
| Sequence ID | 3R_DroMel_CAF1 |
|---|---|
| Location | 14,829,239 – 14,829,382 |
| Length | 143 |
| Max. P | 0.969263 |

| Location | 14,829,239 – 14,829,342 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.52 |
| Mean single sequence MFE | -29.83 |
| Consensus MFE | -24.55 |
| Energy contribution | -25.83 |
| Covariance contribution | 1.28 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.65 |
| Structure conservation index | 0.82 |
| SVM decision value | 1.64 |
| SVM RNA-class probability | 0.969263 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
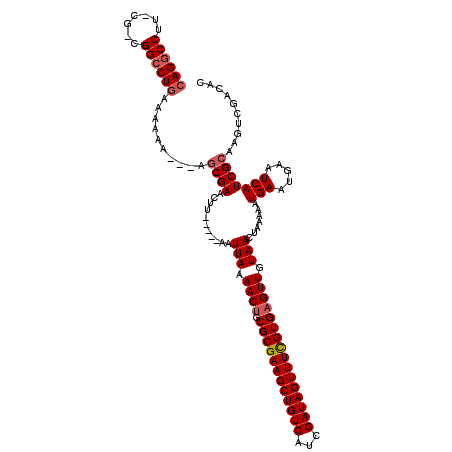
>3R_DroMel_CAF1 14829239 103 + 27905053 CAG------------CCUAAAAAAAAAUAACGAGCUU-----AAUUAAAACAGCGCGAAGCUGUCAUCGAUAGUUUCGUGAGUUGUAACUUAAAAUGAAUGAAUCAUCGCAAGUCGACAC ...------------...............(((.(((-----.......((((((((((((((((...)))))))))))..)))))........((((.....))))...)))))).... ( -21.00) >DroSec_CAF1 7379 112 + 1 CAGGCCUUACGACGGCCUGAAAAAA---AGCGAACUU-----AAUUAAAACUGCGCGAAGCUGUCAUCGAUAGUUUUGUGAGUUGUAACUAAAAAUGAAUGAAUCAUCGCAAGUCGACGC ((((((.......))))))......---.((((..((-----(.(((.((((.((((((((((((...)))))))))))))))).))).)))...(((.....))))))).......... ( -33.50) >DroSim_CAF1 7540 116 + 1 CAGGCCUUUCGGCGGCCUGAAA-AA---AGCGAACUUACCAUAAUUAAAACUGCGCGAAGCUGUCAUCGAUAGUUUCGUGAGUUGUAACUAAAAAUGAAUGAAUCAUCGCAAGUCGACAC ((((((.......))))))...-..---.((((...........(((.((((.((((((((((((...)))))))))))))))).))).......(((.....))))))).......... ( -35.00) >consensus CAGGCCUU_CG_CGGCCUGAAAAAA___AGCGAACUU_____AAUUAAAACUGCGCGAAGCUGUCAUCGAUAGUUUCGUGAGUUGUAACUAAAAAUGAAUGAAUCAUCGCAAGUCGACAC ((((((.......))))))..........((((...........(((.((((.((((((((((((...)))))))))))))))).))).......(((.....))))))).......... (-24.55 = -25.83 + 1.28)



| Location | 14,829,264 – 14,829,382 |
|---|---|
| Length | 118 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 89.80 |
| Mean single sequence MFE | -27.53 |
| Consensus MFE | -23.22 |
| Energy contribution | -23.33 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.551332 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3R_DroMel_CAF1 14829264 118 + 27905053 --AAUUAAAACAGCGCGAAGCUGUCAUCGAUAGUUUCGUGAGUUGUAACUUAAAAUGAAUGAAUCAUCGCAAGUCGACACAAUGUUGCUUUAGUGCUAUUCAUUUCCUUAGAAACUGCGU --.......((((((((((((((((...)))))))))))..)))))..........((((((((...((((((.((((.....)))))))..)))..))))))))............... ( -28.00) >DroSec_CAF1 7413 113 + 1 --AAUUAAAACUGCGCGAAGCUGUCAUCGAUAGUUUUGUGAGUUGUAACUAAAAAUGAAUGAAUCAUCGCAAGUCGACGCUAUGUUGUUUUAGUGCUAAACAAUUCCUUAGAAAU----- --........(((.(((((((((((...)))))))))))(((((((.(((((((....(((..((..(....)..))..).))....))))))).....)))))))..)))....----- ( -25.00) >DroSim_CAF1 7576 115 + 1 AUAAUUAAAACUGCGCGAAGCUGUCAUCGAUAGUUUCGUGAGUUGUAACUAAAAAUGAAUGAAUCAUCGCAAGUCGACACUAUGUUGUUUUAGUGCUGAACAAUUCCUUAGAAAU----- ....(((.((((.((((((((((((...)))))))))))))))).)))((((....((((...(((.((((((.((((.....)))).))).))).)))...)))).))))....----- ( -29.60) >consensus __AAUUAAAACUGCGCGAAGCUGUCAUCGAUAGUUUCGUGAGUUGUAACUAAAAAUGAAUGAAUCAUCGCAAGUCGACACUAUGUUGUUUUAGUGCUAAACAAUUCCUUAGAAAU_____ ...((((((((..((((((((((((...)))))))))))).((((..(((....((((.....))))....)))))))........)))))))).......................... (-23.22 = -23.33 + 0.11)

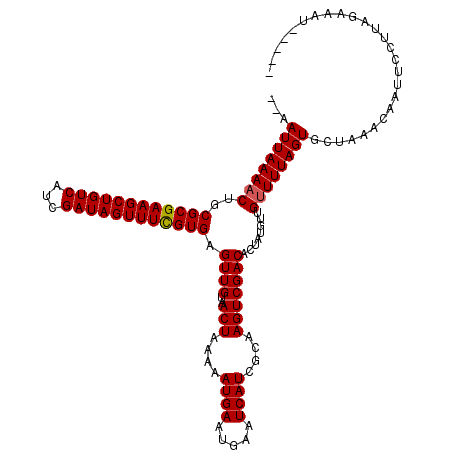

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:00:08 2006