
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 15,374,010 – 15,374,116 |
| Length | 106 |
| Max. P | 0.741446 |

| Location | 15,374,010 – 15,374,116 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 75.91 |
| Mean single sequence MFE | -33.63 |
| Consensus MFE | -16.66 |
| Energy contribution | -17.41 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.50 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.741446 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15374010 106 + 27905053 AGCUUGCUG---AGCG-UUGCCAACUUGCCCAAUGU-----GCCAACU--GUGG--AGCCCUUUGUUAGCCCUGCUAAAUAUUCGUUAACGUGGCGAGCAAAUUUCGUUGGCAAACAGA .((((...)---))).-((((((((..(((((..((-----....)).--.)))--.))..((((((.(((.((.(((.......))).)).))).))))))....))))))))..... ( -28.70) >DroVir_CAF1 152775 107 + 1 AGCUACGCGAAAAUAG-UUGCCAACUUGGCCAAUGU-----GCCAACU--GGGC--CG-CCUUUGUUGGCAUUGCUAAAUGUUCGUUAACGUGGCGCAUAAAUUU-GUUAGCCAACAAA .((((((((((..(((-((((((((.(((((...((-----....)).--.)))--))-.....)))))))..))))....))))....)))))).......(((-(((....)))))) ( -34.30) >DroGri_CAF1 160364 107 + 1 AGCUCCACGAAAAUAG-UUCCCAACUUGGCCAAUGU-----GCCAACU--GGGC--CG-CCUUUGUUAGCGUUGCUAAAUGUUCGUUAACGUGGCGCAUAAAUUU-GUUGGCAAACAAA .(((..(((((.....-..((((..(((((......-----))))).)--))).--((-((..((((((((..((.....)).)))))))).))))......)))-)).)))....... ( -32.10) >DroWil_CAF1 166169 109 + 1 -GUUUGUUU---UCCGUUUGCCAACUUGGCCAAUGUGGU--GCCAAAC--GGGGUUGGCUUUGUGUUUGCCUUGCUAAAUAUUCGUUAACGUGGCGCACA-AUUUCGUUGGCACACAU- -........---...((.(((((((..(((((((.(.((--.....))--.).)))))))((((((..(((.((.(((.......))).)).))))))))-)....))))))).))..- ( -35.10) >DroMoj_CAF1 151101 107 + 1 AGCUUCGCGAAAAUAG-UUGCCAACUUGGCCAAUGG-----GCCAACU--GGGC--CA-ACUCUGUUAGCAUUGCUAAAUGUUCGUUAACGUGGCGCAUAAAUUU-GUUGGCAAACAAA ................-((((((((((((((....)-----))))).(--(.((--((-....(((((((...((.....))..))))))))))).)).......-))))))))..... ( -34.60) >DroAna_CAF1 118877 108 + 1 -----GCUC---AGCG-UUGCCAACUUGGCCAAUGGGCUGGGCCAACGAAGCCG--GGUCCUUUGUUAGCCCUGCUAAAUAUUCGUUAACGUGGCGAACAAAUUUCGUUGGCAAACACU -----((((---(...-..(((.....)))...)))))...(((((((((((.(--(((.........)))).))).....((((((.....))))))......))))))))....... ( -37.00) >consensus AGCUUCCCG___AUAG_UUGCCAACUUGGCCAAUGU_____GCCAACU__GGGC__CG_CCUUUGUUAGCCUUGCUAAAUAUUCGUUAACGUGGCGCACAAAUUU_GUUGGCAAACAAA ...............(.(((((((((((((...........)))))..............(..(((((((..............)))))))..)............)))))))).)... (-16.66 = -17.41 + 0.75)



| Location | 15,374,010 – 15,374,116 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 75.91 |
| Mean single sequence MFE | -31.58 |
| Consensus MFE | -15.98 |
| Energy contribution | -16.74 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.51 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.578261 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15374010 106 - 27905053 UCUGUUUGCCAACGAAAUUUGCUCGCCACGUUAACGAAUAUUUAGCAGGGCUAACAAAGGGCU--CCAC--AGUUGGC-----ACAUUGGGCAAGUUGGCAA-CGCU---CAGCAAGCU .....((((((((.....((((((((((((((((.......))))).(((((.......))).--))..--.).))))-----.....))))))))))))))-.(((---.....))). ( -29.20) >DroVir_CAF1 152775 107 - 1 UUUGUUGGCUAAC-AAAUUUAUGCGCCACGUUAACGAACAUUUAGCAAUGCCAACAAAGG-CG--GCCC--AGUUGGC-----ACAUUGGCCAAGUUGGCAA-CUAUUUUCGCGUAGCU ((((((....)))-))).(((((((((..(((((.......)))))...(((......))-))--))((--(((((((-----......))))).))))...-........)))))).. ( -28.70) >DroGri_CAF1 160364 107 - 1 UUUGUUUGCCAAC-AAAUUUAUGCGCCACGUUAACGAACAUUUAGCAACGCUAACAAAGG-CG--GCCC--AGUUGGC-----ACAUUGGCCAAGUUGGGAA-CUAUUUUCGUGGAGCU ((((((....)))-))).....((.(((((..((..............((((......))-))--.(((--(((((((-----......))))).)))))..-...))..))))).)). ( -31.30) >DroWil_CAF1 166169 109 - 1 -AUGUGUGCCAACGAAAU-UGUGCGCCACGUUAACGAAUAUUUAGCAAGGCAAACACAAAGCCAACCCC--GUUUGGC--ACCACAUUGGCCAAGUUGGCAAACGGA---AAACAAAC- -.(((.(((((((....(-((((.(((..(((((.......)))))..)))...))))).(((((....--..)))))--..............))))))).)))..---........- ( -30.10) >DroMoj_CAF1 151101 107 - 1 UUUGUUUGCCAAC-AAAUUUAUGCGCCACGUUAACGAACAUUUAGCAAUGCUAACAGAGU-UG--GCCC--AGUUGGC-----CCAUUGGCCAAGUUGGCAA-CUAUUUUCGCGAAGCU (((((((((((((-.......((.((((((....)).....(((((...)))))......-))--)).)--).(((((-----(....))))))))))))))-........)))))... ( -30.70) >DroAna_CAF1 118877 108 - 1 AGUGUUUGCCAACGAAAUUUGUUCGCCACGUUAACGAAUAUUUAGCAGGGCUAACAAAGGACC--CGGCUUCGUUGGCCCAGCCCAUUGGCCAAGUUGGCAA-CGCU---GAGC----- ((((((.(((((((((.....)))((((((....))........((.((((((((.(((....--...))).)))))))).))....))))...))))))))-))))---....----- ( -39.50) >consensus UUUGUUUGCCAAC_AAAUUUAUGCGCCACGUUAACGAACAUUUAGCAAGGCUAACAAAGG_CG__CCCC__AGUUGGC_____ACAUUGGCCAAGUUGGCAA_CGAU___CACCAAGCU ...(.((((((((...........((((.(((((.......)))))...((((((.................)))))).........))))...)))))))).)............... (-15.98 = -16.74 + 0.75)

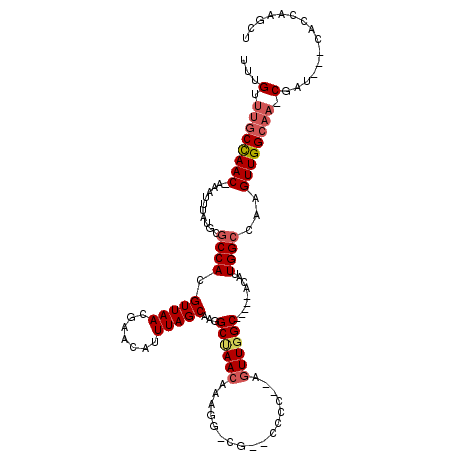

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:06:32 2006