
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 14,900,454 – 14,900,749 |
| Length | 295 |
| Max. P | 0.700469 |

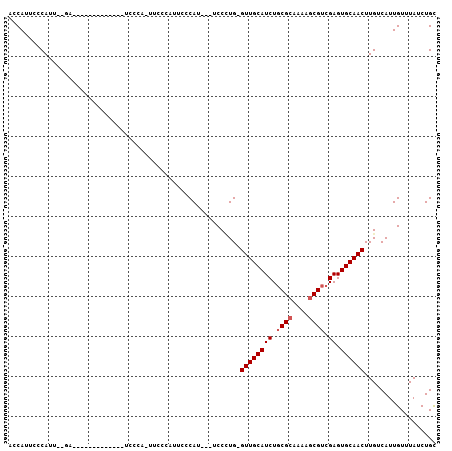
| Location | 14,900,454 – 14,900,551 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 80.07 |
| Mean single sequence MFE | -15.82 |
| Consensus MFE | -12.97 |
| Energy contribution | -13.30 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.27 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.700469 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
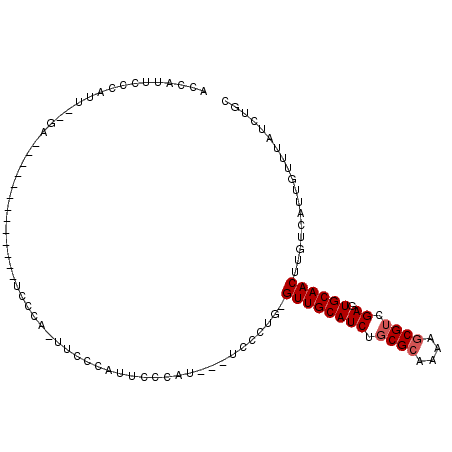
>3R_DroMel_CAF1 14900454 97 + 27905053 ACCAUUCCCAUUACCAU-----UUCCUAUUCCCA-UUCCCAUUCCCAU---UCCCUG-GUUGCAUCUGCGCAAAAGCGUCGAGUGCAACUUGUCAUUGUUUAUCUGC .................-----............-..........((.---.....(-((((((((.((((....)))).)).)))))))......))......... ( -15.90) >DroSec_CAF1 28021 84 + 1 ACCAUUCCCAU------------------UCUUA-UUCCCAUUCCCAU---UCCCUG-GUUGCAUCUGCGCAAAAGCGUCGAGUGCAACUUGUCAUUGUUUAUCUGC ...........------------------.....-..........((.---.....(-((((((((.((((....)))).)).)))))))......))......... ( -15.90) >DroSim_CAF1 27924 85 + 1 ACCAUUCCCAU-----------U------UCCUA-UUCCCAUUCCCAU---UCCCUG-GUUGCAUCUGCGCAAAAGCGUCGAGUGCAACUUGUCAUUGUUUAUCUGC ...........-----------.------.....-..........((.---.....(-((((((((.((((....)))).)).)))))))......))......... ( -15.90) >DroEre_CAF1 27507 103 + 1 CCCAUUCCCAUUUUGAAUCCCAUCCCCAUUCCCAUUUCCUAUUCCCAU---UCCUUG-GUUGCAUCUGCGAAAAAGCGUCGAGUGCAACUUGUCAUUGUUUAUCUGC .............................................((.---.....(-((((((((.(((......))).)).)))))))......))......... ( -12.50) >DroYak_CAF1 34514 97 + 1 ACCAUUCCCAUUUUGAAACC------CAUUCCCAUUUCCUAUUUCCAU---UCCAUG-GUUGCAUCUGCGCCAAAGCGUCGAGUGCAACUUGUCAUUGUUUAUCUGC ....................------...................((.---.....(-((((((((.((((....)))).)).)))))))......))......... ( -15.40) >DroAna_CAF1 26807 94 + 1 UGCACUCCCAUUGCGAU------------UCUGA-CUCUGUCUCCCAUGUCUCCCUGUGUUGCAUCUCCGCCAAAGCGUCGAGUGCAACUUGUCAUUGUUUAUCUGU (((((((....((((((------------.(.((-(............))).....).))))))....(((....)))..))))))).................... ( -19.30) >consensus ACCAUUCCCAUU__GA_____________UCCCA_UUCCCAUUCCCAU___UCCCUG_GUUGCAUCUGCGCAAAAGCGUCGAGUGCAACUUGUCAUUGUUUAUCUGC ..........................................................((((((((.((((....)))).)).)))))).................. (-12.97 = -13.30 + 0.33)


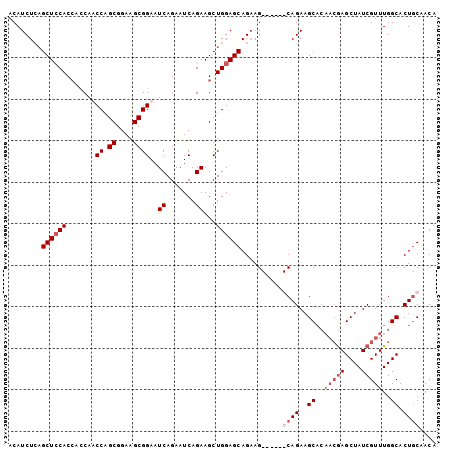
| Location | 14,900,551 – 14,900,651 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 87.91 |
| Mean single sequence MFE | -27.52 |
| Consensus MFE | -18.23 |
| Energy contribution | -20.23 |
| Covariance contribution | 2.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.66 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14900551 100 - 27905053 ACAUCUCAGCUCCACCACCAUCCAGCGGAAGCGGAAUCAGAAUCAGAAGCUGGAGCAGAAGCAGAAGCAGAAGCCG---AAGCUAUCGUUUGGCACUGCCACA ........((((((.(....(((.((....))))).((.......)).).))))))..........((((..(((.---((((....))))))).)))).... ( -29.00) >DroSec_CAF1 28105 97 - 1 ACAUCUCAGCUCCACCACCAACCAGCGGAAGCGGAAUCAGAAUCAGAAGCUGGAGCAGCAG------CAGAAGCACAACGAGCUAUCGUUUGGCACUCCAACA ........((((((.(.....((.((....))))..((.......)).).)))))).((.(------(....)).((((((....))).)))))......... ( -23.60) >DroSim_CAF1 28009 103 - 1 ACAUCUCCGCUCCACCACCAACCAGCGGAAGCGGAAUCAGAAUCAGAAGCUGGAGCAGAAGCAGAAGCAGAAGCACAACGAGCUAUCGUUUGGCACUGCAACA .....((.((((((.(.....((.((....))))..((.......)).).)))))).)).......((((..((..(((((....)))))..)).)))).... ( -29.00) >DroEre_CAF1 27610 94 - 1 ACAUCUGAGCUGCAC---CAUCCAGCGGAAGCGGAAUCAGAAUCAGAAGCUGGAGCAGAAG------CAGAAGCACAACGAGCUAUCGUUUGGCACUGCCACA ...((((..((((.(---(((((.((....))))).((.......))...))).))))...------)))).((..(((((....)))))..))......... ( -28.50) >consensus ACAUCUCAGCUCCACCACCAACCAGCGGAAGCGGAAUCAGAAUCAGAAGCUGGAGCAGAAG______CAGAAGCACAACGAGCUAUCGUUUGGCACUGCAACA ........((((((.......((.((....))))..((.......))...))))))..........((((..((..(((((....)))))..)).)))).... (-18.23 = -20.23 + 2.00)


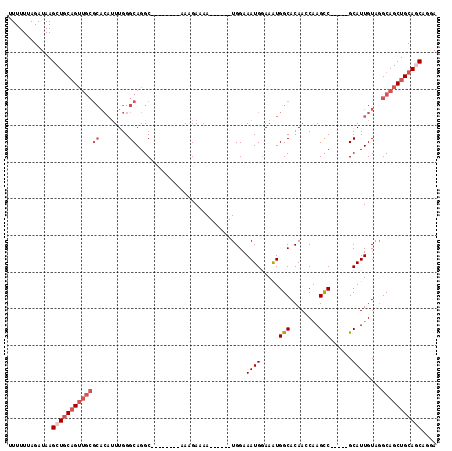
| Location | 14,900,651 – 14,900,749 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 83.35 |
| Mean single sequence MFE | -28.52 |
| Consensus MFE | -18.97 |
| Energy contribution | -20.38 |
| Covariance contribution | 1.42 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.579561 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14900651 98 - 27905053 UUUUUUAGAUAAGCUGCAGUUGCGCACAUUUGGGCAGGC--------AAAGAAAA------UGGAAAUGGAAAUGGCACAAGCAAGCC-----GCAUUGUAGGCAGCUGCAGCAGGA ............(((((((((((..(((.....((.(((--------.......(------(....)).......((....))..)))-----))..)))..))))))))))).... ( -31.50) >DroSec_CAF1 28202 98 - 1 UUUUUUAGAUAAGCUGCAGUUGCGCACAUUUGGGCAGGC--------AAGGAAAA------UGGAAAUGGAAGUGGCACAAGCAAGCC-----GCAUUGUAGGCAGCUGCAGCAGGA ............(((((((((((((........))..((--------((.....(------(....))....(((((........)))-----)).))))..))))))))))).... ( -33.40) >DroSim_CAF1 28112 112 - 1 UUUUUUAGAUAAGCUGCAGUUGCGCACAUUUGGGCAGGCAGGCAGGCAAAGAAAAUGGAAAUGGAAAUGGAAAUGGCACAAGCAAGCC-----GCAUUGUAGGCAGCUGCAGCAGGA ............(((((((((((..(((..(((((..((..((...(.......((....)).......).....))....))..)))-----.)).)))..))))))))))).... ( -34.34) >DroEre_CAF1 27704 101 - 1 -UUUUAAGAUAAGCUGCUGUUGCGCACAUUUGGGCAGGC----AAGCAAAGAAAA------UGGAAAUGGAAAUGGCACAACCAAGUC-----GCAUUGUAGGCAGCUGCAGCAGAA -...........(((((.(((((...(((((..((....----..)).....)))------))..((((((..(((.....)))..))-----.))))....))))).))))).... ( -28.20) >DroYak_CAF1 34702 96 - 1 -UUUUUAGAUAAGCUGCAG----------UUGGGCAGGC----AGGCAAAGAAAA------UGGAAAUGAAAAUGGCACAACCAAGUCAUGUUGCAUUGUAGGCAGCUGCAACAGGA -...........(.(((((----------(...((..((----((((((......------.....((((...(((.....)))..)))).)))).))))..)).)))))).).... ( -23.00) >DroAna_CAF1 26943 97 - 1 CUUUUAAGAUAAGCU-CAGUUGCGCACAUUUGGGCAGGC--------AAGGAAAA------UGGAAAUGGAAAUGGCACAACCGAGUC-----GCAUUGUAUGCAGCUGCAACUUCA .......((......-((((((((.(((.....((.(((--------..((....------(((..((....))..).)).))..)))-----))..))).)))))))).....)). ( -20.70) >consensus UUUUUUAGAUAAGCUGCAGUUGCGCACAUUUGGGCAGGC________AAAGAAAA______UGGAAAUGGAAAUGGCACAACCAAGCC_____GCAUUGUAGGCAGCUGCAGCAGGA ............(((((((((((((........))..............................((((.....(((........)))......))))....))))))))))).... (-18.97 = -20.38 + 1.42)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:01:30 2006