCGUGGCGAAUGUCUU ..(((((((((.((.((((((((.((((((((......))))))))..))))((((((((.(((((.....))))).)))))))).......))...)).)).)))))))))........ ( -73.30) >DroAna_CAF1 4519 120 - 1 GUUGCCGCGAUGUCAGGCUGCCAUGCAGGGGGAAGAAGUACCCUGCGAGGGCGGGGUGGUGGCUCUCAAUGGAAUUGAUGGCUCCAUUAUGUGGCAGUCCUGGAGCGUGGCCAACGUUUU (((((((((...((((((((((((...(((((....(.((((((((....)))))))).)..)))))((((((.........))))))..))))))).)))))..))))).))))..... ( -59.20) >consensus GCUGCCGCGCUGCAAGUCCGCCCAGCAGGGCGACGAGGUGCCCUGCGAGGGCGGCGUCGUGGCCCUCAACGGACGCGACGGCUCCAUUCUGUGGCAAACGUGGAGCGUGGCGAAUGUCUU ...((((((((((....((((((.(((.((((......)))).)))..))))))....))))).................(((((((............))))))))))))......... (-30.61 = -31.62 + 1.01)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:07:16 2006