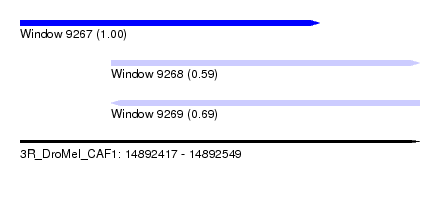
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 14,892,417 – 14,892,549 |
| Length | 132 |
| Max. P | 0.995649 |

| Location | 14,892,417 – 14,892,516 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 95.16 |
| Mean single sequence MFE | -23.88 |
| Consensus MFE | -19.94 |
| Energy contribution | -20.38 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.64 |
| Structure conservation index | 0.84 |
| SVM decision value | 2.60 |
| SVM RNA-class probability | 0.995649 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14892417 99 + 27905053 UGGCAAUUUCUUUUGCCAGUCUGUUUUUUUUUUGUAACAACAGCAAUUAUUAGCUGUUAGUUGUAGGCAACAUGGAAAUUCGAUAUCAAAUUGAAAAAA ((((((......))))))..........((((..(...((((((........)))))).((((....)))))..))))((((((.....)))))).... ( -23.00) >DroSec_CAF1 20049 96 + 1 UGGCAAUUUCUUUUGCCAGUCUGUUUU---UUUGUAACAACAGCAAUUAUUAGCUGUUAGUUGUAGGCAACAUGGAAAUUCGAUAUCAAAUUGAAAAAC ((((((......))))))(((.(..((---((..(...((((((........)))))).((((....)))))..))))..))))............... ( -23.00) >DroSim_CAF1 19921 96 + 1 UGGCAAUUUCUUUUGCCAGUCUGUUUU---UUUGCAACAACAGCAAUUAUUAGCUGUUAGUUGUAGGCAACAUGGAAAUUCGAUAUCAAAUUGAAAAAC ((((((......)))))).(((((.((---((((((((((((((........)))))).)))))))).)).)))))..((((((.....)))))).... ( -27.90) >DroEre_CAF1 19657 95 + 1 UGGCAAUUUCUUUUUCCAGUCUGUUU----UUUGUAACAACAGCAAUUAUUAGCUGUUAGUUGUAGGCAACAUGGAAAUUCGAUAUGAAAUUGAAAAGC ...(((((((..((((((...((((.----((((((((((((((........)))))).)))))))).))))))))))........)))))))...... ( -23.50) >DroYak_CAF1 22623 95 + 1 UGGCAAUUUCUUUUUCCAGUCUGUUU----UUUGUAACUACAGCAAUUAUUAGCUGUUAGUUGUAGGCAACAUGGAAAUUCGAUAUAAAAUUGAAAAAC ............((((((...((((.----((((((((((((((........)))).)))))))))).))))))))))((((((.....)))))).... ( -22.00) >consensus UGGCAAUUUCUUUUGCCAGUCUGUUUU___UUUGUAACAACAGCAAUUAUUAGCUGUUAGUUGUAGGCAACAUGGAAAUUCGAUAUCAAAUUGAAAAAC ((((((......)))))).(((((........((((((((((((........)))))).))))))(....))))))..((((((.....)))))).... (-19.94 = -20.38 + 0.44)



| Location | 14,892,447 – 14,892,549 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 87.84 |
| Mean single sequence MFE | -22.06 |
| Consensus MFE | -16.04 |
| Energy contribution | -17.84 |
| Covariance contribution | 1.80 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.588323 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14892447 102 + 27905053 UUUGUAACAACAGCAAUUAUUAGCUGUUAGUUGUAGGCAACAUGGAAAUUCGAUAUCAAAUUGAAAAAACC-CGAGGGGUAA--GGAAAAGCGAAUGUUGUUUUG ....((((((((((........)))))).)))).(((((((((.....((((((.....))))))...(((-(...))))..--..........))))))))).. ( -23.20) >DroSec_CAF1 20076 104 + 1 UUUGUAACAACAGCAAUUAUUAGCUGUUAGUUGUAGGCAACAUGGAAAUUCGAUAUCAAAUUGAAAAACCCAAGAGGGGCAAGAGGAAAAGCGAAUGUUGUU-UG ....((((((((((........)))))).))))((((((((((.....((((((.....))))))...(((....)))................))))))))-)) ( -24.80) >DroSim_CAF1 19948 104 + 1 UUUGCAACAACAGCAAUUAUUAGCUGUUAGUUGUAGGCAACAUGGAAAUUCGAUAUCAAAUUGAAAAACCCAAGAGGGGCAAGAGGAAAAGCGAAUGUUGUU-UG .(((((((((((((........)))))).)))))))(((((((.....((((((.....))))))...(((....)))................))))))).-.. ( -28.40) >DroEre_CAF1 19683 92 + 1 UUUGUAACAACAGCAAUUAUUAGCUGUUAGUUGUAGGCAACAUGGAAAUUCGAUAUGAAAUUGAAAAGCCC----GGGGUAAG-----AAGUGAAUG---UU-UG .(((((((((((((........)))))).)))))))((..(..((...((((((.....))))))...)).----)..))...-----.........---..-.. ( -16.00) >DroYak_CAF1 22649 92 + 1 UUUGUAACUACAGCAAUUAUUAGCUGUUAGUUGUAGGCAACAUGGAAAUUCGAUAUAAAAUUGAAAAACCC----GCGGUAAG-----AAGUGAGUG---UU-UG .(((((((((((((........)))).)))))))))..(((((((...((((((.....))))))...))(----((......-----..))).)))---))-.. ( -17.90) >consensus UUUGUAACAACAGCAAUUAUUAGCUGUUAGUUGUAGGCAACAUGGAAAUUCGAUAUCAAAUUGAAAAACCC__GAGGGGUAAG_GGAAAAGCGAAUGUUGUU_UG .(((((((((((((........)))))).)))))))(((((((.....((((((.....))))))..((((.....))))..............))))))).... (-16.04 = -17.84 + 1.80)



| Location | 14,892,447 – 14,892,549 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 87.84 |
| Mean single sequence MFE | -17.44 |
| Consensus MFE | -11.64 |
| Energy contribution | -12.04 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.37 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.690321 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14892447 102 - 27905053 CAAAACAACAUUCGCUUUUCC--UUACCCCUCG-GGUUUUUUCAAUUUGAUAUCGAAUUUCCAUGUUGCCUACAACUAACAGCUAAUAAUUGCUGUUGUUACAAA .....((((((..........--..((((...)-)))......((((((....))))))...)))))).....(((.((((((........)))))))))..... ( -18.50) >DroSec_CAF1 20076 104 - 1 CA-AACAACAUUCGCUUUUCCUCUUGCCCCUCUUGGGUUUUUCAAUUUGAUAUCGAAUUUCCAUGUUGCCUACAACUAACAGCUAAUAAUUGCUGUUGUUACAAA ..-..((((((..............((((.....)))).....((((((....))))))...)))))).....(((.((((((........)))))))))..... ( -19.70) >DroSim_CAF1 19948 104 - 1 CA-AACAACAUUCGCUUUUCCUCUUGCCCCUCUUGGGUUUUUCAAUUUGAUAUCGAAUUUCCAUGUUGCCUACAACUAACAGCUAAUAAUUGCUGUUGUUGCAAA ..-..((((((..............((((.....)))).....((((((....))))))...))))))....((((.((((((........)))))))))).... ( -22.20) >DroEre_CAF1 19683 92 - 1 CA-AA---CAUUCACUU-----CUUACCCC----GGGCUUUUCAAUUUCAUAUCGAAUUUCCAUGUUGCCUACAACUAACAGCUAAUAAUUGCUGUUGUUACAAA ..-..---.........-----........----((((........(((.....)))..........))))..(((.((((((........)))))))))..... ( -14.07) >DroYak_CAF1 22649 92 - 1 CA-AA---CACUCACUU-----CUUACCGC----GGGUUUUUCAAUUUUAUAUCGAAUUUCCAUGUUGCCUACAACUAACAGCUAAUAAUUGCUGUAGUUACAAA ..-..---.........-----......(.----.(((.....(((((......)))))........)))..)(((((.((((........)))))))))..... ( -12.72) >consensus CA_AACAACAUUCGCUUUUCC_CUUACCCCUC__GGGUUUUUCAAUUUGAUAUCGAAUUUCCAUGUUGCCUACAACUAACAGCUAAUAAUUGCUGUUGUUACAAA ..................................(((...(((.((.....)).)))..)))..((((....))))(((((((........)))))))....... (-11.64 = -12.04 + 0.40)


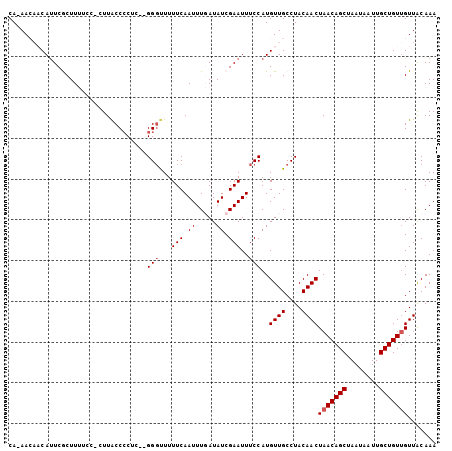
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:01:18 2006