
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 15,152,706 – 15,152,866 |
| Length | 160 |
| Max. P | 0.992946 |

| Location | 15,152,706 – 15,152,826 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.44 |
| Mean single sequence MFE | -28.41 |
| Consensus MFE | -25.25 |
| Energy contribution | -26.37 |
| Covariance contribution | 1.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.620987 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15152706 120 + 27905053 GAUCAAAAUCGAAUGAGUUUCACUUCGCUAAAAACAACUUCUGCUGAAGACGUUUUCAGAGACGGAAAAUGCAUCAUCAUAUGAAUUUUUUUGGCUCUGAUAACUUCUCCAGGACUGUCG (((......((((((.....)).))))................(((.(((.(((.((((((.(((((((..(((......)))...))))))).)))))).))).))).)))....))). ( -25.60) >DroSec_CAF1 76305 120 + 1 GAUCAAAAUCGAAUGAGCUUCACUUCGCUAAAAUCAACUUCUGCUGAAGAAAUUUUCAGAGACUGAAAAUGCAUCAUCAUAUGAAUUUUGUUGGCUCUGAAAACUUCUCCAGGUCUGCCG ((((......((...(((........)))....)).........((.((((.(((((((((.(..(.......((((...))))......)..)))))))))).)))).))))))..... ( -24.32) >DroSim_CAF1 75333 120 + 1 GAUCAAAAUCGAAUGAGCUUCACUUCGCUAAAAUCAACUUCUGUUGAAGAAGUUUUCAGAGACUGAAAAUGCAUCAUCAUAUGAAUUUUUUUGGCUCUGAAAACUUCUCCAGGUCUGCCG ((((.....((((((.....)).))))......(((((....)))))((((((((((((((.(..((((....((((...))))...))))..)))))))))))))))...))))..... ( -35.30) >consensus GAUCAAAAUCGAAUGAGCUUCACUUCGCUAAAAUCAACUUCUGCUGAAGAAGUUUUCAGAGACUGAAAAUGCAUCAUCAUAUGAAUUUUUUUGGCUCUGAAAACUUCUCCAGGUCUGCCG .........((((((.....)).))))................(((.((((((((((((((.(..((((....((((...))))...))))..))))))))))))))).)))........ (-25.25 = -26.37 + 1.11)



| Location | 15,152,706 – 15,152,826 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.44 |
| Mean single sequence MFE | -32.50 |
| Consensus MFE | -30.61 |
| Energy contribution | -31.17 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.94 |
| SVM decision value | 1.37 |
| SVM RNA-class probability | 0.946951 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15152706 120 - 27905053 CGACAGUCCUGGAGAAGUUAUCAGAGCCAAAAAAAUUCAUAUGAUGAUGCAUUUUCCGUCUCUGAAAACGUCUUCAGCAGAAGUUGUUUUUAGCGAAGUGAAACUCAUUCGAUUUUGAUC (((.(((((((((((.(((.((((((.(...(((((.(((......))).)))))..).)))))).))).)))))))..(((...(((((.........)))))...))))))))))... ( -27.10) >DroSec_CAF1 76305 120 - 1 CGGCAGACCUGGAGAAGUUUUCAGAGCCAACAAAAUUCAUAUGAUGAUGCAUUUUCAGUCUCUGAAAAUUUCUUCAGCAGAAGUUGAUUUUAGCGAAGUGAAGCUCAUUCGAUUUUGAUC ........((((((((((((((((((.(...(((((.(((......))).)))))..).))))))))))))))))))..((((((((....(((........)))...)))))))).... ( -35.10) >DroSim_CAF1 75333 120 - 1 CGGCAGACCUGGAGAAGUUUUCAGAGCCAAAAAAAUUCAUAUGAUGAUGCAUUUUCAGUCUCUGAAAACUUCUUCAACAGAAGUUGAUUUUAGCGAAGUGAAGCUCAUUCGAUUUUGAUC .((....))(((((((((((((((((.(...(((((.(((......))).)))))..).)))))))))))))))))...((((((((....(((........)))...)))))))).... ( -35.30) >consensus CGGCAGACCUGGAGAAGUUUUCAGAGCCAAAAAAAUUCAUAUGAUGAUGCAUUUUCAGUCUCUGAAAACUUCUUCAGCAGAAGUUGAUUUUAGCGAAGUGAAGCUCAUUCGAUUUUGAUC ((((....((((((((((((((((((.(...(((((.(((......))).)))))..).)))))))))))))))))).....)))).......(((((((((.....))).))))))... (-30.61 = -31.17 + 0.56)



| Location | 15,152,746 – 15,152,866 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.00 |
| Mean single sequence MFE | -39.21 |
| Consensus MFE | -38.01 |
| Energy contribution | -39.23 |
| Covariance contribution | 1.23 |
| Combinations/Pair | 1.05 |
| Mean z-score | -3.35 |
| Structure conservation index | 0.97 |
| SVM decision value | 2.36 |
| SVM RNA-class probability | 0.992946 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15152746 120 + 27905053 CUGCUGAAGACGUUUUCAGAGACGGAAAAUGCAUCAUCAUAUGAAUUUUUUUGGCUCUGAUAACUUCUCCAGGACUGUCGCUCUUAUGGAGGUGAAACUCCUUCGAGUUUGGGCAUACUA ...(((.(((.(((.((((((.(((((((..(((......)))...))))))).)))))).))).))).)))...((((((((....((((......))))...))))...))))..... ( -35.20) >DroSec_CAF1 76345 120 + 1 CUGCUGAAGAAAUUUUCAGAGACUGAAAAUGCAUCAUCAUAUGAAUUUUGUUGGCUCUGAAAACUUCUCCAGGUCUGCCGCUCCUAUGGAGGUGAAACUCCUUCGAGUUUGGGCAUACUA ...(((.((((.(((((((((.(..(.......((((...))))......)..)))))))))).)))).)))((.((((((((....((((......))))...))))...)))).)).. ( -37.62) >DroSim_CAF1 75373 120 + 1 CUGUUGAAGAAGUUUUCAGAGACUGAAAAUGCAUCAUCAUAUGAAUUUUUUUGGCUCUGAAAACUUCUCCAGGUCUGCCGCUCCUAUGGAGGUGAAACUCCUUCGAGUUUGGGCAUACUA ....((.((((((((((((((.(..((((....((((...))))...))))..))))))))))))))).))(((.((((((((....((((......))))...))))...)))).))). ( -44.80) >consensus CUGCUGAAGAAGUUUUCAGAGACUGAAAAUGCAUCAUCAUAUGAAUUUUUUUGGCUCUGAAAACUUCUCCAGGUCUGCCGCUCCUAUGGAGGUGAAACUCCUUCGAGUUUGGGCAUACUA ...(((.((((((((((((((.(..((((....((((...))))...))))..))))))))))))))).)))((.((((((((....((((......))))...))))...)))).)).. (-38.01 = -39.23 + 1.23)

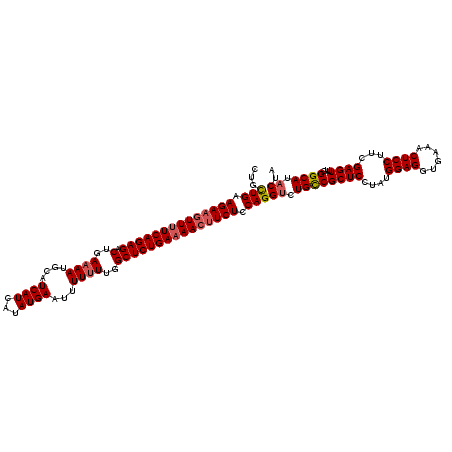

| Location | 15,152,746 – 15,152,866 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.00 |
| Mean single sequence MFE | -38.47 |
| Consensus MFE | -36.75 |
| Energy contribution | -37.87 |
| Covariance contribution | 1.11 |
| Combinations/Pair | 1.02 |
| Mean z-score | -3.93 |
| Structure conservation index | 0.96 |
| SVM decision value | 2.19 |
| SVM RNA-class probability | 0.989884 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15152746 120 - 27905053 UAGUAUGCCCAAACUCGAAGGAGUUUCACCUCCAUAAGAGCGACAGUCCUGGAGAAGUUAUCAGAGCCAAAAAAAUUCAUAUGAUGAUGCAUUUUCCGUCUCUGAAAACGUCUUCAGCAG ..((......((((((....))))))..((((.....))).)))....(((((((.(((.((((((.(...(((((.(((......))).)))))..).)))))).))).)))))))... ( -29.30) >DroSec_CAF1 76345 120 - 1 UAGUAUGCCCAAACUCGAAGGAGUUUCACCUCCAUAGGAGCGGCAGACCUGGAGAAGUUUUCAGAGCCAACAAAAUUCAUAUGAUGAUGCAUUUUCAGUCUCUGAAAAUUUCUUCAGCAG ..((.((((.((((((....))))))...((((...)))).)))).))((((((((((((((((((.(...(((((.(((......))).)))))..).))))))))))))))))))... ( -43.10) >DroSim_CAF1 75373 120 - 1 UAGUAUGCCCAAACUCGAAGGAGUUUCACCUCCAUAGGAGCGGCAGACCUGGAGAAGUUUUCAGAGCCAAAAAAAUUCAUAUGAUGAUGCAUUUUCAGUCUCUGAAAACUUCUUCAACAG ..((.((((.((((((....))))))...((((...)))).)))).)).(((((((((((((((((.(...(((((.(((......))).)))))..).))))))))))))))))).... ( -43.00) >consensus UAGUAUGCCCAAACUCGAAGGAGUUUCACCUCCAUAGGAGCGGCAGACCUGGAGAAGUUUUCAGAGCCAAAAAAAUUCAUAUGAUGAUGCAUUUUCAGUCUCUGAAAACUUCUUCAGCAG .....((((.((((((....))))))...(((.....))).))))...((((((((((((((((((.(...(((((.(((......))).)))))..).))))))))))))))))))... (-36.75 = -37.87 + 1.11)


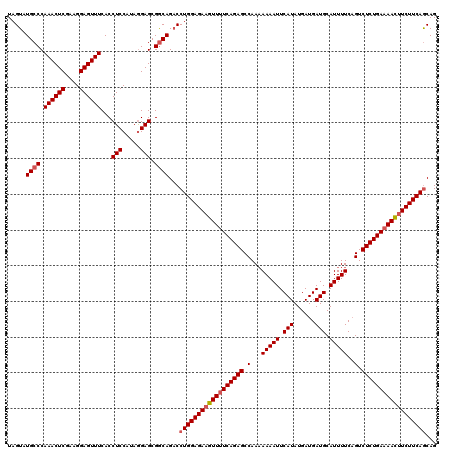
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:05:10 2006