
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 14,993,110 – 14,993,268 |
| Length | 158 |
| Max. P | 0.990093 |

| Location | 14,993,110 – 14,993,228 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.30 |
| Mean single sequence MFE | -35.03 |
| Consensus MFE | -31.97 |
| Energy contribution | -32.30 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.31 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.04 |
| SVM RNA-class probability | 0.905251 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14993110 118 - 27905053 CCAGAAAUUGUUGGUGACAACCACCAGAUGGAAAGCGAAAUGGAAUCGCAUUCUGGCCCGAAGAACCAAAAUCGCGAUUACUCUGUGGCCAGGUGACGAUGGGCAGGCAAAAAAAA--AA .(((((....((((((.....)))))).......((((.......)))).)))))(((((..(.(((....(((((.......)))))...)))..)..)))))............--.. ( -30.10) >DroSim_CAF1 12176 120 - 1 CCAGAAAUUGUUGGUGACAACCACCAGAUGGAAAGCGAAAUGGAAUCGCAUUCUGGCCCGAAGAACCAAAAUCGCGGUUACUCUGUGGGCAGGUGACGAUGGGCUGGCAAAAAAAAAAAA ((((..(((((((....))))((((....((((.((((.......)))).)))).(((((.((((((........)))...))).))))).))))..)))...))))............. ( -37.90) >DroYak_CAF1 12644 113 - 1 CCAGAAAUUGUUGGUGACAACCACCAGAUGGAAAGCGAAAUGGAAUCGCAUUCUGGCCCGAAGAACCAAAAUCGCGGUUACUCUGUGGGCAGGUG---AUGGGCUGGCAAAAAAGU---- ((((.....((((....))))((((....((((.((((.......)))).)))).(((((.((((((........)))...))).))))).))))---.....)))).........---- ( -37.10) >consensus CCAGAAAUUGUUGGUGACAACCACCAGAUGGAAAGCGAAAUGGAAUCGCAUUCUGGCCCGAAGAACCAAAAUCGCGGUUACUCUGUGGGCAGGUGACGAUGGGCUGGCAAAAAAAA__AA .(((((....((((((.....)))))).......((((.......)))).)))))(((((....(((....((((((.....))))))...))).....)))))................ (-31.97 = -32.30 + 0.33)

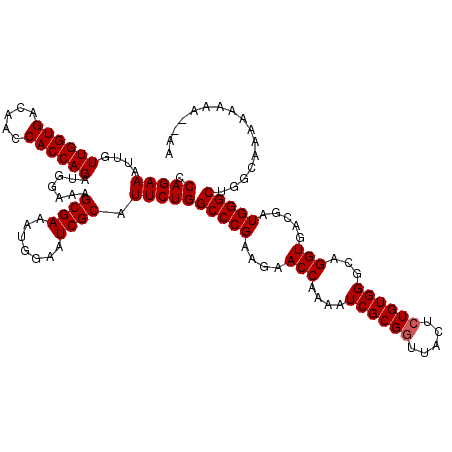
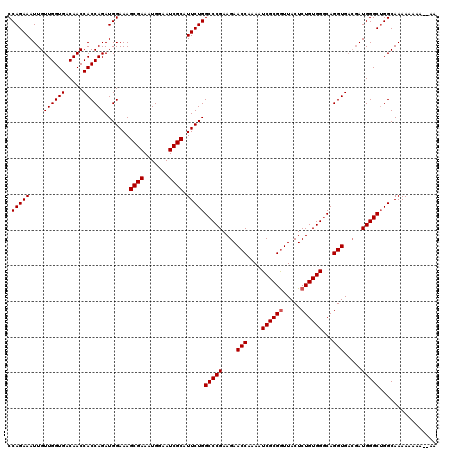
| Location | 14,993,148 – 14,993,268 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.89 |
| Mean single sequence MFE | -34.90 |
| Consensus MFE | -33.91 |
| Energy contribution | -33.47 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.06 |
| Mean z-score | -3.03 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.94 |
| SVM RNA-class probability | 0.983477 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14993148 120 + 27905053 UAAUCGCGAUUUUGGUUCUUCGGGCCAGAAUGCGAUUCCAUUUCGCUUUCCAUCUGGUGGUUGUCACCAACAAUUUCUGGCCGCAAGCGCCUUUUUUGUUUAUCUUGUGAUUUUUUCCGU .((((((.((((((((.(....))))))))))))))).....((((.........((((.....))))(((((.....((((....).)))....)))))......)))).......... ( -34.70) >DroSim_CAF1 12216 120 + 1 UAACCGCGAUUUUGGUUCUUCGGGCCAGAAUGCGAUUCCAUUUCGCUUUCCAUCUGGUGGUUGUCACCAACAAUUUCUGGCCGCAAGCGCCUUUUUUGUUUAUCUUGUGACUUUUUCCGU .((((((.((((((((.(....)))))))))((((.......))))..........))))))(((((.(((((.....((((....).)))....)))))......)))))......... ( -36.10) >DroYak_CAF1 12677 120 + 1 UAACCGCGAUUUUGGUUCUUCGGGCCAGAAUGCGAUUCCAUUUCGCUUUCCAUCUGGUGGUUGUCACCAACAAUUUCUGGCCGCAAGCGCCUUUUUUGUUUAUCUUGUGAUUUUUUCCGU .((((((.((((((((.(....)))))))))((((.......))))..........))))))(((((.(((((.....((((....).)))....)))))......)))))......... ( -33.90) >consensus UAACCGCGAUUUUGGUUCUUCGGGCCAGAAUGCGAUUCCAUUUCGCUUUCCAUCUGGUGGUUGUCACCAACAAUUUCUGGCCGCAAGCGCCUUUUUUGUUUAUCUUGUGAUUUUUUCCGU .((((((.(((((((((.....)))))))))((((.......))))..........))))))(((((.(((((.....((((....).)))....)))))......)))))......... (-33.91 = -33.47 + -0.44)

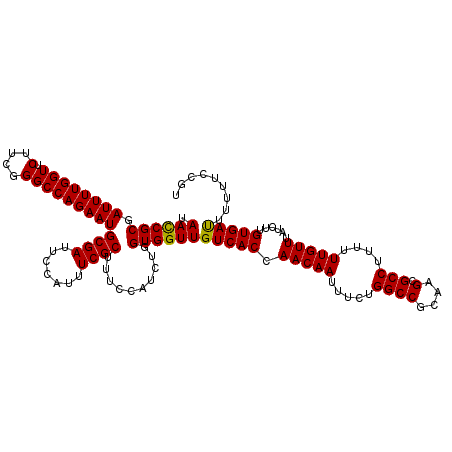

| Location | 14,993,148 – 14,993,268 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.89 |
| Mean single sequence MFE | -32.83 |
| Consensus MFE | -33.05 |
| Energy contribution | -32.83 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.93 |
| Structure conservation index | 1.01 |
| SVM decision value | 2.20 |
| SVM RNA-class probability | 0.990093 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14993148 120 - 27905053 ACGGAAAAAAUCACAAGAUAAACAAAAAAGGCGCUUGCGGCCAGAAAUUGUUGGUGACAACCACCAGAUGGAAAGCGAAAUGGAAUCGCAUUCUGGCCCGAAGAACCAAAAUCGCGAUUA .((((....(((....)))..........((..((((.((((((((....((((((.....)))))).......((((.......)))).))))))))).)))..))....)).)).... ( -32.60) >DroSim_CAF1 12216 120 - 1 ACGGAAAAAGUCACAAGAUAAACAAAAAAGGCGCUUGCGGCCAGAAAUUGUUGGUGACAACCACCAGAUGGAAAGCGAAAUGGAAUCGCAUUCUGGCCCGAAGAACCAAAAUCGCGGUUA .((((....(((....)))..........((..((((.((((((((....((((((.....)))))).......((((.......)))).))))))))).)))..))....)).)).... ( -33.10) >DroYak_CAF1 12677 120 - 1 ACGGAAAAAAUCACAAGAUAAACAAAAAAGGCGCUUGCGGCCAGAAAUUGUUGGUGACAACCACCAGAUGGAAAGCGAAAUGGAAUCGCAUUCUGGCCCGAAGAACCAAAAUCGCGGUUA .((((....(((....)))..........((..((((.((((((((....((((((.....)))))).......((((.......)))).))))))))).)))..))....)).)).... ( -32.80) >consensus ACGGAAAAAAUCACAAGAUAAACAAAAAAGGCGCUUGCGGCCAGAAAUUGUUGGUGACAACCACCAGAUGGAAAGCGAAAUGGAAUCGCAUUCUGGCCCGAAGAACCAAAAUCGCGGUUA .((((....(((....)))..........((..((((.((((((((....((((((.....)))))).......((((.......)))).))))))))).)))..))....)).)).... (-33.05 = -32.83 + -0.22)

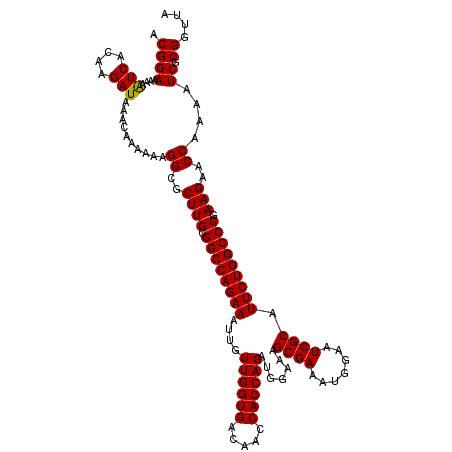

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:04:12 2006