| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 15,045,238 – 15,045,331 |
| Length | 93 |
| Max. P | 0.998991 |

| Location | 15,045,238 – 15,045,331 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 72.24 |
| Mean single sequence MFE | -26.72 |
| Consensus MFE | -8.68 |
| Energy contribution | -9.55 |
| Covariance contribution | 0.87 |
| Combinations/Pair | 1.27 |
| Mean z-score | -2.55 |
| Structure conservation index | 0.32 |
| SVM decision value | 1.53 |
| SVM RNA-class probability | 0.961357 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15045238 93 + 27905053 CUGUGGAGCAGAAAAAAAAAGGA-GUUGGGAAAAUUGUUAUCCU-CGUGACAAGGGGCUU-----UUUGCCCAGUUAACUCUCUUACCUCUUCCACAGUC ((((((((.((.......(((((-(((((.....(((((((...-.))))))).((((..-----...))))..))))))).)))..)).)))))))).. ( -29.80) >DroPse_CAF1 50729 92 + 1 CUGAC----AAAAAA-AAGAAGAAGAGAAGAAAAGUUUCAUUAACUCU---GAGCGGCUUGCGGGUCUCCUCCUUCUCCUCUCUUACCUCUUCCACAGUC .....----......-((((.((.((((((...((((.....))))..---(((.((((....))))..))))))))).))))))............... ( -17.00) >DroSim_CAF1 47444 92 + 1 CUGUGGAGCAGAAAA-AAGAGGA-GUUGGGCAAAUAGUUCUCCU-CGUGACAACGGGCUU-----UUUGCCCAGUUAACUCUCUUACCUCUUCCACAGUC ((((((((.((..((-.((((..-.(((((((((.((.....((-(((....))))))).-----)))))))))....)))).))..)).)))))))).. ( -34.70) >DroEre_CAF1 49553 92 + 1 CUGUGGAGCAGAAAA-AAGGAGA-GUUGGGAAAAUAGUUCUCCA-CAUGACAGGGGGCUU-----UUUCCCCAGUUAACUCUCUUACCUCUUCCACAGUC ((((((((.((....-.((((((-((((((((((.(((((.((.-.......))))))))-----)))))).....)))))))))..)).)))))))).. ( -35.80) >DroYak_CAF1 71362 92 + 1 CUGUGGAACAGAAAA-AAGGAGA-GAUGGAAAAAUAGUUAUCCA-UAUGACAGCGGGCUU-----UUUCUCCAGUUAACUCUCUUACCUCUUCCACAGUC ((((((((.((....-.((((((-((((((.((....)).))))-).((((.(.(((...-----...)))).)))).)))))))..)).)))))))).. ( -26.00) >DroPer_CAF1 50773 91 + 1 CUGAC----AAAA-A-AAGAAGAAGAGAAGAAAAGUUUCAUUAACUCU---GAGCGGCUUGCGGGUCUCCUCCUUCUCCUCUCUUACCUCUUCCACAGUC .....----....-.-((((.((.((((((...((((.....))))..---(((.((((....))))..))))))))).))))))............... ( -17.00) >consensus CUGUGGAGCAGAAAA_AAGAAGA_GAUGGGAAAAUAGUUAUCCA_CAUGACAAGGGGCUU_____UUUCCCCAGUUAACUCUCUUACCUCUUCCACAGUC ((((((((.((.....(((.(((.((((((.......................................))))))....))))))..)).)))))))).. ( -8.68 = -9.55 + 0.87)
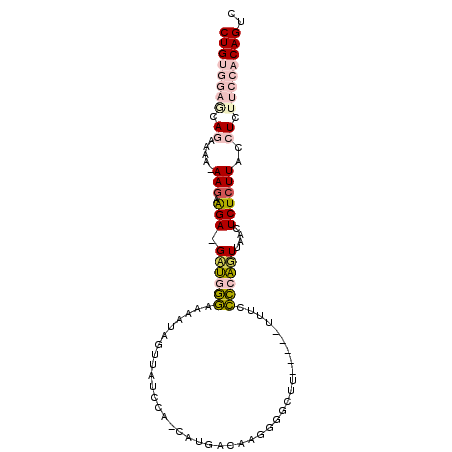



| Location | 15,045,238 – 15,045,331 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 72.24 |
| Mean single sequence MFE | -31.60 |
| Consensus MFE | -16.31 |
| Energy contribution | -15.90 |
| Covariance contribution | -0.41 |
| Combinations/Pair | 1.36 |
| Mean z-score | -3.27 |
| Structure conservation index | 0.52 |
| SVM decision value | 3.32 |
| SVM RNA-class probability | 0.998991 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 15045238 93 - 27905053 GACUGUGGAAGAGGUAAGAGAGUUAACUGGGCAAA-----AAGCCCCUUGUCACG-AGGAUAACAAUUUUCCCAAC-UCCUUUUUUUUUCUGCUCCACAG ..((((((((((((.(((.(((((....((((...-----..))))........(-(((((....))))))..)))-)))))....)))))..))))))) ( -27.80) >DroPse_CAF1 50729 92 - 1 GACUGUGGAAGAGGUAAGAGAGGAGAAGGAGGAGACCCGCAAGCCGCUC---AGAGUUAAUGAAACUUUUCUUCUCUUCUUCUU-UUUUUU----GUCAG (((...((((((((.(((((((((((((...(((...(....)...)))---.............)))))))))))))))))))-))....----))).. ( -32.09) >DroSim_CAF1 47444 92 - 1 GACUGUGGAAGAGGUAAGAGAGUUAACUGGGCAAA-----AAGCCCGUUGUCACG-AGGAGAACUAUUUGCCCAAC-UCCUCUU-UUUUCUGCUCCACAG ..((((((((((((.((((((((....((((((((-----.(((((((....)))-.))....)).))))))))))-).)))))-.)))))..))))))) ( -35.30) >DroEre_CAF1 49553 92 - 1 GACUGUGGAAGAGGUAAGAGAGUUAACUGGGGAAA-----AAGCCCCCUGUCAUG-UGGAGAACUAUUUUCCCAAC-UCUCCUU-UUUUCUGCUCCACAG ..((((((((((((.(((((((((.((.((((...-----....)))).))...(-.(((((.....)))))))))-))).)))-.)))))..))))))) ( -32.70) >DroYak_CAF1 71362 92 - 1 GACUGUGGAAGAGGUAAGAGAGUUAACUGGAGAAA-----AAGCCCGCUGUCAUA-UGGAUAACUAUUUUUCCAUC-UCUCCUU-UUUUCUGUUCCACAG ..((((((((.(((.(((((((.....((((((((-----.((.(((........-)))....)).)))))))).)-))).)))-...))).)))))))) ( -28.90) >DroPer_CAF1 50773 91 - 1 GACUGUGGAAGAGGUAAGAGAGGAGAAGGAGGAGACCCGCAAGCCGCUC---AGAGUUAAUGAAACUUUUCUUCUCUUCUUCUU-U-UUUU----GUCAG (((...((((((((.(((((((((((((...(((...(....)...)))---.............)))))))))))))))))))-)-)...----))).. ( -32.79) >consensus GACUGUGGAAGAGGUAAGAGAGUUAACUGGGGAAA_____AAGCCCCUUGUCACA_UGGAUAACAAUUUUCCCAAC_UCUUCUU_UUUUCUGCUCCACAG ..((((((((((((...(((((.....((((((((...............................)))))))).....)))))..)))))..))))))) (-16.31 = -15.90 + -0.41)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:04:24 2006