
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 14,954,969 – 14,955,117 |
| Length | 148 |
| Max. P | 0.846921 |

| Location | 14,954,969 – 14,955,067 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 80.66 |
| Mean single sequence MFE | -30.61 |
| Consensus MFE | -20.49 |
| Energy contribution | -19.44 |
| Covariance contribution | -1.05 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.636001 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14954969 98 + 27905053 AAAGUC---------CGCAUCGACUGCCAACAGCAGGACCAGCACCUUUGAGCGUAUACCCCAAAAGAAAAGCGAGCCGAAGAUGUCGAUCUCCAGCAAACUGGAGA ...(((---------.(((((..((((.....))))..........((((.((......................))))))))))).)))((((((....)))))). ( -23.25) >DroPse_CAF1 4636 107 + 1 AAAAUCAUUGAAUGUGGCCUCCACGGCCAACAGUCGGACGAGCACCUUUGAGCGCCAGCCGCAGAAGAAGAACGAGCCGAAGAUGGCCAUCUCCAGCAAGCUGGAGA .............((((...))))(((((....((((.((..(..((((..(((.....))).))))..)..))..))))...))))).(((((((....))))))) ( -36.80) >DroEre_CAF1 21865 98 + 1 AAAGUC---------CGCAUCGACUGCCAACAGCAGGACGAGCACCUUUGAGCGCAUACCCCAAAAGAAGAGCGAGUCGAAGAUGUCCAUCUCCAGCAAACUGGAGA ......---------.((.(((.((((.....))))..)))))..((((((.(((................)))..)))))).......(((((((....))))))) ( -28.39) >DroYak_CAF1 5489 98 + 1 AAAGUC---------CGCAUCGGCUGCCAACAGUAGGACCAGCACAUUUGAGCGCGCGCCCCAAAAGAAUAGCGAAGCGAAGAUGUCCAUCUCCAGCAAACUGGAGA ...(((---------(......((((....)))).))))..(.((((((..((...(((............)))..))..)))))).).(((((((....))))))) ( -29.10) >DroAna_CAF1 5361 98 + 1 CAAGGC---------GGCCUCCACGGCCAAUAGUCGGACCAGCACCUUUGAGCGCCAGCCCCAGAAGAUGAGUGAACCGAAGAUGUCGAUUUCCAGCAAGUUGGAAA ...(((---------((((.....)))).((..((((..((.((.((((..((....))....)))).))..))..))))..)))))..(((((((....))))))) ( -27.90) >DroPer_CAF1 4640 107 + 1 AAAAUCAUUGAAUGUGGCCUCCACGGCCAACAGUCGGACGAGCACCUUUGAGCGCCAGCCGCAGAAGAAGAGCGAGCCGAAGAUGGCCAUCUCCAGCAAGCUGGAGA .............((((...))))(((((....((((.((..(..((((..(((.....))).))))..)..))..))))...))))).(((((((....))))))) ( -38.20) >consensus AAAGUC_________CGCAUCCACGGCCAACAGUAGGACCAGCACCUUUGAGCGCCAGCCCCAAAAGAAGAGCGAGCCGAAGAUGUCCAUCUCCAGCAAACUGGAGA ................((.....((((.....)))).....))..(((((..(((................)))...))))).......(((((((....))))))) (-20.49 = -19.44 + -1.05)


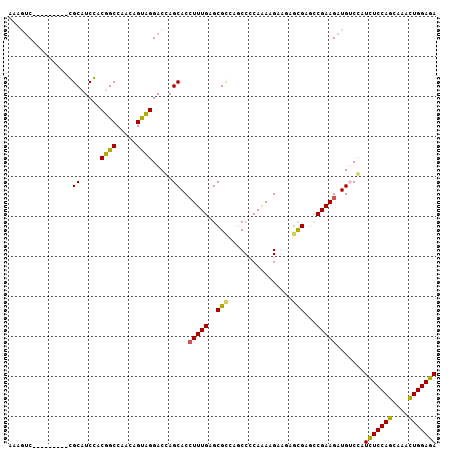
| Location | 14,954,969 – 14,955,067 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 80.66 |
| Mean single sequence MFE | -37.19 |
| Consensus MFE | -22.33 |
| Energy contribution | -22.00 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.730525 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14954969 98 - 27905053 UCUCCAGUUUGCUGGAGAUCGACAUCUUCGGCUCGCUUUUCUUUUGGGGUAUACGCUCAAAGGUGCUGGUCCUGCUGUUGGCAGUCGAUGCG---------GACUUU (((((((....)))))))(((.((((...((((.((....(((((((((....).)))))))).)).))))((((.....))))..))))))---------)..... ( -36.00) >DroPse_CAF1 4636 107 - 1 UCUCCAGCUUGCUGGAGAUGGCCAUCUUCGGCUCGUUCUUCUUCUGCGGCUGGCGCUCAAAGGUGCUCGUCCGACUGUUGGCCGUGGAGGCCACAUUCAAUGAUUUU (((((((....)))))))(((((.(((.(((((..(..........((((.((((((....)))))).).)))...)..))))).)))))))).............. ( -41.22) >DroEre_CAF1 21865 98 - 1 UCUCCAGUUUGCUGGAGAUGGACAUCUUCGACUCGCUCUUCUUUUGGGGUAUGCGCUCAAAGGUGCUCGUCCUGCUGUUGGCAGUCGAUGCG---------GACUUU (((((((....))))))).(..((((((.((..(((.((((....))))...))).)).))))))..)((((.((.((((.....)))))))---------)))... ( -34.30) >DroYak_CAF1 5489 98 - 1 UCUCCAGUUUGCUGGAGAUGGACAUCUUCGCUUCGCUAUUCUUUUGGGGCGCGCGCUCAAAUGUGCUGGUCCUACUGUUGGCAGCCGAUGCG---------GACUUU (((((((....))))))).((((.....(((((((.........))))))).((((......))))..))))....(((.(((.....))).---------)))... ( -35.00) >DroAna_CAF1 5361 98 - 1 UUUCCAACUUGCUGGAAAUCGACAUCUUCGGUUCACUCAUCUUCUGGGGCUGGCGCUCAAAGGUGCUGGUCCGACUAUUGGCCGUGGAGGCC---------GCCUUG ((((((......))))))...........(((..............((((..(((((....)))))..)))).......((((.....))))---------)))... ( -32.90) >DroPer_CAF1 4640 107 - 1 UCUCCAGCUUGCUGGAGAUGGCCAUCUUCGGCUCGCUCUUCUUCUGCGGCUGGCGCUCAAAGGUGCUCGUCCGACUGUUGGCCGUGGAGGCCACAUUCAAUGAUUUU (((((((....)))))))(((((.(((.(((((.((..........((((.((((((....)))))).).)))...)).))))).)))))))).............. ( -43.72) >consensus UCUCCAGCUUGCUGGAGAUGGACAUCUUCGGCUCGCUCUUCUUCUGGGGCUGGCGCUCAAAGGUGCUCGUCCGACUGUUGGCAGUCGAGGCC_________GACUUU (((((((....)))))))............................((((..((((......))))..))))........(((.....)))................ (-22.33 = -22.00 + -0.33)



| Location | 14,955,027 – 14,955,117 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 80.03 |
| Mean single sequence MFE | -35.38 |
| Consensus MFE | -23.84 |
| Energy contribution | -23.87 |
| Covariance contribution | 0.03 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.77 |
| SVM RNA-class probability | 0.846921 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14955027 90 - 27905053 ACUGCUGCGCUGGGCAGC---AUCCGCUGCAUCC---AUGGAUCGCUGCUCGAAGAUCUCCAGUUUGCUGGAGAUCGACAUCUUCGGCUCGCUUUU ...((.(.((((((((((---(((((.((....)---)))))).)))))))(..(((((((((....)))))))))..)......))).))).... ( -38.90) >DroPse_CAF1 4703 93 - 1 GCUGCUGCGCUGGGCCGCCGUGUCCGUGGCAUCG---CUGGAUCGCUGCUCGAAGAUCUCCAGCUUGCUGGAGAUGGCCAUCUUCGGCUCGUUCUU ...((((.....((((((.(((((((((....))---).))).))).))......((((((((....)))))))))))).....))))........ ( -37.30) >DroGri_CAF1 4725 90 - 1 GCUGCUGCGCUGAGCGCU---GUCCGUGCUGUCG---UGGGCUCGCUGCUCGAAGAUCUCCAGCUUGCUGGAUAUGGACAUCUUUGUCUCAGCUUU ((....))((((((((.(---(((((((..(((.---(((((.....)))))..))).(((((....)))))))))))))....)).))))))... ( -33.70) >DroEre_CAF1 21923 90 - 1 ACUGCUUCGCUGGGCAGC---AUCCGCUGCAUCC---AUGGAUCGUUGCUCGAAGAUCUCCAGUUUGCUGGAGAUGGACAUCUUCGACUCGCUCUU ........((.(((((((---(((((.((....)---)))))).)))))((((((((((((((....)))))))......))))))))).)).... ( -32.50) >DroAna_CAF1 5419 93 - 1 GCUGCUCCUUUGGGCAGC---AUCUGUGGCGUCGUUGGUGGAACGCUGCUCGAAGAUUUCCAACUUGCUGGAAAUCGACAUCUUCGGUUCACUCAU (((((((....)))))))---....((((.(((.((((..(....)..).))).((((((((......)))))))))))..(....).)))).... ( -30.90) >DroPer_CAF1 4707 93 - 1 GCUGCUGCGCUGGGCCGCCGUGUCCGUGGCAUCG---CUGGAUCGCUGCUCGAAGAUCUCCAGCUUGCUGGAGAUGGCCAUCUUCGGCUCGCUCUU ((.((((.....((((((.(((((((((....))---).))).))).))......((((((((....)))))))))))).....))))..)).... ( -39.00) >consensus GCUGCUGCGCUGGGCAGC___AUCCGUGGCAUCG___AUGGAUCGCUGCUCGAAGAUCUCCAGCUUGCUGGAGAUGGACAUCUUCGGCUCGCUCUU ...........(((((((...((((..............)))).)))))))((((((((((((....)))))))......)))))........... (-23.84 = -23.87 + 0.03)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:03:35 2006