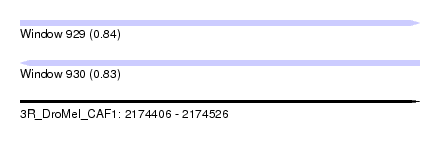

| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 2,174,406 – 2,174,526 |
| Length | 120 |
| Max. P | 0.837857 |

| Location | 2,174,406 – 2,174,526 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.25 |
| Mean single sequence MFE | -40.60 |
| Consensus MFE | -38.38 |
| Energy contribution | -39.12 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.74 |
| SVM RNA-class probability | 0.837857 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 2174406 120 + 27905053 GGUCCGCAAUAGUAGUAGAUGAUUGUCACUUUUGACGCGCCACGCCCCCUGCCAGGGGCGUCUCGGGCAAAUAAAAAUCCUCAAAUCACGCUUUGGCACAAGGCCCUACUACACUGGGAG ..(((.((...(((((((.(((((((((....))))..((((((((((......)))))))....)))...............))))).((((((...)))))).)))))))..))))). ( -41.10) >DroSec_CAF1 316824 120 + 1 GGUCCGCAAUGGUAGUAGAUGAUUGUCACUUUUGACGUGCCACGCCCCCUGCCAGGGGCGUUUCGGGCAAAUAAAAAUCCUCAAAUCACGCUUUGGCACAAGGCACUACUACACUGGGAG ..(((.((...(((((((.(((((((((....)))).(((((((((((......)))))))....))))..............))))).((((((...)))))).)))))))..))))). ( -43.90) >DroSim_CAF1 333667 120 + 1 GGUCCGCAAUAGUAGUAGAUGAUUGUCACUUUUGACGCGCCACGCCCCCUGCCAGGGGCGUCUCGGGCAAAUAAAAAUCCUCAAAUCACGCUUUGGCACAAGGCACUACUACACUGGGAG ..(((.((...(((((((.(((((((((....))))..((((((((((......)))))))....)))...............))))).((((((...)))))).)))))))..))))). ( -42.20) >DroYak_CAF1 320465 120 + 1 GGUCCGCAAUAGUAGUAGAUGAUUGUCACUUUUGACGCGCCACGCCCCCUGCCAGGGGCGUCUCGGGCAAAUAGAAAUCCUCAAUUCACGCUUUGGCACAAGGCACUAAUACACUGGCAA .....((..((((.(((((.((((((((....))))..((((((((((......)))))))....))).......)))).)).......((((((...)))))).....))))))))).. ( -35.20) >consensus GGUCCGCAAUAGUAGUAGAUGAUUGUCACUUUUGACGCGCCACGCCCCCUGCCAGGGGCGUCUCGGGCAAAUAAAAAUCCUCAAAUCACGCUUUGGCACAAGGCACUACUACACUGGGAG ..(((.((...(((((((.(((((((((....))))..((((((((((......)))))))....)))...............))))).((((((...)))))).)))))))..))))). (-38.38 = -39.12 + 0.75)

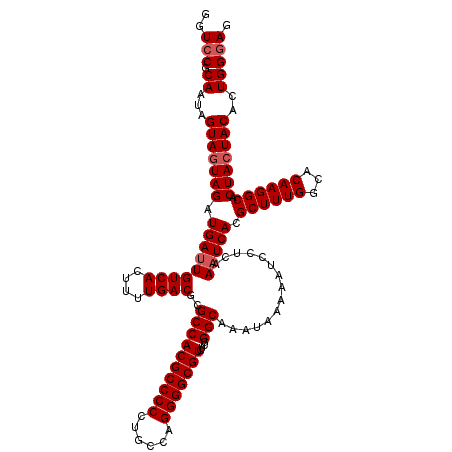
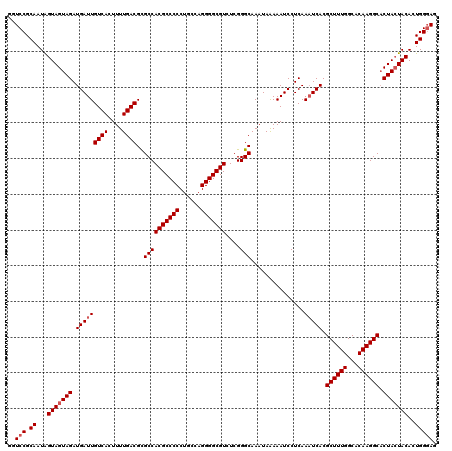
| Location | 2,174,406 – 2,174,526 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.25 |
| Mean single sequence MFE | -43.88 |
| Consensus MFE | -41.99 |
| Energy contribution | -42.43 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.828245 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 2174406 120 - 27905053 CUCCCAGUGUAGUAGGGCCUUGUGCCAAAGCGUGAUUUGAGGAUUUUUAUUUGCCCGAGACGCCCCUGGCAGGGGGCGUGGCGCGUCAAAAGUGACAAUCAUCUACUACUAUUGCGGACC .(((((((((((((((.....(((((.........((((.((...........))))))((((((((....)))))))))))))((((....)))).....)))))))).)))).))).. ( -46.40) >DroSec_CAF1 316824 120 - 1 CUCCCAGUGUAGUAGUGCCUUGUGCCAAAGCGUGAUUUGAGGAUUUUUAUUUGCCCGAAACGCCCCUGGCAGGGGGCGUGGCACGUCAAAAGUGACAAUCAUCUACUACCAUUGCGGACC .((((((((((((((((....(((((.........((((.((...........))))))((((((((....)))))))))))))((((....))))...)).)))))).))))).))).. ( -45.90) >DroSim_CAF1 333667 120 - 1 CUCCCAGUGUAGUAGUGCCUUGUGCCAAAGCGUGAUUUGAGGAUUUUUAUUUGCCCGAGACGCCCCUGGCAGGGGGCGUGGCGCGUCAAAAGUGACAAUCAUCUACUACUAUUGCGGACC .((((((((((((((((....(((((.........((((.((...........))))))((((((((....)))))))))))))((((....))))...)).))))))).)))).))).. ( -45.30) >DroYak_CAF1 320465 120 - 1 UUGCCAGUGUAUUAGUGCCUUGUGCCAAAGCGUGAAUUGAGGAUUUCUAUUUGCCCGAGACGCCCCUGGCAGGGGGCGUGGCGCGUCAAAAGUGACAAUCAUCUACUACUAUUGCGGACC .......((((.(((((....(((((...(.(..(((.(((...))).)))..).)...((((((((....)))))))))))))((((....))))..........))))).)))).... ( -37.90) >consensus CUCCCAGUGUAGUAGUGCCUUGUGCCAAAGCGUGAUUUGAGGAUUUUUAUUUGCCCGAGACGCCCCUGGCAGGGGGCGUGGCGCGUCAAAAGUGACAAUCAUCUACUACUAUUGCGGACC .((((((((((((((......(((((.........((((.((...........))))))((((((((....)))))))))))))((((....))))......))))))).)))).))).. (-41.99 = -42.43 + 0.44)

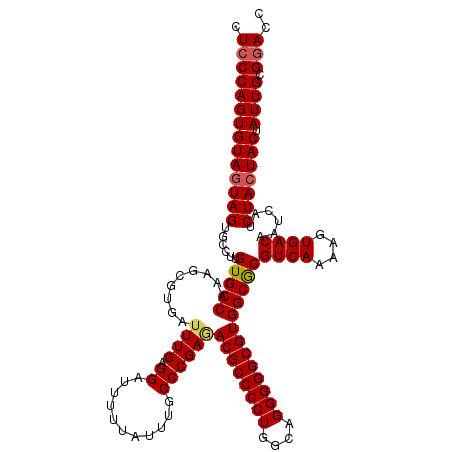
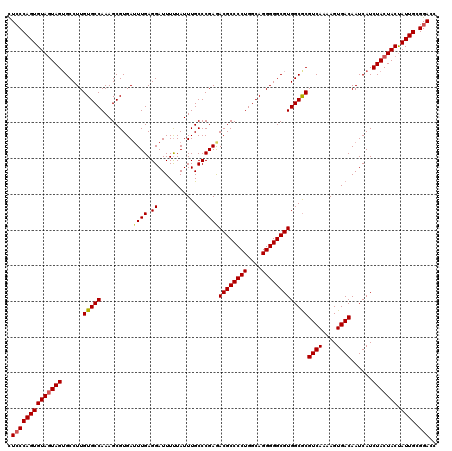
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:46:42 2006